

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0055.14
(267 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
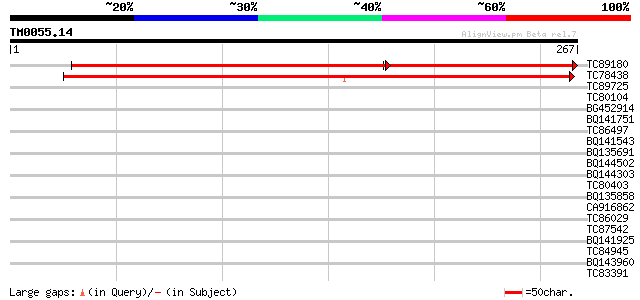
Sequences producing significant alignments: (bits) Value
TC89180 weakly similar to GP|13174241|gb|AAK14415.1 putative glu... 258 e-106
TC78438 weakly similar to GP|3738324|gb|AAC63665.1| unknown prot... 320 4e-88
TC89725 similar to PIR|T05494|T05494 glycine-rich protein T19K4.... 33 0.088
TC80104 similar to GP|20197699|gb|AAD20910.3 putative LRR recept... 33 0.15
BG452914 similar to GP|13374061|emb ZF-HD homeobox protein {Flav... 32 0.20
BQ141751 similar to SP|P33485|VNUA Probable nuclear antigen. [st... 32 0.26
TC86497 similar to GP|21535793|emb|CAC85725. putative carbamoyl ... 32 0.26
BQ141543 weakly similar to GP|6523547|emb| hydroxyproline-rich g... 32 0.26
BQ135691 similar to PIR|E96636|E966 hypothetical protein T7P1.21... 32 0.26
BQ144502 weakly similar to GP|6523547|emb hydroxyproline-rich gl... 32 0.26
BQ144303 weakly similar to GP|6523547|emb| hydroxyproline-rich g... 32 0.33
TC80403 similar to PIR|E96661|E96661 GMP synthase 61700-64653 [... 31 0.44
BQ135858 similar to GP|23491723|db formin homology protein A {Di... 31 0.44
CA916862 GP|12805231|gb Similar to mesenchyme homeobox 2 {Mus mu... 31 0.44
TC86029 homologue to GP|5802244|gb|AAD51625.1| seed maturation p... 25 0.56
TC87542 weakly similar to GP|22136436|gb|AAM91296.1 putative pro... 31 0.57
BQ141925 homologue to GP|10727150|gb| Rm62 gene product {alt 1} ... 30 0.74
TC84945 similar to GP|20259551|gb|AAM13895.1 unknown protein {Ar... 30 0.74
BQ143960 weakly similar to SP|P21997|SSGP_ Sulfated surface glyc... 30 0.74
TC83391 similar to GP|21555032|gb|AAM63759.1 unknown {Arabidopsi... 30 0.74
>TC89180 weakly similar to GP|13174241|gb|AAK14415.1 putative glutamine
synthetase {Oryza sativa}, partial (26%)
Length = 989
Score = 258 bits (659), Expect(2) = e-106
Identities = 117/150 (78%), Positives = 134/150 (89%)
Frame = +1
Query: 30 KRYALLMCGEDSEYLLKKHGGCFGFFTRMLAEEGERWDLYKVVNGEFPRHDELPLYDGFV 89
KRYA+LMCGEDSEYLLK+HGGC+GFFT+MLAEEGE WDLYKVVN EFP D++ YDGFV
Sbjct: 79 KRYAILMCGEDSEYLLKRHGGCYGFFTKMLAEEGETWDLYKVVNQEFPEDDDVDFYDGFV 258
Query: 90 ITGSCHDAHANHPWIHDLLSLVHTLDSINKKILGICFGHQILGRVLGGKVGRSSTGWDIG 149
ITGSC DAH+N PWIH LL+LVHTL+S+NKKILGICFGHQI+GR LGGKV RS+ GWDIG
Sbjct: 259 ITGSCKDAHSNDPWIHQLLTLVHTLNSLNKKILGICFGHQIIGRALGGKVVRSAAGWDIG 438
Query: 150 VTTMKLASSSSIALNLPSELSIFKCHRDEV 179
V+T+ L SSS +LNLPS+LS+FKCHRDEV
Sbjct: 439 VSTINLLQSSSSSLNLPSKLSLFKCHRDEV 528
Score = 144 bits (362), Expect(2) = e-106
Identities = 65/91 (71%), Positives = 79/91 (86%)
Frame = +2
Query: 177 DEVRNLPPKAEIIGWSEKTGIEMFRYGDHMLGIQGHPEFTIDILFHFIDRLTSRNLLQEC 236
D V +LP +AE+IGWSEKTGIEMFRYG+HMLGIQGHPEF IDI HFIDR+T+RNL+QE
Sbjct: 620 D*VLDLPAEAEVIGWSEKTGIEMFRYGNHMLGIQGHPEFNIDIFLHFIDRITNRNLIQEA 799
Query: 237 FAVDVKVKAALQEPNTEAWKRLCVNFLKDRL 267
FA DVK+KA L++P+T+AWK LC+ FLK +L
Sbjct: 800 FASDVKMKATLRDPDTDAWKTLCLTFLKGQL 892
>TC78438 weakly similar to GP|3738324|gb|AAC63665.1| unknown protein
{Arabidopsis thaliana}, partial (72%)
Length = 1163
Score = 320 bits (820), Expect = 4e-88
Identities = 151/244 (61%), Positives = 187/244 (75%), Gaps = 3/244 (1%)
Frame = +1
Query: 26 GGKGKRYALLMCGEDSEYLLKKHGGCFGFFTRMLAEEGERWDLYKVVNGEFPRHDELPLY 85
GG GKR+ +L+C +DSEY+ K +GG FG F RML EEGE WD+YKV GEFP ++L LY
Sbjct: 172 GGSGKRFGVLLCADDSEYVKKMYGGYFGVFVRMLEEEGESWDVYKVARGEFPDDEDLNLY 351
Query: 86 DGFVITGSCHDAHANHPWIHDLLSLVHTLDSINKKILGICFGHQILGRVLGGKVGRSSTG 145
DGFVITGSC DAH N W+ LL+L+ L+ +NKKILGICFGHQILGR +GGKV RS TG
Sbjct: 352 DGFVITGSCSDAHGNDTWVSQLLNLLKKLNDMNKKILGICFGHQILGRAIGGKVTRSPTG 531
Query: 146 WDIGVTTMKLA---SSSSIALNLPSELSIFKCHRDEVRNLPPKAEIIGWSEKTGIEMFRY 202
WDIGV + L+ S +L LP++LSI +CHRDE+R LP KAE+I S+KTGIEMFRY
Sbjct: 532 WDIGVRNITLSPSLPSPLSSLELPTKLSIIQCHRDELRELPAKAEVIAKSDKTGIEMFRY 711
Query: 203 GDHMLGIQGHPEFTIDILFHFIDRLTSRNLLQECFAVDVKVKAALQEPNTEAWKRLCVNF 262
GDH++GIQGHPE++ DIL + IDRL RN + E FA+ +K KA + EP+ EAWKRLC++F
Sbjct: 712 GDHIMGIQGHPEYSKDILLNIIDRLIQRNFITENFAMRLKEKAGMWEPDKEAWKRLCISF 891
Query: 263 LKDR 266
LK R
Sbjct: 892 LKGR 903
>TC89725 similar to PIR|T05494|T05494 glycine-rich protein T19K4.150 -
Arabidopsis thaliana, partial (17%)
Length = 378
Score = 33.5 bits (75), Expect = 0.088
Identities = 15/27 (55%), Positives = 17/27 (62%)
Frame = +1
Query: 17 GCGGGGGGGGGKGKRYALLMCGEDSEY 43
G GGGGGGGGG G Y+ CGE +
Sbjct: 94 GGGGGGGGGGGGGSCYS---CGESGHF 165
>TC80104 similar to GP|20197699|gb|AAD20910.3 putative LRR receptor protein
kinase {Arabidopsis thaliana}, partial (28%)
Length = 1549
Score = 32.7 bits (73), Expect = 0.15
Identities = 13/19 (68%), Positives = 14/19 (73%)
Frame = -3
Query: 14 MESGCGGGGGGGGGKGKRY 32
+E G GGGGGGGGG G Y
Sbjct: 1007 IEGGGGGGGGGGGGGGTEY 951
Score = 27.3 bits (59), Expect = 6.3
Identities = 11/16 (68%), Positives = 12/16 (74%)
Frame = -3
Query: 12 LEMESGCGGGGGGGGG 27
+E G GGGGGGGGG
Sbjct: 1007 IEGGGGGGGGGGGGGG 960
>BG452914 similar to GP|13374061|emb ZF-HD homeobox protein {Flaveria
bidentis}, partial (29%)
Length = 454
Score = 32.3 bits (72), Expect = 0.20
Identities = 19/47 (40%), Positives = 24/47 (50%), Gaps = 8/47 (17%)
Frame = +2
Query: 11 ELEMESGCGGGGGGGGGKG-----KRYALLMCGEDSEYLL---KKHG 49
++ S GGGGGGGGG G KR+ E E LL ++HG
Sbjct: 221 DMSNPSSSGGGGGGGGGSGGSGSRKRFRTKFTQEQKEKLLAFAEEHG 361
>BQ141751 similar to SP|P33485|VNUA Probable nuclear antigen. [strain
Kaplan PRV] {Pseudorabies virus}, partial (2%)
Length = 1162
Score = 32.0 bits (71), Expect = 0.26
Identities = 13/23 (56%), Positives = 17/23 (73%)
Frame = -1
Query: 7 LLERELEMESGCGGGGGGGGGKG 29
LL+ ++ +E G GG GGGGGG G
Sbjct: 70 LLKIQVGVEQGAGGAGGGGGGGG 2
>TC86497 similar to GP|21535793|emb|CAC85725. putative carbamoyl phosphate
synthase small subunit {Nicotiana tabacum}, partial (64%)
Length = 1199
Score = 32.0 bits (71), Expect = 0.26
Identities = 15/29 (51%), Positives = 20/29 (68%), Gaps = 2/29 (6%)
Frame = +2
Query: 111 VHTLDSINKKI--LGICFGHQILGRVLGG 137
V T+ +I K+ GIC GHQ+LG+ LGG
Sbjct: 1064 VETVKNIIGKVPVFGICMGHQLLGQALGG 1150
>BQ141543 weakly similar to GP|6523547|emb| hydroxyproline-rich glycoprotein
DZ-HRGP {Volvox carteri f. nagariensis}, partial (15%)
Length = 1271
Score = 32.0 bits (71), Expect = 0.26
Identities = 14/19 (73%), Positives = 15/19 (78%), Gaps = 2/19 (10%)
Frame = +1
Query: 13 EMESGCGGGGG--GGGGKG 29
E E+GCGGGGG GGGG G
Sbjct: 475 EAEAGCGGGGGEVGGGGDG 531
>BQ135691 similar to PIR|E96636|E966 hypothetical protein T7P1.21
[imported] - Arabidopsis thaliana, partial (4%)
Length = 1215
Score = 32.0 bits (71), Expect = 0.26
Identities = 13/21 (61%), Positives = 15/21 (70%)
Frame = -3
Query: 9 ERELEMESGCGGGGGGGGGKG 29
+R E + G GGGGGGGGG G
Sbjct: 64 QRRKEQKRGRGGGGGGGGGGG 2
Score = 29.6 bits (65), Expect = 1.3
Identities = 12/21 (57%), Positives = 14/21 (66%)
Frame = -1
Query: 9 ERELEMESGCGGGGGGGGGKG 29
+ E + G GGGGGGGGG G
Sbjct: 63 KEEKSKKGGGGGGGGGGGGGG 1
>BQ144502 weakly similar to GP|6523547|emb hydroxyproline-rich glycoprotein
DZ-HRGP {Volvox carteri f. nagariensis}, partial (39%)
Length = 1358
Score = 32.0 bits (71), Expect = 0.26
Identities = 11/12 (91%), Positives = 12/12 (99%)
Frame = +3
Query: 18 CGGGGGGGGGKG 29
CGGGGGGGGG+G
Sbjct: 132 CGGGGGGGGGRG 167
Score = 27.7 bits (60), Expect = 4.8
Identities = 11/15 (73%), Positives = 12/15 (79%)
Frame = +1
Query: 17 GCGGGGGGGGGKGKR 31
G GGGG GGGG+G R
Sbjct: 526 GDGGGGTGGGGRGGR 570
Score = 26.9 bits (58), Expect = 8.2
Identities = 11/17 (64%), Positives = 12/17 (69%)
Frame = +2
Query: 15 ESGCGGGGGGGGGKGKR 31
E G GGGGGGG +G R
Sbjct: 665 EEGGGGGGGGGWRRGAR 715
Score = 26.9 bits (58), Expect = 8.2
Identities = 12/24 (50%), Positives = 14/24 (58%)
Frame = +1
Query: 6 MLLERELEMESGCGGGGGGGGGKG 29
+ L R SG GGGG GGG +G
Sbjct: 625 IFLARGTRSHSGGGGGGRGGGRRG 696
>BQ144303 weakly similar to GP|6523547|emb| hydroxyproline-rich glycoprotein
DZ-HRGP {Volvox carteri f. nagariensis}, partial (11%)
Length = 1158
Score = 31.6 bits (70), Expect = 0.33
Identities = 12/20 (60%), Positives = 16/20 (80%)
Frame = +2
Query: 8 LERELEMESGCGGGGGGGGG 27
+ER+ ++ G GGGGGGGGG
Sbjct: 410 IERQYQVLGGRGGGGGGGGG 469
>TC80403 similar to PIR|E96661|E96661 GMP synthase 61700-64653 [imported] -
Arabidopsis thaliana, partial (24%)
Length = 558
Score = 31.2 bits (69), Expect = 0.44
Identities = 26/98 (26%), Positives = 45/98 (45%), Gaps = 3/98 (3%)
Frame = +2
Query: 89 VITGSCHDAHA-NHPWIHDLLSLVHTLDSINKKILGICFGHQILGRVLGGKVGRSSTGWD 147
+++G H H + P D + S +LGIC+G Q+L LGG+V R +
Sbjct: 269 ILSGGPHSVHTPDSPSFPD--GFLQWAQSNAVPVLGICYGLQLLVSRLGGEV-RVGNTQE 439
Query: 148 IGVTTMKLASSSSI--ALNLPSELSIFKCHRDEVRNLP 183
G +K+ ++S++ + ++ H DE LP
Sbjct: 440 YGRMEIKVXNASALFPVEKVGHSQLVWMSHGDEAVTLP 553
>BQ135858 similar to GP|23491723|db formin homology protein A {Dictyostelium
discoideum}, partial (4%)
Length = 1033
Score = 31.2 bits (69), Expect = 0.44
Identities = 11/11 (100%), Positives = 11/11 (100%)
Frame = -1
Query: 17 GCGGGGGGGGG 27
GCGGGGGGGGG
Sbjct: 712 GCGGGGGGGGG 680
>CA916862 GP|12805231|gb Similar to mesenchyme homeobox 2 {Mus musculus},
partial (5%)
Length = 727
Score = 31.2 bits (69), Expect = 0.44
Identities = 12/24 (50%), Positives = 15/24 (62%)
Frame = -1
Query: 6 MLLERELEMESGCGGGGGGGGGKG 29
M++ + GCGG GGGGGG G
Sbjct: 439 MMMMMMMNCWGGCGGSGGGGGGGG 368
>TC86029 homologue to GP|5802244|gb|AAD51625.1| seed maturation protein PM37
{Glycine max}, complete
Length = 1572
Score = 25.4 bits (54), Expect(2) = 0.56
Identities = 13/27 (48%), Positives = 16/27 (59%)
Frame = +1
Query: 20 GGGGGGGGKGKRYALLMCGEDSEYLLK 46
GGGGGG +G+R GED + LK
Sbjct: 388 GGGGGGSSRGRRQRR---GEDVVHPLK 459
Score = 23.9 bits (50), Expect(2) = 0.56
Identities = 8/12 (66%), Positives = 10/12 (82%)
Frame = +1
Query: 14 MESGCGGGGGGG 25
++ G GGGGGGG
Sbjct: 319 LKEGMGGGGGGG 354
>TC87542 weakly similar to GP|22136436|gb|AAM91296.1 putative protein
{Arabidopsis thaliana}, partial (39%)
Length = 2234
Score = 30.8 bits (68), Expect = 0.57
Identities = 13/19 (68%), Positives = 15/19 (78%)
Frame = -3
Query: 12 LEMESGCGGGGGGGGGKGK 30
L ESG GGGGGGGG+G+
Sbjct: 450 LTRESGI*GGGGGGGGEGE 394
>BQ141925 homologue to GP|10727150|gb| Rm62 gene product {alt 1}
[Drosophila melanogaster], partial (3%)
Length = 1059
Score = 30.4 bits (67), Expect = 0.74
Identities = 12/20 (60%), Positives = 14/20 (70%)
Frame = -3
Query: 10 RELEMESGCGGGGGGGGGKG 29
+ + E G GGGGGGGGG G
Sbjct: 73 KRIGKEKGGGGGGGGGGGGG 14
Score = 28.9 bits (63), Expect = 2.2
Identities = 11/16 (68%), Positives = 13/16 (80%)
Frame = -3
Query: 15 ESGCGGGGGGGGGKGK 30
+ G GGGGGGGGG G+
Sbjct: 55 KGGGGGGGGGGGGGGE 8
Score = 27.7 bits (60), Expect = 4.8
Identities = 11/20 (55%), Positives = 13/20 (65%)
Frame = -1
Query: 11 ELEMESGCGGGGGGGGGKGK 30
E E G GGGGG GGG+ +
Sbjct: 63 EKRKEGGGGGGGGAGGGENQ 4
Score = 27.3 bits (59), Expect = 6.3
Identities = 12/22 (54%), Positives = 13/22 (58%)
Frame = -1
Query: 9 ERELEMESGCGGGGGGGGGKGK 30
E LE GGGGGGG G G+
Sbjct: 75 ENGLEKRKEGGGGGGGGAGGGE 10
Score = 26.9 bits (58), Expect = 8.2
Identities = 12/27 (44%), Positives = 17/27 (62%)
Frame = -2
Query: 4 RTMLLERELEMESGCGGGGGGGGGKGK 30
R +L+ + + E GGGGGGG G G+
Sbjct: 89 RKGILKTDWKRERRGGGGGGGGRGGGR 9
>TC84945 similar to GP|20259551|gb|AAM13895.1 unknown protein {Arabidopsis
thaliana}, partial (18%)
Length = 785
Score = 30.4 bits (67), Expect = 0.74
Identities = 12/15 (80%), Positives = 12/15 (80%)
Frame = -1
Query: 15 ESGCGGGGGGGGGKG 29
ES CGGGGGGGG G
Sbjct: 146 ESDCGGGGGGGGRGG 102
>BQ143960 weakly similar to SP|P21997|SSGP_ Sulfated surface glycoprotein
185 (SSG 185). {Volvox carteri}, partial (13%)
Length = 1023
Score = 30.4 bits (67), Expect = 0.74
Identities = 12/15 (80%), Positives = 13/15 (86%)
Frame = -3
Query: 17 GCGGGGGGGGGKGKR 31
G GGGGGGGGG+G R
Sbjct: 70 GGGGGGGGGGGEGGR 26
Score = 28.1 bits (61), Expect = 3.7
Identities = 13/30 (43%), Positives = 18/30 (59%)
Frame = -3
Query: 1 MKGRTMLLERELEMESGCGGGGGGGGGKGK 30
++G + E+ E G GGGGGGGG G+
Sbjct: 115 VRGEKKMGEKWGGNEGGGGGGGGGGGEGGR 26
Score = 27.7 bits (60), Expect = 4.8
Identities = 11/15 (73%), Positives = 12/15 (79%)
Frame = -3
Query: 17 GCGGGGGGGGGKGKR 31
G GGGGGG GG+G R
Sbjct: 61 GGGGGGGGEGGRGGR 17
>TC83391 similar to GP|21555032|gb|AAM63759.1 unknown {Arabidopsis
thaliana}, partial (82%)
Length = 781
Score = 30.4 bits (67), Expect = 0.74
Identities = 12/17 (70%), Positives = 12/17 (70%)
Frame = -3
Query: 19 GGGGGGGGGKGKRYALL 35
GGGGGGGGG G Y L
Sbjct: 119 GGGGGGGGGNGNVYVCL 69
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.322 0.143 0.450
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 9,816,661
Number of Sequences: 36976
Number of extensions: 157223
Number of successful extensions: 4365
Number of sequences better than 10.0: 134
Number of HSP's better than 10.0 without gapping: 1563
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 3221
length of query: 267
length of database: 9,014,727
effective HSP length: 94
effective length of query: 173
effective length of database: 5,538,983
effective search space: 958244059
effective search space used: 958244059
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 57 (26.6 bits)
Lotus: description of TM0055.14


