

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0074.9
(351 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
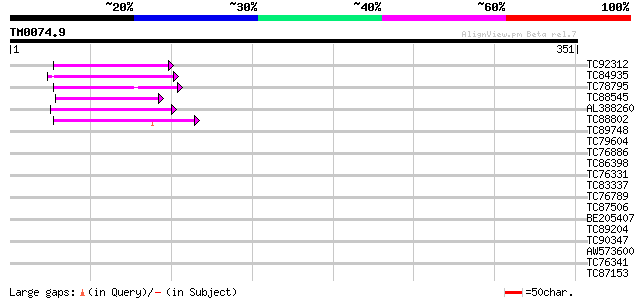
Score E
Sequences producing significant alignments: (bits) Value
TC92312 weakly similar to GP|4567275|gb|AAD23688.1| expressed pr... 44 9e-05
TC84935 similar to PIR|G96631|G96631 probable RNA-binding protei... 44 9e-05
TC78795 similar to GP|15081787|gb|AAK82548.1 AT4g36960/C7A10_400... 42 4e-04
TC88545 similar to GP|13560783|gb|AAK30205.1 poly(A)-binding pro... 42 4e-04
AL388260 weakly similar to SP|Q08937|ROC2_ 29 kDa ribonucleoprot... 41 8e-04
TC88802 similar to GP|20804774|dbj|BAB92459. hypothetical protei... 41 8e-04
TC89748 homologue to PIR|B84565|B84565 probable spliceosome asso... 40 0.001
TC79604 similar to GP|18175762|gb|AAL59923.1 unknown protein {Ar... 40 0.001
TC76886 similar to PIR|T01932|T01932 RNA binding protein homolog... 40 0.001
TC86398 similar to GP|16226341|gb|AAL16140.1 AT3g11400/F24K9_7 {... 40 0.002
TC76331 similar to GP|7673355|gb|AAF66823.1| poly(A)-binding pro... 39 0.002
TC83337 similar to GP|15081787|gb|AAK82548.1 AT4g36960/C7A10_400... 39 0.003
TC76789 similar to GP|15294254|gb|AAK95304.1 AT4g24770/F22K18_30... 39 0.004
TC87506 similar to GP|9843655|emb|CAC03601.1 SC35-like splicing ... 39 0.004
BE205407 weakly similar to GP|10716605|gb putative RNA binding p... 38 0.005
TC89204 similar to GP|16226341|gb|AAL16140.1 AT3g11400/F24K9_7 {... 38 0.007
TC90347 similar to GP|10334491|emb|CAC10207. putative splicing f... 37 0.011
AW573600 similar to PIR|T01932|T01 RNA binding protein homolog -... 37 0.011
TC76341 similar to GP|7528270|gb|AAF63202.1| poly(A)-binding pro... 37 0.011
TC87153 similar to GP|20804774|dbj|BAB92459. hypothetical protei... 36 0.019
>TC92312 weakly similar to GP|4567275|gb|AAD23688.1| expressed protein
{Arabidopsis thaliana}, partial (29%)
Length = 1319
Score = 43.9 bits (102), Expect = 9e-05
Identities = 23/74 (31%), Positives = 39/74 (52%)
Frame = +1
Query: 28 RLFVDGLDRDTKHSQVRKLFERFGTVVSVFVPKTIKRVRREKFGFVLFQSKRDGIAAMRS 87
+L V L K +++R F GTV VF+P+ + F FV F K+D A+R
Sbjct: 277 KLIVRNLPFKAKENEIRDAFSSAGTVWEVFIPQKSDTGLSKGFAFVKFTCKQDAENAIRK 456
Query: 88 LHGTRLNSFFLSIN 101
L+G++ S ++++
Sbjct: 457 LNGSKFGSRLIAVD 498
>TC84935 similar to PIR|G96631|G96631 probable RNA-binding protein F8A5.17
[imported] - Arabidopsis thaliana, partial (41%)
Length = 552
Score = 43.9 bits (102), Expect = 9e-05
Identities = 24/81 (29%), Positives = 40/81 (48%)
Frame = +2
Query: 24 EENIRLFVDGLDRDTKHSQVRKLFERFGTVVSVFVPKTIKRVRREKFGFVLFQSKRDGIA 83
EEN R+FV GL D Q+ F+R+G ++ + R FGF+ F +R
Sbjct: 131 EEN-RIFVGGLGWDVTERQLEHAFDRYGKILECQIMMERDTGRPRGFGFITFSDRRGMED 307
Query: 84 AMRSLHGTRLNSFFLSINPAR 104
A++ + G + +S+N A+
Sbjct: 308 AIKEMDGREIGDRIISVNKAQ 370
>TC78795 similar to GP|15081787|gb|AAK82548.1 AT4g36960/C7A10_400 {Arabidopsis
thaliana}, partial (77%)
Length = 1751
Score = 42.0 bits (97), Expect = 4e-04
Identities = 25/80 (31%), Positives = 40/80 (49%)
Frame = +3
Query: 28 RLFVDGLDRDTKHSQVRKLFERFGTVVSVFVPKTIKRVRREKFGFVLFQSKRDGIAAMRS 87
++FV L + +R+ F RFG + V++P+ KR FGFV F +G+A S
Sbjct: 954 KIFVGRLPPEANSEDLRQYFGRFGRIEDVYIPRDPKRTGHRGFGFVTFAD--EGVADRVS 1127
Query: 88 LHGTRLNSFFLSINPARFVD 107
L + ++I+ A VD
Sbjct: 1128 LRPHEICGHEVAIDSATPVD 1187
Score = 29.3 bits (64), Expect = 2.4
Identities = 16/70 (22%), Positives = 28/70 (39%)
Frame = +3
Query: 23 AEENIRLFVDGLDRDTKHSQVRKLFERFGTVVSVFVPKTIKRVRREKFGFVLFQSKRDGI 82
A++ R+FV + R FE++G + +++PK GF+ F +
Sbjct: 540 AKKVTRIFVARIPPSVTEETFRSHFEKYGDITDLYMPKDQGSKMHRGIGFITFGNAESVE 719
Query: 83 AAMRSLHGTR 92
M H R
Sbjct: 720 NLMSETHELR 749
>TC88545 similar to GP|13560783|gb|AAK30205.1 poly(A)-binding protein
{Daucus carota}, partial (35%)
Length = 866
Score = 42.0 bits (97), Expect = 4e-04
Identities = 24/67 (35%), Positives = 36/67 (52%)
Frame = +2
Query: 29 LFVDGLDRDTKHSQVRKLFERFGTVVSVFVPKTIKRVRREKFGFVLFQSKRDGIAAMRSL 88
L+V LD D SQ+ LF + G VVSV + + + + +G+V F + D AM L
Sbjct: 185 LYVGDLDHDVTDSQLYDLFNQIGQVVSVRICRDLASQQSLGYGYVNFSNPHDAAKAMDVL 364
Query: 89 HGTRLNS 95
+ T LN+
Sbjct: 365 NFTPLNN 385
>AL388260 weakly similar to SP|Q08937|ROC2_ 29 kDa ribonucleoprotein B
chloroplast precursor (CP29B). [Wood tobacco] {Nicotiana
sylvestris}, partial (26%)
Length = 346
Score = 40.8 bits (94), Expect = 8e-04
Identities = 23/78 (29%), Positives = 38/78 (48%)
Frame = +3
Query: 26 NIRLFVDGLDRDTKHSQVRKLFERFGTVVSVFVPKTIKRVRREKFGFVLFQSKRDGIAAM 85
+++LFV GL T +R FE +GTV V + R FGFV F S + A+
Sbjct: 81 SVKLFVGGLSWGTDDRTLRSKFEEYGTVEDAVVIRDRDTGRSRGFGFVTFASNEEAEIAI 260
Query: 86 RSLHGTRLNSFFLSINPA 103
++L+ + + ++ A
Sbjct: 261 QNLNDAEFDGRNIKVDRA 314
>TC88802 similar to GP|20804774|dbj|BAB92459. hypothetical protein~similar
to Arabidopsis thaliana chromosome 1 F10C21_14, partial
(26%)
Length = 1200
Score = 40.8 bits (94), Expect = 8e-04
Identities = 24/94 (25%), Positives = 45/94 (47%), Gaps = 4/94 (4%)
Frame = +3
Query: 28 RLFVDGLDRDTKHSQVRKLFERFGTVVSVFVPKTIKRVRREKFGFVLFQSKRDGIAAMRS 87
++FV GL +TK +++ F++FG ++ V + + +GFV F+ I A ++
Sbjct: 327 KIFVGGLAWETKRDTLKRYFDQFGDILEAVVITDRTTGKSKGYGFVTFKDPNSAILACQN 506
Query: 88 ----LHGTRLNSFFLSINPARFVDRKYKWNPPAK 117
+ G R N P + K K+N P++
Sbjct: 507 PNPMIDGRRANCNLAYQKPDPSLTGKQKFNSPSR 608
>TC89748 homologue to PIR|B84565|B84565 probable spliceosome associated
protein [imported] - Arabidopsis thaliana, partial (56%)
Length = 707
Score = 40.0 bits (92), Expect = 0.001
Identities = 24/86 (27%), Positives = 44/86 (50%)
Frame = +2
Query: 18 ERSEVAEENIRLFVDGLDRDTKHSQVRKLFERFGTVVSVFVPKTIKRVRREKFGFVLFQS 77
+ S ++ +V LD + +LF + G VV+V+VPK + + +GFV F+S
Sbjct: 137 QHSAERNQDATAYVGNLDPQISEELLWELFVQAGPVVNVYVPKDRVTNQHQGYGFVEFRS 316
Query: 78 KRDGIAAMRSLHGTRLNSFFLSINPA 103
+ D A++ L+ +L + +N A
Sbjct: 317 EEDADYAIKVLNMIKLYGKPIRVNKA 394
>TC79604 similar to GP|18175762|gb|AAL59923.1 unknown protein {Arabidopsis
thaliana}, partial (56%)
Length = 727
Score = 40.0 bits (92), Expect = 0.001
Identities = 24/84 (28%), Positives = 39/84 (45%)
Frame = +2
Query: 21 EVAEENIRLFVDGLDRDTKHSQVRKLFERFGTVVSVFVPKTIKRVRREKFGFVLFQSKRD 80
++AE + LFV GL + T +R+ F++FG VV V + FGFV + + D
Sbjct: 221 QMAEPSTNLFVSGLSKRTTTETLREAFQKFGEVVHARVVTDRVSGYSKGFGFVKYATLED 400
Query: 81 GIAAMRSLHGTRLNSFFLSINPAR 104
+ + G L + + AR
Sbjct: 401 AAKGIEGMDGKFLEGWVIFAEYAR 472
>TC76886 similar to PIR|T01932|T01932 RNA binding protein homolog - common
tobacco (fragment), partial (67%)
Length = 1896
Score = 40.0 bits (92), Expect = 0.001
Identities = 24/73 (32%), Positives = 38/73 (51%)
Frame = +2
Query: 19 RSEVAEENIRLFVDGLDRDTKHSQVRKLFERFGTVVSVFVPKTIKRVRREKFGFVLFQSK 78
+SE N +FV GLD D +R+ F +FG V+SV +P + GFV +
Sbjct: 1178 QSEGDSNNTTIFVGGLDSDISDEDLRQPFLQFGDVISVKIPV------GKGCGFVQLADR 1339
Query: 79 RDGIAAMRSLHGT 91
++ A++ L+GT
Sbjct: 1340 KNAEEAIQGLNGT 1378
>TC86398 similar to GP|16226341|gb|AAL16140.1 AT3g11400/F24K9_7 {Arabidopsis
thaliana}, partial (75%)
Length = 1170
Score = 39.7 bits (91), Expect = 0.002
Identities = 25/77 (32%), Positives = 37/77 (47%)
Frame = +3
Query: 24 EENIRLFVDGLDRDTKHSQVRKLFERFGTVVSVFVPKTIKRVRREKFGFVLFQSKRDGIA 83
E ++R V L DT+ + +LF FG V V+V K FGFV F S+ D
Sbjct: 684 ENSVR--VTNLSEDTREPDLLELFRPFGAVSRVYVAIDQKTGMSRGFGFVNFVSREDAQR 857
Query: 84 AMRSLHGTRLNSFFLSI 100
A+ L+G ++ L +
Sbjct: 858 AINKLNGYGYDNLILRV 908
>TC76331 similar to GP|7673355|gb|AAF66823.1| poly(A)-binding protein
{Nicotiana tabacum}, partial (97%)
Length = 2581
Score = 39.3 bits (90), Expect = 0.002
Identities = 23/67 (34%), Positives = 37/67 (54%)
Frame = +3
Query: 29 LFVDGLDRDTKHSQVRKLFERFGTVVSVFVPKTIKRVRREKFGFVLFQSKRDGIAAMRSL 88
L+V LD + SQ+ LF + G VVSV V + + R +G+V + + +D A+ L
Sbjct: 450 LYVGDLDMNVTDSQLYDLFNQLGQVVSVRVCRDLTTRRSLGYGYVNYSNPQDAARALDVL 629
Query: 89 HGTRLNS 95
+ T LN+
Sbjct: 630 NFTPLNN 650
>TC83337 similar to GP|15081787|gb|AAK82548.1 AT4g36960/C7A10_400
{Arabidopsis thaliana}, partial (24%)
Length = 678
Score = 38.9 bits (89), Expect = 0.003
Identities = 17/48 (35%), Positives = 27/48 (55%)
Frame = +1
Query: 28 RLFVDGLDRDTKHSQVRKLFERFGTVVSVFVPKTIKRVRREKFGFVLF 75
++FV L + +R F RFG ++ V++P+ +KR FGFV F
Sbjct: 13 KIFVGRLPPEATTEDLRLYFGRFGHILDVYIPRDVKRPGHRGFGFVTF 156
>TC76789 similar to GP|15294254|gb|AAK95304.1 AT4g24770/F22K18_30
{Arabidopsis thaliana}, partial (62%)
Length = 1294
Score = 38.5 bits (88), Expect = 0.004
Identities = 24/83 (28%), Positives = 38/83 (44%)
Frame = +3
Query: 21 EVAEENIRLFVDGLDRDTKHSQVRKLFERFGTVVSVFVPKTIKRVRREKFGFVLFQSKRD 80
E E++++FV L D ++ +LFE+ GTV V R FGFV + +
Sbjct: 435 EEPSEDLKIFVGNLPFDVDSEKLAQLFEQSGTVEIAEVIYNRDTDRSRGFGFVTMSTSEE 614
Query: 81 GIAAMRSLHGTRLNSFFLSINPA 103
A+ G L+ L++N A
Sbjct: 615 VERAVNKFSGFELDGRLLTVNKA 683
>TC87506 similar to GP|9843655|emb|CAC03601.1 SC35-like splicing factor
SCL28 28 kD {Arabidopsis thaliana}, partial (51%)
Length = 1558
Score = 38.5 bits (88), Expect = 0.004
Identities = 27/100 (27%), Positives = 40/100 (40%), Gaps = 11/100 (11%)
Frame = +3
Query: 3 PTGSARAPPPRWSCHERSEVAEENIR-----------LFVDGLDRDTKHSQVRKLFERFG 51
P +R PPPR + +N L V L D + +R FER+G
Sbjct: 300 PRRRSRTPPPRGRKRYEDDRYPDNRSRRDRRSPLPSGLLVRNLPLDARPEDLRGPFERYG 479
Query: 52 TVVSVFVPKTIKRVRREKFGFVLFQSKRDGIAAMRSLHGT 91
V V++P+ FGFV ++ D A + L+ T
Sbjct: 480 PVKDVYLPRNYYTGEPRGFGFVKYRHGEDAAEAKQQLNHT 599
>BE205407 weakly similar to GP|10716605|gb putative RNA binding protein
{Oryza sativa}, partial (23%)
Length = 608
Score = 38.1 bits (87), Expect = 0.005
Identities = 25/74 (33%), Positives = 38/74 (50%), Gaps = 5/74 (6%)
Frame = +1
Query: 21 EVAEENIR---LFVDGLDRDTKHSQVRKLFERFGTVVSVFVPKTIKRVRREKFGFVLFQS 77
E E+IR +FV GL + + ++ FERFG + V V + + FGF+ F S
Sbjct: 379 ECGNEHIRTKKIFVGGLPANMSVEEFKRYFERFGRITDVVVMQDSLTHKPRGFGFITFDS 558
Query: 78 KRDGIAA--MRSLH 89
+ D + + MRS H
Sbjct: 559 E-DSVQSVMMRSFH 597
>TC89204 similar to GP|16226341|gb|AAL16140.1 AT3g11400/F24K9_7 {Arabidopsis
thaliana}, partial (80%)
Length = 1087
Score = 37.7 bits (86), Expect = 0.007
Identities = 23/77 (29%), Positives = 37/77 (47%)
Frame = +1
Query: 24 EENIRLFVDGLDRDTKHSQVRKLFERFGTVVSVFVPKTIKRVRREKFGFVLFQSKRDGIA 83
E ++R V L DT+ + +LF FG+V V+V FGFV F ++ D
Sbjct: 610 ENSVR--VTNLSEDTREPDLHELFGTFGSVSRVYVAIDKNTGTSRGFGFVNFVNREDAQR 783
Query: 84 AMRSLHGTRLNSFFLSI 100
A+ L+G ++ L +
Sbjct: 784 AINKLNGYGYDNLILKV 834
>TC90347 similar to GP|10334491|emb|CAC10207. putative splicing factor
{Cicer arietinum}, partial (53%)
Length = 908
Score = 37.0 bits (84), Expect = 0.011
Identities = 23/63 (36%), Positives = 36/63 (56%)
Frame = +1
Query: 28 RLFVDGLDRDTKHSQVRKLFERFGTVVSVFVPKTIKRVRREKFGFVLFQSKRDGIAAMRS 87
+L+V L + SQ+R++FE FGTV V +P ++ + FGF+ F AA +S
Sbjct: 316 KLYVGNLHFNMTESQLREIFEPFGTVEVVQLPLDLETGHCKGFGFIQFAHIEHAKAA-QS 492
Query: 88 LHG 90
L+G
Sbjct: 493 LNG 501
>AW573600 similar to PIR|T01932|T01 RNA binding protein homolog - common
tobacco (fragment), partial (23%)
Length = 522
Score = 37.0 bits (84), Expect = 0.011
Identities = 21/67 (31%), Positives = 36/67 (53%)
Frame = +3
Query: 25 ENIRLFVDGLDRDTKHSQVRKLFERFGTVVSVFVPKTIKRVRREKFGFVLFQSKRDGIAA 84
++ +FV GLD D +R+ F +FG V+SV +P + GFV +++ A
Sbjct: 99 QSTTIFVGGLDSDISDEDLRQPFLQFGDVISVKIPV------GKGCGFVQLADRKNAEEA 260
Query: 85 MRSLHGT 91
++ L+GT
Sbjct: 261 IQGLNGT 281
>TC76341 similar to GP|7528270|gb|AAF63202.1| poly(A)-binding protein
{Cucumis sativus}, partial (90%)
Length = 2501
Score = 37.0 bits (84), Expect = 0.011
Identities = 22/67 (32%), Positives = 37/67 (54%)
Frame = +3
Query: 29 LFVDGLDRDTKHSQVRKLFERFGTVVSVFVPKTIKRVRREKFGFVLFQSKRDGIAAMRSL 88
L+V L+ + SQ+ LF + G VVSV V + + R +G+V F + +D A+ L
Sbjct: 300 LYVGDLEVNVNDSQLYDLFNQVGQVVSVRVCRDLATRRSLGYGYVNFTNPQDAARALDVL 479
Query: 89 HGTRLNS 95
+ T +N+
Sbjct: 480 NFTPMNN 500
>TC87153 similar to GP|20804774|dbj|BAB92459. hypothetical protein~similar
to Arabidopsis thaliana chromosome 1 F10C21_14, partial
(50%)
Length = 920
Score = 36.2 bits (82), Expect = 0.019
Identities = 26/84 (30%), Positives = 41/84 (47%), Gaps = 7/84 (8%)
Frame = +2
Query: 28 RLFVDGLDRDTKHSQVRKLFERFGTVVSVFVPKTIKRVRREKFGFVLFQSKRDGIAAMRS 87
++FV GL +T+ ++K FE+FG ++ V R + +GFV F R+ AAMR+
Sbjct: 323 KVFVGGLAWETQKETMKKYFEQFGEILEAVVITDKATGRSKGYGFVTF---REPDAAMRA 493
Query: 88 -------LHGTRLNSFFLSINPAR 104
+ G R N S+ R
Sbjct: 494 CVDAAPIIDGRRANCNLASLGVQR 565
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.321 0.136 0.408
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 11,249,555
Number of Sequences: 36976
Number of extensions: 149493
Number of successful extensions: 877
Number of sequences better than 10.0: 76
Number of HSP's better than 10.0 without gapping: 871
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 877
length of query: 351
length of database: 9,014,727
effective HSP length: 97
effective length of query: 254
effective length of database: 5,428,055
effective search space: 1378725970
effective search space used: 1378725970
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 59 (27.3 bits)
Lotus: description of TM0074.9


