

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0062.9
(271 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
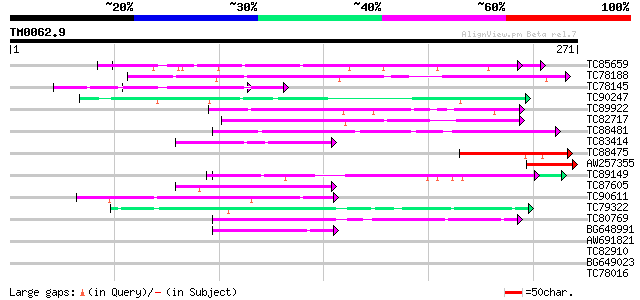
Sequences producing significant alignments: (bits) Value
TC85659 SP|O24076|GBLP_MEDSA Guanine nucleotide-binding protein ... 63 1e-10
TC78188 similar to PIR|C84870|C84870 probable splicing factor [i... 61 5e-10
TC78145 similar to GP|15294240|gb|AAK95297.1 AT3g18860/MCB22_3 {... 52 2e-07
TC90247 weakly similar to GP|21592898|gb|AAM64848.1 unknown {Ara... 51 5e-07
TC89922 weakly similar to GP|15912305|gb|AAL08286.1 AT4g18900/F1... 50 1e-06
TC82717 weakly similar to GP|21539583|gb|AAM53344.1 putative coa... 48 3e-06
TC88481 similar to PIR|T00806|T02445 probable U4/U6 small nuclea... 47 1e-05
TC83414 weakly similar to GP|17381206|gb|AAL36415.1 unknown prot... 47 1e-05
TC88475 similar to GP|20260442|gb|AAM13119.1 unknown protein {Ar... 47 1e-05
AW257355 similar to GP|22022579|gb| AT3g05090/T12H1_5 {Arabidops... 45 2e-05
TC89149 similar to GP|20466231|gb|AAM20433.1 cell cycle switch p... 45 3e-05
TC87605 homologue to PIR|G96563|G96563 probable coatomer complex... 44 7e-05
TC90611 similar to GP|10177223|dbj|BAB10298. contains similarity... 43 1e-04
TC79322 similar to GP|13605815|gb|AAK32893.1 At2g32700/F24L7.16 ... 42 2e-04
TC80769 similar to GP|11141605|gb|AAG32022.1 LEUNIG {Arabidopsis... 42 3e-04
BG648991 similar to PIR|C86239|C86 protein T10O24.21 [imported] ... 41 6e-04
AW691821 similar to GP|10177570|db contains similarity to regula... 40 0.001
TC82910 similar to GP|19347844|gb|AAL86002.1 unknown protein {Ar... 40 0.001
BG649023 similar to GP|10178169|db contains similarity to WD-con... 40 0.001
TC78016 similar to GP|21537191|gb|AAM61532.1 PRL1 protein {Arabi... 40 0.001
>TC85659 SP|O24076|GBLP_MEDSA Guanine nucleotide-binding protein beta
subunit-like protein. [Alfalfa] {Medicago sativa},
complete
Length = 1261
Score = 62.8 bits (151), Expect = 1e-10
Identities = 69/220 (31%), Positives = 96/220 (43%), Gaps = 17/220 (7%)
Frame = +2
Query: 43 VLKSAVSDGSDYLFTGSRDGRLKRW-ALGEDAATCSATFE---SHVDWVNDAVLVGDSTL 98
VL A S + + + SRD +K W LGE C T + +H DWV+ V STL
Sbjct: 383 VLSVAFSIDNRQIVSASRDRTIKLWNTLGE----CKYTIQDGDAHSDWVS-CVRFSPSTL 547
Query: 99 ----VSCSSDTTLKTWDAFSTGTCTRTLRQHSDYVTCLAASEKNSNIVASGGLGGEVFVW 154
VS S D T+K W+ + TL HS YV +A S S + ASGG G + +W
Sbjct: 548 QPTIVSASWDRTVKVWN-LTNCKLRNTLAGHSGYVNTVAVSPDGS-LCASGGKDGVILLW 721
Query: 155 DLEAAHA----SAAKCIDATDDDTSNG--INGSGNSLPLTSLRSISSSNSISVHT-TQNQ 207
DL A I A + + +S+ + L S S + V T+
Sbjct: 722 DLAEGKRLYSLDAGSIIHALCFSPNRYWLCAATESSIKIWDLESKSIVEDLKVDLKTEAD 901
Query: 208 GYIPIAAKGHKESVYALAMN--EGGTLLVSGGTEKVVRVW 245
I K+ +Y ++N G+ L SG T+ VVRVW
Sbjct: 902 AAIGGDTTTKKKVIYCTSLNWSADGSTLFSGYTDGVVRVW 1021
Score = 55.5 bits (132), Expect = 2e-08
Identities = 60/211 (28%), Positives = 88/211 (41%), Gaps = 4/211 (1%)
Frame = +2
Query: 50 DGSDYLFTGSRDGRLKRWALGEDAATCSAT---FESHVDWVNDAVLVGDSTL-VSCSSDT 105
D SD + T SRD + W L ++ T H +V D VL D +S S D
Sbjct: 137 DNSDMIVTASRDKSIILWHLTKEDKTYGVPRRRLTGHSHFVQDVVLSSDGQFALSGSWDG 316
Query: 106 TLKTWDAFSTGTCTRTLRQHSDYVTCLAASEKNSNIVASGGLGGEVFVWDLEAAHASAAK 165
L+ WD + GT R H+ V +A S N IV S + +W+ +
Sbjct: 317 ELRLWD-LNAGTSARRFVGHTKDVLSVAFSIDNRQIV-SASRDRTIKLWNTLGECKYTIQ 490
Query: 166 CIDATDDDTSNGINGSGNSLPLTSLRSISSSNSISVHTTQNQGYIPIAAKGHKESVYALA 225
DA D S + S ++L T + S S ++ V N A GH V +A
Sbjct: 491 DGDAHSDWVSC-VRFSPSTLQPTIV-SASWDRTVKVWNLTNCKLRNTLA-GHSGYVNTVA 661
Query: 226 MNEGGTLLVSGGTEKVVRVWDPRSGSKTMKL 256
++ G+L SGG + V+ +WD G + L
Sbjct: 662 VSPDGSLCASGGKDGVILLWDLAEGKRLYSL 754
Score = 39.3 bits (90), Expect = 0.002
Identities = 33/165 (20%), Positives = 62/165 (37%), Gaps = 1/165 (0%)
Frame = +2
Query: 100 SCSSDTTLKTWDAFSTGTCTR-TLRQHSDYVTCLAASEKNSNIVASGGLGGEVFVWDLEA 158
SC+ ++ + + G R T+R H+D VT +A NS+++ + + +W L
Sbjct: 23 SCNCNSIPQKPPIMAEGLVLRGTMRAHTDVVTAIATPIDNSDMIVTASRDKSIILWHL-- 196
Query: 159 AHASAAKCIDATDDDTSNGINGSGNSLPLTSLRSISSSNSISVHTTQNQGYIPIAAKGHK 218
T +D + G+ P L GH
Sbjct: 197 -----------TKEDKTYGV-------PRRRLT------------------------GHS 250
Query: 219 ESVYALAMNEGGTLLVSGGTEKVVRVWDPRSGSKTMKLKGHTDNI 263
V + ++ G +SG + +R+WD +G+ + GHT ++
Sbjct: 251 HFVQDVVLSSDGQFALSGSWDGELRLWDLNAGTSARRFVGHTKDV 385
>TC78188 similar to PIR|C84870|C84870 probable splicing factor [imported] -
Arabidopsis thaliana, partial (95%)
Length = 1436
Score = 60.8 bits (146), Expect = 5e-10
Identities = 52/227 (22%), Positives = 95/227 (40%), Gaps = 15/227 (6%)
Frame = +1
Query: 57 TGSRDGRLKRWALGEDAATCSATFESHVDWVNDAVLVGDST-LVSCSSDTTLKTWDAFST 115
+GS D + W + D + H + V D D T ++S S D TL+ WD T
Sbjct: 364 SGSHDKEIFLWNVHGDCKNFMV-LKGHKNAVLDLHWTSDGTQIISASPDKTLRLWDT-ET 537
Query: 116 GTCTRTLRQHSDYVTCLAASEKNSNIVASGGLGGEVFVWDL----------EAAHASAAK 165
G + + +H YV + + +V SG G +WD+ + +A
Sbjct: 538 GKQIKKMVEHLSYVNSCCPTRRGPPLVVSGSDDGTAKLWDMRQRGSIQTFPDKYQITAVS 717
Query: 166 CIDATDDDTSNGINGSGNSLPLTSLRSISSSNSISVHTTQNQGYIPIAAKGHKESVYALA 225
DA+D + GI+ N + + LR +G + + +GH++ + ++
Sbjct: 718 FSDASDKIYTGGID---NDVKIWDLR---------------KGEVTMTLQGHQDMITSMQ 843
Query: 226 MNEGGTLLVSGGTEKVVRVWDPRSGSKTMK----LKGHTDNIRALLL 268
++ G+ L++ G + + +WD R + + L+GH N LL
Sbjct: 844 LSPDGSYLLTNGMDCKLCIWDMRPYAPQNRCVKILEGHQHNFEKNLL 984
Score = 36.2 bits (82), Expect = 0.014
Identities = 16/49 (32%), Positives = 28/49 (56%), Gaps = 1/49 (2%)
Frame = +1
Query: 216 GHKESVYALAMNEGGTLLVSGGTEKVVRVWDPRSGSKT-MKLKGHTDNI 263
GH+ +VY + N G+++ SG +K + +W+ K M LKGH + +
Sbjct: 307 GHQSAVYTMKFNPTGSVVASGSHDKEIFLWNVHGDCKNFMVLKGHKNAV 453
>TC78145 similar to GP|15294240|gb|AAK95297.1 AT3g18860/MCB22_3 {Arabidopsis
thaliana}, partial (93%)
Length = 2633
Score = 52.0 bits (123), Expect = 2e-07
Identities = 39/112 (34%), Positives = 54/112 (47%)
Frame = +3
Query: 22 LTYVLNDSDDTKHCAGINCLAVLKSAVSDGSDYLFTGSRDGRLKRWALGEDAATCSATFE 81
L + LN + + G + L V V DG L + S D L+RW G+ C T+E
Sbjct: 366 LVWDLNSGEKVQSLKG-HQLQVTGIVVDDGD--LVSSSMDCTLRRWRNGQ----CIETWE 524
Query: 82 SHVDWVNDAVLVGDSTLVSCSSDTTLKTWDAFSTGTCTRTLRQHSDYVTCLA 133
+H + + + LV+ SSDTTLKTW TC T + H+D V LA
Sbjct: 525 AHKGPIQAVIKLPTGELVTGSSDTTLKTWKG---KTCLHTFQGHADTVRGLA 671
Score = 41.2 bits (95), Expect = 4e-04
Identities = 22/62 (35%), Positives = 29/62 (46%)
Frame = +3
Query: 55 LFTGSRDGRLKRWALGEDAATCSATFESHVDWVNDAVLVGDSTLVSCSSDTTLKTWDAFS 114
L TGS D LK W TC TF+ H D V ++ D ++S S D +L+ W
Sbjct: 573 LVTGSSDTTLKTWK----GKTCLHTFQGHADTVRGLAVMSDLGILSASHDGSLRLWAVSG 740
Query: 115 TG 116
G
Sbjct: 741 EG 746
Score = 38.5 bits (88), Expect = 0.003
Identities = 33/107 (30%), Positives = 47/107 (43%), Gaps = 8/107 (7%)
Frame = +3
Query: 171 DDDTSNGINGSGNSLPLTSLRSISSSNSISVHTTQNQGYI-PIAAKGHKESVYALA---- 225
DD I G N TS R +I + T N+ ++ GH V +A
Sbjct: 144 DDVRGITICGGDNGGIATSSRD----KTIRLWTRDNRKFVLDKVLVGHTSFVGPIAWIPP 311
Query: 226 ---MNEGGTLLVSGGTEKVVRVWDPRSGSKTMKLKGHTDNIRALLLD 269
+ GG +VSGG + +V VWD SG K LKGH + +++D
Sbjct: 312 NPDLPHGG--VVSGGMDTLVLVWDLNSGEKVQSLKGHQLQVTGIVVD 446
Score = 37.4 bits (85), Expect = 0.006
Identities = 47/202 (23%), Positives = 81/202 (39%), Gaps = 14/202 (6%)
Frame = +3
Query: 83 HVDWVNDAVLVGDST--LVSCSSDTTLKTWDAFSTG-TCTRTLRQHSDYVTCLAASEKNS 139
H D V + G + + S D T++ W + + L H+ +V +A N
Sbjct: 138 HEDDVRGITICGGDNGGIATSSRDKTIRLWTRDNRKFVLDKVLVGHTSFVGPIAWIPPNP 317
Query: 140 NI----VASGGLGGEVFVWDLEAAHASAAKCIDATDDDTSNGINGSGNSLPLTSL----- 190
++ V SGG+ V VWDL + + G+ L +T +
Sbjct: 318 DLPHGGVVSGGMDTLVLVWDLNSGEKVQSL---------------KGHQLQVTGIVVDDG 452
Query: 191 RSISSSNSISVHTTQNQGYIPI--AAKGHKESVYALAMNEGGTLLVSGGTEKVVRVWDPR 248
+SSS ++ +N I A KG ++V L E LV+G ++ ++ W +
Sbjct: 453 DLVSSSMDCTLRRWRNGQCIETWEAHKGPIQAVIKLPTGE----LVTGSSDTTLKTWKGK 620
Query: 249 SGSKTMKLKGHTDNIRALLLDS 270
+ T +GH D +R L + S
Sbjct: 621 TCLHT--FQGHADTVRGLAVMS 680
>TC90247 weakly similar to GP|21592898|gb|AAM64848.1 unknown {Arabidopsis
thaliana}, partial (60%)
Length = 1041
Score = 50.8 bits (120), Expect = 5e-07
Identities = 53/229 (23%), Positives = 88/229 (38%), Gaps = 13/229 (5%)
Frame = +1
Query: 34 HCAGINCLAVLKSAVSDGSDYLFTGSRDGRLKRWAL---------GEDAATCSATFESHV 84
H + CL SD L +GS DG ++ W+L E+ +F H
Sbjct: 439 HLRAVTCLVF-----SDDDSLLISGSEDGYVRVWSLFSLFDDLRSREEKNLYEYSFSEHT 603
Query: 85 DWVNDAVLVG---DSTLVSCSSDTTLKTWDAFSTGTCTRTLRQHSDYVTCLAASEKNSNI 141
V D V+ ++ +VS S D T K W + S GT R++ S V + ++
Sbjct: 604 MRVTDVVIGNGGCNAIIVSASDDRTCKVW-SLSRGTLLRSIVFPS--VINAIVLDPAEHV 774
Query: 142 VASGGLGGEVFVWDLEAAHASAAKCIDATDDDTSNGINGSGNSLPLTSLRSISSSNSISV 201
+G G++F I++ N+ +
Sbjct: 775 FYAGSSDGKIF----------------------------------------IAALNTERI 834
Query: 202 HTTQNQGYIPIAA-KGHKESVYALAMNEGGTLLVSGGTEKVVRVWDPRS 249
T N G I + H ++V LA ++G LL+SG + +VRVW+ ++
Sbjct: 835 TTIDNYGMHIIGSFSNHSKAVTCLAYSKGRNLLISGSEDGIVRVWNAKT 981
>TC89922 weakly similar to GP|15912305|gb|AAL08286.1 AT4g18900/F13C5_70
{Arabidopsis thaliana}, partial (48%)
Length = 864
Score = 49.7 bits (117), Expect = 1e-06
Identities = 43/161 (26%), Positives = 76/161 (46%), Gaps = 10/161 (6%)
Frame = +1
Query: 96 STLVSCSSDTTLKTWDAFSTGTCTRTLRQHSDYVTCLAASEKNSNIVASGGLGGEVFVWD 155
+TL S S+D +K WD + G CT T+ HSD V +A + + I+ SG V + D
Sbjct: 133 NTLASASADKRVKIWDIVA-GKCTITMDHHSDKVQAVAWNHRAQQILLSGSFDHTVALKD 309
Query: 156 LEA-AHASAAKCIDATDD--------DTSNGINGSGNSLPLTSLRSISSSNSISVHTTQN 206
+ +H+ + A + + S ++ ++ +R+ + SN+ SV QN
Sbjct: 310 VRTPSHSGYTWSVSADVESLAWDPHTEHSFAVSLEDGTIQCFDVRT-AMSNATSV---QN 477
Query: 207 QGYIPIAAKGHKESVYALAMNEGG-TLLVSGGTEKVVRVWD 246
+ H +SV +++ N LL +G +K V++WD
Sbjct: 478 ATF---TLHAHDKSVTSVSYNTAAPNLLATGSMDKTVKLWD 591
Score = 36.2 bits (82), Expect = 0.014
Identities = 18/51 (35%), Positives = 29/51 (56%), Gaps = 1/51 (1%)
Frame = +1
Query: 217 HKESVYALAMN-EGGTLLVSGGTEKVVRVWDPRSGSKTMKLKGHTDNIRAL 266
H +SV LA N E L S +K V++WD +G T+ + H+D ++A+
Sbjct: 88 HTDSVLGLAWNKEYSNTLASASADKRVKIWDIVAGKCTITMDHHSDKVQAV 240
Score = 30.0 bits (66), Expect = 0.98
Identities = 29/116 (25%), Positives = 49/116 (42%), Gaps = 11/116 (9%)
Frame = +1
Query: 125 HSDYVTCLAASEKNSNIVASGGLGGEVFVWDLEAAHASAAKCIDATDDDTSNGINGSGNS 184
H+D V LA +++ SN +AS V +WD+ A KC D + + N
Sbjct: 88 HTDSVLGLAWNKEYSNTLASASADKRVKIWDI-----VAGKCTITMDHHSDKVQAVAWNH 252
Query: 185 LPLTSLRSISSSNSIS---VHTTQNQGYI--------PIAAKGHKESVYALAMNEG 229
L S S ++++ V T + GY +A H E +A+++ +G
Sbjct: 253 RAQQILLSGSFDHTVALKDVRTPSHSGYTWSVSADVESLAWDPHTEHSFAVSLEDG 420
>TC82717 weakly similar to GP|21539583|gb|AAM53344.1 putative coatomer
complex subunit {Arabidopsis thaliana}, partial (29%)
Length = 1118
Score = 48.1 bits (113), Expect = 3e-06
Identities = 32/153 (20%), Positives = 63/153 (40%), Gaps = 8/153 (5%)
Frame = +1
Query: 102 SSDTTLKTWDAFSTGTCTRTLRQHSDYVTCLAASEKNSNIVASGGLGGEVFVWDLEAA-- 159
S D LK W+ +C T +S YV +A + K+ + AS L G + +W ++++
Sbjct: 442 SDDQVLKLWNWKKGWSCDETFEGNSHYVMQVAFNPKDPSTFASASLDGTLKIWTIDSSAP 621
Query: 160 ------HASAAKCIDATDDDTSNGINGSGNSLPLTSLRSISSSNSISVHTTQNQGYIPIA 213
H C+D + + + SG+ + S N +
Sbjct: 622 NFTFEGHLKGMNCVDYFESNDKQYLL-SGSDDYTAKVWDYDSKNCVQT------------ 762
Query: 214 AKGHKESVYALAMNEGGTLLVSGGTEKVVRVWD 246
+GHK +V A+ + ++++ + V++WD
Sbjct: 763 LEGHKNNVTAICAHPEIPIIITASEDSTVKIWD 861
>TC88481 similar to PIR|T00806|T02445 probable U4/U6 small nuclear
ribonucleoprotein [imported] - Arabidopsis thaliana,
partial (60%)
Length = 1836
Score = 46.6 bits (109), Expect = 1e-05
Identities = 42/166 (25%), Positives = 67/166 (40%)
Frame = +1
Query: 98 LVSCSSDTTLKTWDAFSTGTCTRTLRQHSDYVTCLAASEKNSNIVASGGLGGEVFVWDLE 157
L +CS K W + + TL+ H+ T +A S + N +A+ W+ +
Sbjct: 994 LATCSFTGATKLWSMPNVKKVS-TLKGHTQRATDVAYSPVHKNHLATASADRTAKYWNDQ 1170
Query: 158 AAHASAAKCIDATDDDTSNGINGSGNSLPLTSLRSISSSNSISVHTTQNQGYIPIAAKGH 217
A K + + SG L S + V T + + +GH
Sbjct: 1171 GALLGTFK--GHLERLARIAFHPSGKYLGTASYDK--TWRLWDVETEEEL----LLQEGH 1326
Query: 218 KESVYALAMNEGGTLLVSGGTEKVVRVWDPRSGSKTMKLKGHTDNI 263
SVY L + G+L S G + + RVWD R+G + L+GH +I
Sbjct: 1327 SRSVYGLDFHHDGSLAASCGLDALARVWDLRTGRSVLALEGHVKSI 1464
Score = 40.0 bits (92), Expect = 0.001
Identities = 45/222 (20%), Positives = 83/222 (37%), Gaps = 13/222 (5%)
Frame = +1
Query: 55 LFTGSRDGRLKRWALGEDAATCSATFESHVDWVNDAVL--VGDSTLVSCSSDTTLKTWDA 112
L T S G K W++ +T + H D V + L + S+D T K W+
Sbjct: 994 LATCSFTGATKLWSMPNVKKV--STLKGHTQRATDVAYSPVHKNHLATASADRTAKYWN- 1164
Query: 113 FSTGTCTRTLRQHSDYVTCLAASEKNSNIVASGGLGGEVFVWDLEA--------AHASAA 164
G T + H + + +A + + + +WD+E H+ +
Sbjct: 1165 -DQGALLGTFKGHLERLARIAF-HPSGKYLGTASYDKTWRLWDVETEEELLLQEGHSRSV 1338
Query: 165 KCIDATDDDT---SNGINGSGNSLPLTSLRSISSSNSISVHTTQNQGYIPIAAKGHKESV 221
+D D + S G++ L + RS+ +A +GH +S+
Sbjct: 1339 YGLDFHHDGSLAASCGLDALARVWDLRTGRSV------------------LALEGHVKSI 1464
Query: 222 YALAMNEGGTLLVSGGTEKVVRVWDPRSGSKTMKLKGHTDNI 263
++ + G L +GG + R+WD R + H++ I
Sbjct: 1465 LGISFSPNGYHLATGGEDNTCRIWDLRKKKSLYTIPAHSNLI 1590
Score = 34.7 bits (78), Expect = 0.040
Identities = 51/223 (22%), Positives = 76/223 (33%), Gaps = 9/223 (4%)
Frame = +1
Query: 53 DYLFTGSRDGRLKRWALGEDAATCSATFESHVDWV-NDAVLVGDSTLVSCSSDTTLKTWD 111
++L T S D K W D TF+ H++ + A L + S D T + WD
Sbjct: 1117 NHLATASADRTAKYW---NDQGALLGTFKGHLERLARIAFHPSGKYLGTASYDKTWRLWD 1287
Query: 112 AFSTGTCTRTLRQHSDYVTCLAASEKNSNIVASGGLGGEVFVWDLEAA--------HASA 163
T HS V L S + AS GL VWDL H +
Sbjct: 1288 V-ETEEELLLQEGHSRSVYGLDFHHDGS-LAASCGLDALARVWDLRTGRSVLALEGHVKS 1461
Query: 164 AKCIDATDDDTSNGINGSGNSLPLTSLRSISSSNSISVHTTQNQGYIPIAAKGHKESVYA 223
I + + G N+ + LR S +I H+ + ++ E
Sbjct: 1462 ILGISFSPNGYHLATGGEDNTCRIWDLRKKKSLYTIPAHSN-------LISQVKFEP--- 1611
Query: 224 LAMNEGGTLLVSGGTEKVVRVWDPRSGSKTMKLKGHTDNIRAL 266
+ G LV+ + +VW R L GH + +L
Sbjct: 1612 ----QEGYFLVTASYDMTAKVWSSRDFKPVKTLSGHEAKVTSL 1728
>TC83414 weakly similar to GP|17381206|gb|AAL36415.1 unknown protein
{Arabidopsis thaliana}, partial (28%)
Length = 1088
Score = 46.6 bits (109), Expect = 1e-05
Identities = 24/77 (31%), Positives = 38/77 (49%)
Frame = +2
Query: 80 FESHVDWVNDAVLVGDSTLVSCSSDTTLKTWDAFSTGTCTRTLRQHSDYVTCLAASEKNS 139
F+ H D + D L+S S D T++ W C R H++YVTC+ + N
Sbjct: 284 FQGHSDDILDLAWSKSGFLLSSSVDKTVRLWQV-GISKCLRVF-SHNNYVTCVNFNPVND 457
Query: 140 NIVASGGLGGEVFVWDL 156
N SG + G+V +W++
Sbjct: 458 NFFISGSIDGKVRIWEV 508
>TC88475 similar to GP|20260442|gb|AAM13119.1 unknown protein {Arabidopsis
thaliana}, partial (96%)
Length = 1315
Score = 46.6 bits (109), Expect = 1e-05
Identities = 24/58 (41%), Positives = 36/58 (61%), Gaps = 4/58 (6%)
Frame = +3
Query: 216 GHKESVYALAMNEGGTLLVSGGTEKVVRVW--DPRSGSKT--MKLKGHTDNIRALLLD 269
GHK+ V+++A N GT L SG ++ R+W DP + K ++LKGHTD++ L D
Sbjct: 195 GHKKKVHSVAWNCIGTKLASGSVDQTARIWHIDPHAHGKVKDIELKGHTDSVDQLCWD 368
Score = 33.5 bits (75), Expect = 0.089
Identities = 21/55 (38%), Positives = 30/55 (54%), Gaps = 3/55 (5%)
Frame = +3
Query: 212 IAAKGHKESVYALAMN-EGGTLLVSGGTEKVVRVWDPRSG--SKTMKLKGHTDNI 263
I KGH +SV L + + L+ + +K VR+WD RSG S+ +L G NI
Sbjct: 321 IELKGHTDSVDQLCWDPKHPDLIATASGDKTVRLWDARSGKCSQQAELSGENINI 485
>AW257355 similar to GP|22022579|gb| AT3g05090/T12H1_5 {Arabidopsis
thaliana}, partial (11%)
Length = 264
Score = 45.4 bits (106), Expect = 2e-05
Identities = 21/24 (87%), Positives = 22/24 (91%)
Frame = +1
Query: 248 RSGSKTMKLKGHTDNIRALLLDST 271
RSGSK +KLKGHTDN RALLLDST
Sbjct: 4 RSGSKILKLKGHTDNFRALLLDST 75
Score = 33.9 bits (76), Expect = 0.068
Identities = 16/52 (30%), Positives = 27/52 (51%)
Frame = +1
Query: 215 KGHKESVYALAMNEGGTLLVSGGTEKVVRVWDPRSGSKTMKLKGHTDNIRAL 266
KGH ++ AL ++ G +SG + ++R+WD HTD++ AL
Sbjct: 31 KGHTDNFRALLLDSTGRYCLSGSSYSMIRLWDIGQQRCVHSYAVHTDSVWAL 186
>TC89149 similar to GP|20466231|gb|AAM20433.1 cell cycle switch protein
{Arabidopsis thaliana}, partial (71%)
Length = 1283
Score = 45.1 bits (105), Expect = 3e-05
Identities = 45/169 (26%), Positives = 70/169 (40%), Gaps = 10/169 (5%)
Frame = +2
Query: 95 DSTLVSCSSDTTLKTWDAFSTGTCTRTLRQHSDYVTCLAASEKNSNIVASGGLGGEVFVW 154
D L S +D L W+ S R L +H+ V +A S SN++ SGG
Sbjct: 581 DRELASGGNDNQLLVWNQHSQQPTLR-LTEHTAAVKAIAWSPHQSNLLVSGG-------- 733
Query: 155 DLEAAHASAAKCIDATDDDTSNGINGSGNSLPLTSLRSISSSNS-ISVHT-TQNQGYI-- 210
+A +CI + + +N + +L + N +S H +QNQ +
Sbjct: 734 ------GTADRCIRFWNTTNGHQLNSVDTGSQVCNLAWSKNVNELVSTHGYSQNQIMVWK 895
Query: 211 -PIAAK-----GHKESVYALAMNEGGTLLVSGGTEKVVRVWDPRSGSKT 253
P AK GH V LAM+ G +V+G ++ +R W+ KT
Sbjct: 896 YPSLAKVATLTGHSMRVLYLAMSPDGQTIVTGAGDETLRFWNVFPSMKT 1042
Score = 42.7 bits (99), Expect = 1e-04
Identities = 45/180 (25%), Positives = 65/180 (36%), Gaps = 11/180 (6%)
Frame = +2
Query: 98 LVSCSSDTTLKTWDAFSTGTCTRTLRQHSDYVT-----------CLAASEKNSNIVASGG 146
LV SS TL A GTC + VT C K + ++ G
Sbjct: 188 LVDWSSQNTL----AAGLGTCVYLWSASNSKVTKLCDLGPYDGVCSVQWTKEGSFISIGT 355
Query: 147 LGGEVFVWDLEAAHASAAKCIDATDDDTSNGINGSGNSLPLTSLRSISSSNSISVHTTQN 206
GG+V +WD K T G+ + + L S S +I H +
Sbjct: 356 NGGQVQIWD----GTKCKKVRTMGGHQTRTGVLAWNSRI----LASGSRDRNILQHDMRV 511
Query: 207 QGYIPIAAKGHKESVYALAMNEGGTLLVSGGTEKVVRVWDPRSGSKTMKLKGHTDNIRAL 266
GHK V L + L SGG + + VW+ S T++L HT ++A+
Sbjct: 512 PSDFIGKLVGHKSEVCGLKWSCDDRELASGGNDNQLLVWNQHSQQPTLRLTEHTAAVKAI 691
>TC87605 homologue to PIR|G96563|G96563 probable coatomer complex subunit
33791-27676 [imported] - Arabidopsis thaliana, partial
(23%)
Length = 765
Score = 43.9 bits (102), Expect = 7e-05
Identities = 25/78 (32%), Positives = 39/78 (49%), Gaps = 1/78 (1%)
Frame = +3
Query: 80 FESHVDWVND-AVLVGDSTLVSCSSDTTLKTWDAFSTGTCTRTLRQHSDYVTCLAASEKN 138
FE+H D++ AV ++S S D +K WD CT+ HS YV + + K+
Sbjct: 408 FEAHTDYIRCVAVHPTLPYVLSSSDDMLIKLWDWEKGWICTQIFEGHSHYVMQVTFNPKD 587
Query: 139 SNIVASGGLGGEVFVWDL 156
+N AS L + +W+L
Sbjct: 588 TNTFASASLDRTIKIWNL 641
>TC90611 similar to GP|10177223|dbj|BAB10298. contains similarity to
GTP-binding regulatory protein and WD-repeat
protein~gene_id:MPF21.14, partial (40%)
Length = 855
Score = 42.7 bits (99), Expect = 1e-04
Identities = 34/135 (25%), Positives = 57/135 (42%), Gaps = 10/135 (7%)
Frame = +1
Query: 33 KHCAGINCLAVLKS-AVSDGSDYLFTGSRDGRLKRWALGEDAATCSATFESHVDWVNDAV 91
K C ++ + + S A+S +L++ S D +K W +D + +H D +N V
Sbjct: 457 KKCTWVHHVDTVSSLALSKDGKFLYSVSWDRTIKIWRT-KDFTCLESITNAHDDAINAVV 633
Query: 92 LVGDSTLVSCSSDTTLKTWDAFS---------TGTCTRTLRQHSDYVTCLAASEKNSNIV 142
+ D T S S+D +K W + TL +H+ + L S S I+
Sbjct: 634 VSYDGTFYSGSADKRIKVWKNLQGDDHNKKKHKHSLVDTLEKHNSGINALVLSSDES-IL 810
Query: 143 ASGGLGGEVFVWDLE 157
SG + VW+ E
Sbjct: 811 YSGACDRSILVWEKE 855
>TC79322 similar to GP|13605815|gb|AAK32893.1 At2g32700/F24L7.16 {Arabidopsis
thaliana}, partial (59%)
Length = 1824
Score = 42.0 bits (97), Expect = 2e-04
Identities = 47/203 (23%), Positives = 81/203 (39%), Gaps = 1/203 (0%)
Frame = +2
Query: 49 SDGSDYLFTGSRDGRLKRWALGEDAATCSATFESHVDWVNDAVLVGDSTLVSCSS-DTTL 107
SDG L + D ++ W + + +T E H + D +ST ++ SS DTT+
Sbjct: 830 SDGK-LLASAGHDKKVVLWNM--ETLKTQSTPEEHTVIITDVRFRPNSTQLATSSFDTTV 1000
Query: 108 KTWDAFSTGTCTRTLRQHSDYVTCLAASEKNSNIVASGGLGGEVFVWDLEAAHASAAKCI 167
+ WDA + H+ +V L K +++ S E+ W++ S C
Sbjct: 1001 RLWDAADPSVSLQAYSGHTSHVASLDFHPKKNDLFCSCDDNNEIRFWNI-----SQYSCT 1165
Query: 168 DATDDDTSNGINGSGNSLPLTSLRSISSSNSISVHTTQNQGYIPIAAKGHKESVYALAMN 227
S + S L + +S N +S+ + + + +GH V+ + +
Sbjct: 1166 RVFKGG-STQVRFQPRS---GHLLAAASGNVVSLFDVETDRQMH-SLQGHSGEVHCVCWD 1330
Query: 228 EGGTLLVSGGTEKVVRVWDPRSG 250
G L S E V+VW SG
Sbjct: 1331 TNGDYLASVSQES-VKVWSLASG 1396
>TC80769 similar to GP|11141605|gb|AAG32022.1 LEUNIG {Arabidopsis thaliana},
partial (40%)
Length = 1362
Score = 41.6 bits (96), Expect = 3e-04
Identities = 34/148 (22%), Positives = 61/148 (40%)
Frame = +3
Query: 98 LVSCSSDTTLKTWDAFSTGTCTRTLRQHSDYVTCLAASEKNSNIVASGGLGGEVFVWDLE 157
L + S D T++ WD + G RT HS V L +++ S GE+ W +
Sbjct: 537 LATSSFDKTVRVWDVDNPGYSLRTFTGHSTSVMSLDFHPNKDDLICSCDGDGEIRYWSI- 713
Query: 158 AAHASAAKCIDATDDDTSNGINGSGNSLPLTSLRSISSSNSISVHTTQNQGYIPIAAKGH 217
+ C+ + G L + ++ N +S+ + Q + KGH
Sbjct: 714 ----NNGSCVRV----SKGGTTQMRFQPRLGRYLAAAAENIVSILDVETQA-CRYSLKGH 866
Query: 218 KESVYALAMNEGGTLLVSGGTEKVVRVW 245
+++ ++ + G LL S +E VR+W
Sbjct: 867 TKTIDSVCWDPSGELLAS-VSEDSVRIW 947
>BG648991 similar to PIR|C86239|C86 protein T10O24.21 [imported] -
Arabidopsis thaliana, partial (35%)
Length = 702
Score = 40.8 bits (94), Expect = 6e-04
Identities = 20/60 (33%), Positives = 29/60 (48%)
Frame = +3
Query: 98 LVSCSSDTTLKTWDAFSTGTCTRTLRQHSDYVTCLAASEKNSNIVASGGLGGEVFVWDLE 157
++S DT +K WD F+TG C RT HS V + + + + S G + WD E
Sbjct: 375 ILSAGMDTKVKIWDVFNTGKCMRTYMGHSKAVRDICFTNDGTKFL-SAGYDKNIKYWDTE 551
Score = 30.8 bits (68), Expect = 0.57
Identities = 14/59 (23%), Positives = 24/59 (39%)
Frame = +3
Query: 192 SISSSNSISVHTTQNQGYIPIAAKGHKESVYALAMNEGGTLLVSGGTEKVVRVWDPRSG 250
S + + N G GH ++V + GT +S G +K ++ WD +G
Sbjct: 381 SAGMDTKVKIWDVFNTGKCMRTYMGHSKAVRDICFTNDGTKFLSAGYDKNIKYWDTETG 557
>AW691821 similar to GP|10177570|db contains similarity to regulatory protein
Nedd1~gene_id:K18J17.16 {Arabidopsis thaliana}, partial
(19%)
Length = 648
Score = 40.0 bits (92), Expect = 0.001
Identities = 21/68 (30%), Positives = 39/68 (56%), Gaps = 5/68 (7%)
Frame = +2
Query: 205 QNQGYIPIAAK----GHKESVYALAM-NEGGTLLVSGGTEKVVRVWDPRSGSKTMKLKGH 259
Q+ G IP+A ++ES+ A++ N+ + SGG+ +VV++WD + LKGH
Sbjct: 353 QSMGTIPVAGMDSVDNNEESILAISFSNKASRYVCSGGSGQVVKIWDLQKKRCIKWLKGH 532
Query: 260 TDNIRALL 267
T+ + ++
Sbjct: 533 TNTVTGVM 556
Score = 28.5 bits (62), Expect = 2.9
Identities = 18/66 (27%), Positives = 30/66 (45%), Gaps = 6/66 (9%)
Frame = +2
Query: 98 LVSCSSDTTLKTW--DAFSTGTCT----RTLRQHSDYVTCLAASEKNSNIVASGGLGGEV 151
+ S D + W + S GT ++ + + + ++ S K S V SGG G V
Sbjct: 302 VASAGDDKRISLWRKNGQSMGTIPVAGMDSVDNNEESILAISFSNKASRYVCSGGSGQVV 481
Query: 152 FVWDLE 157
+WDL+
Sbjct: 482 KIWDLQ 499
>TC82910 similar to GP|19347844|gb|AAL86002.1 unknown protein {Arabidopsis
thaliana}, partial (19%)
Length = 821
Score = 40.0 bits (92), Expect = 0.001
Identities = 34/152 (22%), Positives = 68/152 (44%), Gaps = 10/152 (6%)
Frame = +1
Query: 129 VTCLAASEKNSNIVASGGLGGEVFVWDLEA--------AHASAAKCIDATDDDTSNGING 180
+TCLA + S I +G + G V + ++ +H+S+ +C+ G
Sbjct: 52 ITCLAINS-TSTIALTGSVDGSVHIVNITTGRVVSTLPSHSSSIECV---------GFAP 201
Query: 181 SGNSLPLTSLRSISSSNSISVHTTQNQGYIPIAAKGHKESVYALAMNE--GGTLLVSGGT 238
SG S +I + +++ ++ A+ E Y + + G + + +G
Sbjct: 202 SG------SWAAIGGMDKMTIWDVEHS-----LARSICEHEYGVTCSTWLGTSYVATGSN 348
Query: 239 EKVVRVWDPRSGSKTMKLKGHTDNIRALLLDS 270
+ VR+WD RSG +GH+++I++L L +
Sbjct: 349 DGAVRLWDSRSGECVRTFRGHSESIQSLSLSA 444
Score = 35.4 bits (80), Expect = 0.023
Identities = 33/133 (24%), Positives = 56/133 (41%)
Frame = +1
Query: 114 STGTCTRTLRQHSDYVTCLAASEKNSNIVASGGLGGEVFVWDLEAAHASAAKCIDATDDD 173
+TG TL HS + C+ + S A GG+ ++ +WD+E H+ A +
Sbjct: 133 TTGRVVSTLPSHSSSIECVGFAPSGS-WAAIGGMD-KMTIWDVE--HSLARSICEHEYGV 300
Query: 174 TSNGINGSGNSLPLTSLRSISSSNSISVHTTQNQGYIPIAAKGHKESVYALAMNEGGTLL 233
T + G TS + S++ G +GH ES+ +L+++ L
Sbjct: 301 TCSTWLG-------TSYVATGSNDGAVRLWDSRSGECVRTFRGHSESIQSLSLSANQEYL 459
Query: 234 VSGGTEKVVRVWD 246
VS + RV+D
Sbjct: 460 VSASLDHTARVFD 498
>BG649023 similar to GP|10178169|db contains similarity to WD-containing
protein~gene_id:K18G13.8 {Arabidopsis thaliana}, partial
(25%)
Length = 635
Score = 39.7 bits (91), Expect = 0.001
Identities = 23/74 (31%), Positives = 37/74 (49%)
Frame = +3
Query: 83 HVDWVNDAVLVGDSTLVSCSSDTTLKTWDAFSTGTCTRTLRQHSDYVTCLAASEKNSNIV 142
++D V D L+S S D T++ W S+ +C + HSDYVTC+ + +
Sbjct: 21 YLDDVLDLSWSKSQHLLSSSMDKTVRLWH-LSSKSCLKIF-SHSDYVTCIHFNPVDDRYF 194
Query: 143 ASGGLGGEVFVWDL 156
SG L +V +W +
Sbjct: 195 ISGSLDAKVRIWSI 236
>TC78016 similar to GP|21537191|gb|AAM61532.1 PRL1 protein {Arabidopsis
thaliana}, partial (81%)
Length = 1759
Score = 39.7 bits (91), Expect = 0.001
Identities = 33/144 (22%), Positives = 61/144 (41%), Gaps = 13/144 (9%)
Frame = +1
Query: 138 NSNIVASGGLGGEVFVWDLEAAHASAAKCIDATDDDTS-NGINGSGNSLPLTSLRSISSS 196
+SN++A G G DL+ A A + + T+ NG+ G + + S S
Sbjct: 316 SSNVLALPGPGDSK---DLQKGGAHNALVVGPSMPSTATNGLGFQGKNTVVVSTSGSSER 486
Query: 197 N---SISVHTTQNQGYIPI---------AAKGHKESVYALAMNEGGTLLVSGGTEKVVRV 244
N S + ++ P+ GH V ++A++ T +G ++ +++
Sbjct: 487 NFSTSALMERMPSKWPRPVWHAPWKNYRVISGHLGWVRSVAVDPSNTWFATGSADRTIKI 666
Query: 245 WDPRSGSKTMKLKGHTDNIRALLL 268
WD SG + L GH + +R L +
Sbjct: 667 WDLASGVLKLTLTGHIEQVRGLAI 738
Score = 37.0 bits (84), Expect = 0.008
Identities = 18/64 (28%), Positives = 27/64 (42%)
Frame = +1
Query: 208 GYIPIAAKGHKESVYALAMNEGGTLLVSGGTEKVVRVWDPRSGSKTMKLKGHTDNIRALL 267
G + + GH E V LA++ T + S G +K V+ WD GH + L
Sbjct: 682 GVLKLTLTGHIEQVRGLAISHKHTYMFSAGDDKQVKCWDLEQNKVIRSYHGHLSGVYCLA 861
Query: 268 LDST 271
+ T
Sbjct: 862 IHPT 873
Score = 33.1 bits (74), Expect = 0.12
Identities = 14/54 (25%), Positives = 27/54 (49%)
Frame = +3
Query: 213 AAKGHKESVYALAMNEGGTLLVSGGTEKVVRVWDPRSGSKTMKLKGHTDNIRAL 266
A GH+ +V ++ +V+G + +++WD R G L H ++RA+
Sbjct: 951 ALSGHENTVCSVFTRPTDPQVVTGSHDSTIKMWDLRYGKTMSTLTNHKKSVRAM 1112
Score = 32.7 bits (73), Expect = 0.15
Identities = 24/88 (27%), Positives = 34/88 (38%), Gaps = 1/88 (1%)
Frame = +1
Query: 47 AVSDGSDYLFTGSRDGRLKRWALGEDAATCSATFESHVDWVNDAVLVGDST-LVSCSSDT 105
AV + + TGS D +K W L + T H++ V + T + S D
Sbjct: 607 AVDPSNTWFATGSADRTIKIWDLA--SGVLKLTLTGHIEQVRGLAISHKHTYMFSAGDDK 780
Query: 106 TLKTWDAFSTGTCTRTLRQHSDYVTCLA 133
+K WD R+ H V CLA
Sbjct: 781 QVKCWD-LEQNKVIRSYHGHLSGVYCLA 861
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.312 0.127 0.372
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 8,195,393
Number of Sequences: 36976
Number of extensions: 111745
Number of successful extensions: 684
Number of sequences better than 10.0: 119
Number of HSP's better than 10.0 without gapping: 561
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 664
length of query: 271
length of database: 9,014,727
effective HSP length: 95
effective length of query: 176
effective length of database: 5,502,007
effective search space: 968353232
effective search space used: 968353232
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.9 bits)
S2: 57 (26.6 bits)
Lotus: description of TM0062.9


