

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0074.12
(369 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
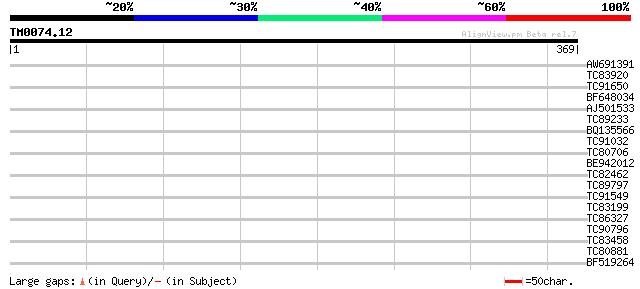
Score E
Sequences producing significant alignments: (bits) Value
AW691391 similar to GP|16604643|gb unknown protein {Arabidopsis ... 32 0.30
TC83920 similar to PIR|D86429|D86429 hypothetical protein AAF197... 32 0.39
TC91650 similar to GP|8809635|dbj|BAA97186.1 gb|AAD21751.1~gene_... 31 0.66
BF648034 similar to GP|16604643|gb unknown protein {Arabidopsis ... 31 0.87
AJ501533 30 1.1
TC89233 homologue to GP|18734|emb|CAA36734.1|| DNA-directed RNA ... 30 1.5
BQ135566 30 1.5
TC91032 weakly similar to GP|20196995|gb|AAB91977.2 expressed pr... 29 2.5
TC80706 similar to GP|13183101|gb|AAK15052.1 mutant PhcA {Ralsto... 29 2.5
BE942012 weakly similar to GP|15408731|dbj hypothetical protein~... 28 4.3
TC82462 similar to GP|9759387|dbj|BAB10038.1 dihydropyrimidinase... 28 5.6
TC89797 GP|13775543|gb|AAK39351.1 Hypothetical protein Y73B3A.17... 28 7.3
TC91549 similar to GP|17063189|gb|AAL32989.1 unknown protein {Ar... 28 7.3
TC83199 similar to GP|20466774|gb|AAM20704.1 putative protein {A... 28 7.3
TC86327 similar to GP|14335106|gb|AAK59832.1 At2g41900/T6D20.20 ... 27 9.6
TC90796 similar to GP|3395431|gb|AAC28763.1| unknown protein {Ar... 27 9.6
TC83458 homologue to PIR|T07796|T07796 DNA-directed RNA polymera... 27 9.6
TC80881 similar to PIR|T11622|T11622 extensin class 1 precursor ... 27 9.6
BF519264 similar to GP|15293221|gb| unknown protein {Arabidopsis... 27 9.6
>AW691391 similar to GP|16604643|gb unknown protein {Arabidopsis thaliana},
partial (30%)
Length = 646
Score = 32.3 bits (72), Expect = 0.30
Identities = 22/72 (30%), Positives = 38/72 (52%), Gaps = 7/72 (9%)
Frame = -3
Query: 196 ESHESSTSEDSSLITTPVDSTPYATRSDAITPDVNKT-------FVLKTTDLVLRNPRQV 248
E H S T+ SS++ PVDS ++RS +++ D N+T +T+ VL +P
Sbjct: 617 EKHLSDTT--SSILENPVDSNSSSSRSGSVSRDANRTQKGLSSENTHCSTEKVLESPPSH 444
Query: 249 FGNLLEGKFVQK 260
G++ G V++
Sbjct: 443 VGDIGRGNSVRE 408
>TC83920 similar to PIR|D86429|D86429 hypothetical protein AAF19748.1
[imported] - Arabidopsis thaliana, partial (13%)
Length = 845
Score = 32.0 bits (71), Expect = 0.39
Identities = 11/23 (47%), Positives = 15/23 (64%)
Frame = -3
Query: 25 PPSQPNTSPYYCHNVPSYYPQTH 47
P + P+ PY+CH + S YPQ H
Sbjct: 111 PGNHPHHCPYFCHAMLSVYPQAH 43
>TC91650 similar to GP|8809635|dbj|BAA97186.1
gb|AAD21751.1~gene_id:MMI9.5~similar to unknown protein
{Arabidopsis thaliana}, partial (18%)
Length = 754
Score = 31.2 bits (69), Expect = 0.66
Identities = 14/43 (32%), Positives = 21/43 (48%)
Frame = +3
Query: 24 SPPSQPNTSPYYCHNVPSYYPQTHSSLQLSEFNGENPEYWLFM 66
SPP P + P+ + + T SL FN +NP W+F+
Sbjct: 81 SPPHSPESQPFKLSTTKTSFKTTFQSL----FNLQNPRAWIFL 197
>BF648034 similar to GP|16604643|gb unknown protein {Arabidopsis thaliana},
partial (28%)
Length = 652
Score = 30.8 bits (68), Expect = 0.87
Identities = 21/72 (29%), Positives = 38/72 (52%), Gaps = 7/72 (9%)
Frame = -3
Query: 196 ESHESSTSEDSSLITTPVDSTPYATRSDAITPDVNKT-------FVLKTTDLVLRNPRQV 248
+ H S T+ SS++ PVDS ++RS +++ D N+T +T+ VL +P
Sbjct: 638 KKHLSDTT--SSILENPVDSNSSSSRSGSVSRDANRTQKGLSSENTHCSTEKVLESPPSH 465
Query: 249 FGNLLEGKFVQK 260
G++ G V++
Sbjct: 464 VGDIGRGNSVRE 429
>AJ501533
Length = 671
Score = 30.4 bits (67), Expect = 1.1
Identities = 15/36 (41%), Positives = 21/36 (57%), Gaps = 5/36 (13%)
Frame = +2
Query: 25 PPSQPNTSP-YYCHNVPS----YYPQTHSSLQLSEF 55
PP P+ SP YY PS YYPQ+H +++L +
Sbjct: 452 PPHVPSPSPMYYSGGCPSFFGPYYPQSHYNMELPRY 559
>TC89233 homologue to GP|18734|emb|CAA36734.1|| DNA-directed RNA polymerase
{Glycine max}, partial (21%)
Length = 648
Score = 30.0 bits (66), Expect = 1.5
Identities = 18/43 (41%), Positives = 23/43 (52%), Gaps = 3/43 (6%)
Frame = +3
Query: 23 YSP--PSQPNTSPYYCHNVPSYYPQTHSSLQLSEFN-GENPEY 62
YSP PS TSP Y PSY P + + S +N G +P+Y
Sbjct: 75 YSPTSPSYSPTSPSYSPTSPSYSPSSPTYSPSSPYNSGTSPDY 203
>BQ135566
Length = 930
Score = 30.0 bits (66), Expect = 1.5
Identities = 15/35 (42%), Positives = 21/35 (59%), Gaps = 3/35 (8%)
Frame = +2
Query: 18 PRSIF---YSPPSQPNTSPYYCHNVPSYYPQTHSS 49
P S+F Y P QP+ PY+ +P Y+P +HSS
Sbjct: 395 PHSLFPFCYPSPFQPHLLPYH---IPFYHPLSHSS 490
>TC91032 weakly similar to GP|20196995|gb|AAB91977.2 expressed protein
{Arabidopsis thaliana}, partial (37%)
Length = 1366
Score = 29.3 bits (64), Expect = 2.5
Identities = 14/35 (40%), Positives = 21/35 (60%), Gaps = 5/35 (14%)
Frame = -2
Query: 22 FYSPPSQPNTSPY-----YCHNVPSYYPQTHSSLQ 51
F +PPS P+T P + +PS+ P +HS+LQ
Sbjct: 516 FSTPPSTPSTLPQQPSTSHSSTLPSHSPTSHSTLQ 412
>TC80706 similar to GP|13183101|gb|AAK15052.1 mutant PhcA {Ralstonia
solanacearum}, partial (8%)
Length = 729
Score = 29.3 bits (64), Expect = 2.5
Identities = 16/48 (33%), Positives = 22/48 (45%)
Frame = -1
Query: 29 PNTSPYYCHNVPSYYPQTHSSLQLSEFNGENPEYWLFMADCYFLQNPY 76
P TSPY+ S+Y Q H S +L F P + + Q+PY
Sbjct: 555 PGTSPYH*ARCQSFYHQQHDS-ELLHFQLAGPRHGFLSHQSF*HQHPY 415
>BE942012 weakly similar to GP|15408731|dbj hypothetical protein~similar to
receptor serine/threonine kinase PR5K, partial (7%)
Length = 561
Score = 28.5 bits (62), Expect = 4.3
Identities = 23/85 (27%), Positives = 44/85 (51%), Gaps = 3/85 (3%)
Frame = +3
Query: 134 GFEKSSLVEIQRSSEQVSPHRVVFSETTLEDGAQLPFE---ASSDVHLLPEVVSRVELGS 190
G + ++VE RSSE PH V++ + LE G L + S++ + + + ++ V L
Sbjct: 27 GRKNVNIVEASRSSELYFPHLVIYKK--LEKGNDLELDDGVMSNEENEIAKKLTMVGLWC 200
Query: 191 AETEIESHESSTSEDSSLITTPVDS 215
+T I +H + S+ ++ +DS
Sbjct: 201 IQT-IPTHRPTISKVIDMLDGSMDS 272
>TC82462 similar to GP|9759387|dbj|BAB10038.1 dihydropyrimidinase
{Arabidopsis thaliana}, partial (28%)
Length = 693
Score = 28.1 bits (61), Expect = 5.6
Identities = 12/34 (35%), Positives = 21/34 (61%)
Frame = -2
Query: 257 FVQKNNEAQFEKHFDKLLMQFHNSSTTVDVQKIG 290
++ NN+A F+ + LL+ ++STT D Q +G
Sbjct: 308 WINSNNDAVFDINISNLLVMSIDNSTTFDQQLVG 207
>TC89797 GP|13775543|gb|AAK39351.1 Hypothetical protein Y73B3A.17
{Caenorhabditis elegans}, partial (1%)
Length = 958
Score = 27.7 bits (60), Expect = 7.3
Identities = 13/40 (32%), Positives = 19/40 (47%)
Frame = +3
Query: 21 IFYSPPSQPNTSPYYCHNVPSYYPQTHSSLQLSEFNGENP 60
I+Y PPS P SP P ++ +H+S L + P
Sbjct: 207 IYYFPPSFPFPSPEQPQ*FPPFHESSHASQPLHSASSSKP 326
>TC91549 similar to GP|17063189|gb|AAL32989.1 unknown protein {Arabidopsis
thaliana}, partial (31%)
Length = 1499
Score = 27.7 bits (60), Expect = 7.3
Identities = 20/42 (47%), Positives = 24/42 (56%), Gaps = 3/42 (7%)
Frame = +3
Query: 177 HLLPEVVSRVE-LGSAET--EIESHESSTSEDSSLITTPVDS 215
HL EV R++ L ET +I S + SEDSS TTP DS
Sbjct: 1050 HLCDEVDRRIKPLLKLETVLDISSSSAPVSEDSSSHTTPPDS 1175
>TC83199 similar to GP|20466774|gb|AAM20704.1 putative protein {Arabidopsis
thaliana}, partial (18%)
Length = 748
Score = 27.7 bits (60), Expect = 7.3
Identities = 14/43 (32%), Positives = 25/43 (57%), Gaps = 1/43 (2%)
Frame = -1
Query: 13 SAQRDPRSIFYSPPSQPNTSPYYCH-NVPSYYPQTHSSLQLSE 54
S +R P ++ S P+ T+ CH N+P +P+TH Q+++
Sbjct: 370 SRRRRP-NLRISTPNHGTTTTRVCHQNLPLKFPKTHKKTQMTQ 245
>TC86327 similar to GP|14335106|gb|AAK59832.1 At2g41900/T6D20.20 {Arabidopsis
thaliana}, partial (50%)
Length = 3018
Score = 27.3 bits (59), Expect = 9.6
Identities = 15/54 (27%), Positives = 28/54 (51%)
Frame = +1
Query: 65 FMADCYFLQNPYPWETRFQLLAIHLGGKALTWFRLSHQQIHSWEEFKVAFRLYF 118
F YF+ P+ T +++L++HL W +S++ I +E + +LYF
Sbjct: 2746 FFLSFYFISF*VPFPTLYKILSVHLCCLCRKWD*VSYEGI--YELLNIYCKLYF 2901
>TC90796 similar to GP|3395431|gb|AAC28763.1| unknown protein {Arabidopsis
thaliana}, partial (20%)
Length = 965
Score = 27.3 bits (59), Expect = 9.6
Identities = 12/35 (34%), Positives = 17/35 (48%)
Frame = +2
Query: 20 SIFYSPPSQPNTSPYYCHNVPSYYPQTHSSLQLSE 54
S Y+P + P SPY + P PQ+ S Q +
Sbjct: 773 SYSYAPQAYPGNSPYNSYVAPWAQPQSKSEFQTQQ 877
>TC83458 homologue to PIR|T07796|T07796 DNA-directed RNA polymerase (EC
2.7.7.6) largest chain - soybean (fragment), partial
(63%)
Length = 1220
Score = 27.3 bits (59), Expect = 9.6
Identities = 16/33 (48%), Positives = 17/33 (51%), Gaps = 2/33 (6%)
Frame = +2
Query: 18 PRSIFYSP--PSQPNTSPYYCHNVPSYYPQTHS 48
P S YSP PS TSP Y PSY P + S
Sbjct: 1004 PTSPSYSPTSPSYSPTSPSYSPTSPSYSPTSSS 1102
>TC80881 similar to PIR|T11622|T11622 extensin class 1 precursor - cowpea,
partial (34%)
Length = 597
Score = 27.3 bits (59), Expect = 9.6
Identities = 14/36 (38%), Positives = 17/36 (46%), Gaps = 8/36 (22%)
Frame = +2
Query: 25 PPSQPNTSPYYCHNVPSYY--------PQTHSSLQL 52
PP P+ P Y H P YY P T SS++L
Sbjct: 245 PPPSPSPPPPYVHKYPPYYYKSPPPPSPFTTSSIRL 352
>BF519264 similar to GP|15293221|gb| unknown protein {Arabidopsis thaliana},
partial (7%)
Length = 416
Score = 27.3 bits (59), Expect = 9.6
Identities = 15/32 (46%), Positives = 17/32 (52%)
Frame = -3
Query: 18 PRSIFYSPPSQPNTSPYYCHNVPSYYPQTHSS 49
P IFYSP S N PS +PQTH+S
Sbjct: 123 PFEIFYSPNSS---------NDPSEFPQTHTS 55
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.318 0.134 0.395
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 12,372,509
Number of Sequences: 36976
Number of extensions: 196860
Number of successful extensions: 1117
Number of sequences better than 10.0: 38
Number of HSP's better than 10.0 without gapping: 1097
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1111
length of query: 369
length of database: 9,014,727
effective HSP length: 98
effective length of query: 271
effective length of database: 5,391,079
effective search space: 1460982409
effective search space used: 1460982409
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 59 (27.3 bits)
Lotus: description of TM0074.12


