

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0074.3
(267 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
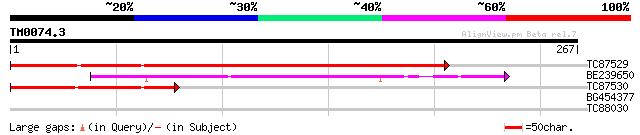
TC87529 similar to GP|15293029|gb|AAK93625.1 unknown protein {Ar... 324 2e-89
BE239650 similar to PIR|D87615|D876 conserved hypothetical prote... 129 1e-30
TC87530 118 2e-27
BG454377 similar to PIR|A85055|A85 probable leucyl tRNA syntheta... 28 4.8
TC88030 similar to PIR|F86416|F86416 probable RNA-binding protei... 27 8.2
>TC87529 similar to GP|15293029|gb|AAK93625.1 unknown protein {Arabidopsis
thaliana}, partial (54%)
Length = 687
Score = 324 bits (831), Expect = 2e-89
Identities = 154/207 (74%), Positives = 182/207 (87%)
Frame = +1
Query: 1 MALKDTFYISHGSPTLSIDESLVARKFLQSWKKEVFPPRPTSILVISGHWDTAVPTVNVV 60
MALK+TFYISHGSPTL+IDE++ A KFL SWK EVFP RP++ILVISGHWDT+VPTVNVV
Sbjct: 73 MALKETFYISHGSPTLAIDETIPAWKFLTSWK-EVFPERPSAILVISGHWDTSVPTVNVV 249
Query: 61 DSTNDTIYDFYGFPKPMYQLKYPAPGAPHLAKRVKELLKEGGFSRVDEDKKRGLDHGAWV 120
+ N+TI+DF GFP+ MY+LKYPAPGAP LAKRVKEL++ G SRVDEDKKRGLDHG WV
Sbjct: 250 NH-NETIHDFGGFPRSMYKLKYPAPGAPKLAKRVKELIEASGLSRVDEDKKRGLDHGTWV 426
Query: 121 PLLLMYPEADIPVCQLSVQSNLDGTHHYNIGKALAPLKDEGVLIVGSGSAVHNLRALERH 180
PL+LMYPEADIPVCQLSV SN +GT+HYN+GKA+APLKDEGVLI+GSGSA HN+RA+
Sbjct: 427 PLMLMYPEADIPVCQLSVSSNRNGTYHYNLGKAIAPLKDEGVLIIGSGSATHNMRAIGPR 606
Query: 181 ATVAAPWAVEFDNWLKEALLEGRYEDV 207
+ PWA+ FD+WLKE+L+EGRYED+
Sbjct: 607 ESPPPPWALAFDSWLKESLVEGRYEDI 687
>BE239650 similar to PIR|D87615|D876 conserved hypothetical protein CC2958
[imported] - Caulobacter crescentus, partial (10%)
Length = 628
Score = 129 bits (324), Expect = 1e-30
Identities = 74/203 (36%), Positives = 113/203 (55%), Gaps = 6/203 (2%)
Frame = +3
Query: 39 RPTSILVISGHWDTAVPTVNVVDST--NDTIYDFYGFPKPMYQLKYPAPGAPHLAKRVKE 96
+P +IL++S HW T + V N+ +D+ GFPK MY+L+Y APG+P LAK+V
Sbjct: 45 KPKAILIVSAHWITNNKILKVTSKKQHNELYFDYGGFPKEMYELQYNAPGSPELAKKVIG 224
Query: 97 LLKEGGFSRVDEDKKRGLDHGAWVPLLLMYPEADIPVCQLSVQSNLDGTHHYNIGKALAP 156
LL + G + +ED KR DH + PL+ ++P DIP+ +++ N H +GKAL
Sbjct: 225 LLSDAGI-KAEEDTKRNFDH*VFGPLMEIWPSTDIPIVEINFD*NY*SKFHIPVGKALPS 401
Query: 157 LKDEGVLIVGSGSAVHN----LRALERHATVAAPWAVEFDNWLKEALLEGRYEDVNHYEQ 212
++EG+ I+GSG H+ + + AT P +E DN +E R + + +E
Sbjct: 402 FREEGIFIIGSGEITHHFSGGITGAKTFATAINP-VIENDN------IEERNKGLIEWE- 557
Query: 213 KAPHAKKAHPWPDHFYPLHVAIG 235
K A+++H DH PLHVA G
Sbjct: 558 KLNGARESHAQEDHLIPLHVAAG 626
>TC87530
Length = 699
Score = 118 bits (296), Expect = 2e-27
Identities = 58/80 (72%), Positives = 71/80 (88%)
Frame = +3
Query: 1 MALKDTFYISHGSPTLSIDESLVARKFLQSWKKEVFPPRPTSILVISGHWDTAVPTVNVV 60
MALK+TFYISHGSPTL+IDE++ A KFL SW KEVFP RP++ILVISGHWDT+VPTVNVV
Sbjct: 48 MALKETFYISHGSPTLAIDETIPAWKFLTSW-KEVFPERPSAILVISGHWDTSVPTVNVV 224
Query: 61 DSTNDTIYDFYGFPKPMYQL 80
+ N+TI+DF GFP+ MY++
Sbjct: 225 NH-NETIHDFGGFPRSMYKV 281
>BG454377 similar to PIR|A85055|A85 probable leucyl tRNA synthetase
[imported] - Arabidopsis thaliana, partial (11%)
Length = 670
Score = 27.7 bits (60), Expect = 4.8
Identities = 15/35 (42%), Positives = 19/35 (53%)
Frame = -1
Query: 223 WPDHFYPLHVAIGAAGENSKAKLIHSSIDLGSLSY 257
WP + GA G+NSK K S+ DLG LS+
Sbjct: 181 WPRRDHSSSAMCGAYGDNSKTKA--STTDLGCLSH 83
>TC88030 similar to PIR|F86416|F86416 probable RNA-binding protein MEI2
36123-32976 [imported] - Arabidopsis thaliana, partial
(23%)
Length = 1549
Score = 26.9 bits (58), Expect = 8.2
Identities = 12/30 (40%), Positives = 17/30 (56%)
Frame = +3
Query: 49 HWDTAVPTVNVVDSTNDTIYDFYGFPKPMY 78
HW TVNV DS T+++ YG + +Y
Sbjct: 900 HWLFETSTVNVEDSELRTLFEQYGDIRTLY 989
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.317 0.136 0.418
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 8,906,249
Number of Sequences: 36976
Number of extensions: 131041
Number of successful extensions: 663
Number of sequences better than 10.0: 10
Number of HSP's better than 10.0 without gapping: 618
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 644
length of query: 267
length of database: 9,014,727
effective HSP length: 94
effective length of query: 173
effective length of database: 5,538,983
effective search space: 958244059
effective search space used: 958244059
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 57 (26.6 bits)
Lotus: description of TM0074.3


