

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
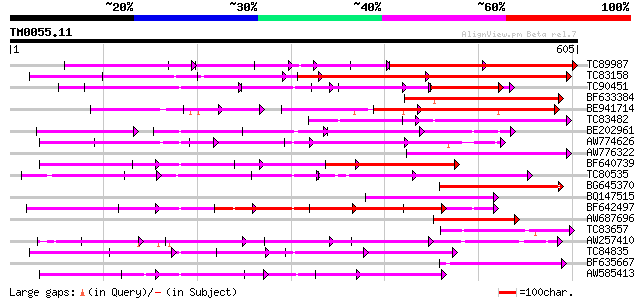
Query= TM0055.11
(605 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC89987 weakly similar to PIR|D86443|D86443 probable PPR-repeat ... 162 1e-59
TC83158 weakly similar to PIR|H86191|H86191 hypothetical protein... 214 8e-56
TC90451 similar to PIR|H84508|H84508 hypothetical protein At2g13... 126 3e-48
BF633384 similar to PIR|T46179|T46 hypothetical protein T8H10.30... 147 1e-35
BE941714 weakly similar to GP|11994734|dbj selenium-binding prot... 146 3e-35
TC83482 weakly similar to GP|9758453|dbj|BAB08982.1 selenium-bin... 140 2e-33
BE202961 weakly similar to GP|6016735|gb|A hypothetical protein ... 129 4e-30
AW774626 similar to PIR|T47909|T47 hypothetical protein T20K12.7... 124 1e-28
AW776322 weakly similar to PIR|H84859|H84 hypothetical protein A... 122 5e-28
BF640739 weakly similar to GP|7363286|dbj Similar to Arabidopsis... 118 7e-27
TC80535 weakly similar to GP|14335136|gb|AAK59848.1 At2g15690/F9... 113 2e-25
BG645370 weakly similar to PIR|T00405|T004 hypothetical protein ... 111 7e-25
BQ147515 weakly similar to PIR|T05355|T05 hypothetical protein F... 104 9e-23
BF642497 weakly similar to PIR|H96713|H96 hypothetical protein T... 102 4e-22
AW687696 weakly similar to PIR|T49965|T499 hypothetical protein ... 101 9e-22
TC83657 weakly similar to GP|11994382|dbj|BAB02341. gb|AAB82628.... 94 2e-19
AW257410 weakly similar to GP|10177533|db selenium-binding prote... 93 3e-19
TC84835 weakly similar to PIR|T02328|T02328 hypothetical protein... 85 9e-17
BF635667 weakly similar to PIR|C85359|C853 hypothetical protein ... 84 2e-16
AW585413 weakly similar to PIR|B85153|B851 hypothetical protein ... 77 1e-14
>TC89987 weakly similar to PIR|D86443|D86443 probable PPR-repeat protein
[imported] - Arabidopsis thaliana, partial (62%)
Length = 1559
Score = 162 bits (409), Expect(2) = 1e-59
Identities = 77/200 (38%), Positives = 126/200 (62%)
Frame = +2
Query: 406 LEGSLQIFLEMQNRDSISWSTMISAYGLHGHGEQALKLFSEMKERGVKPDALTFLAVLSA 465
L L++F M ++ S++ MIS +HG G++ALK+FSEM E G+ PD + ++ V SA
Sbjct: 563 LRKGLRVFKNMSEKNRYSYTVMISGLAIHGRGKEALKVFSEMIEEGLAPDDVVYVGVFSA 742
Query: 466 CNHAGLVTEGQQVFKQVNADYEIPLTIEHYACLVDLLGRSGKVEDALEIMRTMPMKPSTR 525
C+HAGLV EG Q FK + +++I T++HY C+VDLLGR G +++A E++++M +KP+
Sbjct: 743 CSHAGLVEEGLQCFKSMQFEHKIEPTVQHYGCMVDLLGRFGMLKEAYELIKSMSIKPNDV 922
Query: 526 IWSSLVSACKLHGRLDIAEMLGPQLIRSEPDNAANYTLLNMIYAEHGHWLHKEQVMEAMK 585
IW SL+SACK+H L+I ++ L +N+ +Y +L +YA+ W ++ +
Sbjct: 923 IWRSLLSACKVHHNLEIGKIAAENLFMLNQNNSGDYLVLANMYAKAQKWDDVAKIRTKLA 1102
Query: 586 LQRLKKCYGFSRIETGNQCF 605
+ L + GFS IE + +
Sbjct: 1103ERNLVQTPGFSLIEAKRKVY 1162
Score = 86.7 bits (213), Expect(2) = 1e-59
Identities = 51/147 (34%), Positives = 82/147 (55%), Gaps = 1/147 (0%)
Frame = +1
Query: 262 ACAKPGFVKHGKEIHGYAFRHGIESSHIFSSALINMYCQSGESLHLAEQIFERSSFRDVV 321
AC+ G V G ++HG+ F+ G+E I ++LINMY + GE + + +F + V
Sbjct: 130 ACSLLGVVDEGIQVHGHVFKMGLEGDVIVQNSLINMYGKCGEIKNACD-VFNGMDEKSVA 306
Query: 322 LWSSIIGSYSQRGDSYKALTLFNKMLTE-ETEPNYVTLLAVISACTNLSSLKHGCGLHGF 380
WS+IIG+++ + L L KM +E TL+ V+SACT+L S G +HG
Sbjct: 307 SWSAIIGAHACVEMWNECLMLLGKMSSEGRCRVEESTLVNVLSACTHLGSPDLGKCIHGI 486
Query: 381 ILKFGLSFSISLSNALMNMYAKCGCLE 407
+L+ ++ + +L++MY K GCLE
Sbjct: 487 LLRNISELNVVVKTSLIDMYVKSGCLE 567
Score = 72.0 bits (175), Expect = 6e-13
Identities = 43/147 (29%), Positives = 72/147 (48%), Gaps = 1/147 (0%)
Frame = +1
Query: 364 ACTNLSSLKHGCGLHGFILKFGLSFSISLSNALMNMYAKCGCLEGSLQIFLEMQNRDSIS 423
AC+ L + G +HG + K GL + + N+L+NMY KCG ++ + +F M + S
Sbjct: 130 ACSLLGVVDEGIQVHGHVFKMGLEGDVIVQNSLINMYGKCGEIKNACDVFNGMDEKSVAS 309
Query: 424 WSTMISAYGLHGHGEQALKLFSEMKERG-VKPDALTFLAVLSACNHAGLVTEGQQVFKQV 482
WS +I A+ + L L +M G + + T + VLSAC H G G+ + +
Sbjct: 310 WSAIIGAHACVEMWNECLMLLGKMSSEGRCRVEESTLVNVLSACTHLGSPDLGKCIHGIL 489
Query: 483 NADYEIPLTIEHYACLVDLLGRSGKVE 509
+ L + L+D+ +SG +E
Sbjct: 490 LRNIS-ELNVVVKTSLIDMYVKSGCLE 567
Score = 62.4 bits (150), Expect = 5e-10
Identities = 42/134 (31%), Positives = 67/134 (49%), Gaps = 1/134 (0%)
Frame = +1
Query: 170 GRQIHALVVVDGRVEHSVVLSTALVDFYFRCDDALMASRVFDGMEVKNEVSWTAMISGCI 229
G Q+H V G +E V++ +L++ Y +C + A VF+GM+ K+ SW+A+I
Sbjct: 160 GIQVHGHVFKMG-LEGDVIVQNSLINMYGKCGEIKNACDVFNGMDEKSVASWSAIIGAHA 336
Query: 230 ANLDYDAAFARFRAMQVEG-VDPNRVTLIALLPACAKPGFVKHGKEIHGYAFRHGIESSH 288
++ M EG TL+ +L AC G GK IHG R+ E +
Sbjct: 337 CVEMWNECLMLLGKMSSEGRCRVEESTLVNVLSACTHLGSPDLGKCIHGILLRNISELNV 516
Query: 289 IFSSALINMYCQSG 302
+ ++LI+MY +SG
Sbjct: 517 VVKTSLIDMYVKSG 558
Score = 62.4 bits (150), Expect(2) = 3e-19
Identities = 43/142 (30%), Positives = 71/142 (49%), Gaps = 1/142 (0%)
Frame = +1
Query: 59 ACSSLHSHTLGTLLHGLALTTGSHSDPVVSNSLISLYAKFSDIQSARQVFDTMPHRDPIS 118
ACS L G +HG G D +V NSLI++Y K +I++A VF+ M + S
Sbjct: 130 ACSLLGVVDEGIQVHGHVFKMGLEGDVIVQNSLINMYGKCGEIKNACDVFNGMDEKSVAS 309
Query: 119 WNSMINAYLQNGCLEEALEMLKGVYLLGLVPKPE-LLASMVSLCGRKMGSRIGRQIHALV 177
W+++I A+ E L +L + G E L +++S C +G+ IH +
Sbjct: 310 WSAIIGAHACVEMWNECLMLLGKMSSEGRCRVEESTLVNVLSACTHLGSPDLGKCIHG-I 486
Query: 178 VVDGRVEHSVVLSTALVDFYFR 199
++ E +VV+ T+L+D Y +
Sbjct: 487 LLRNISELNVVVKTSLIDMYVK 552
Score = 50.8 bits (120), Expect(2) = 3e-19
Identities = 37/132 (28%), Positives = 57/132 (43%), Gaps = 1/132 (0%)
Frame = +2
Query: 199 RCDDALMASRVFDGMEVKNEVSWTAMISGCIANLDYDAAFARFRAMQVEGVDPNRVTLIA 258
R D RVF M KN S+T MISG + A F M EG+ P+ V +
Sbjct: 551 RVDVLRKGLRVFKNMSEKNRYSYTVMISGLAIHGRGKEALKVFSEMIEEGLAPDDVVYVG 730
Query: 259 LLPACAKPGFVKHGKE-IHGYAFRHGIESSHIFSSALINMYCQSGESLHLAEQIFERSSF 317
+ AC+ G V+ G + F H IE + ++++ + G E I S
Sbjct: 731 VFSACSHAGLVEEGLQCFKSMQFEHKIEPTVQHYGCMVDLLGRFGMLKEAYELIKSMSIK 910
Query: 318 RDVVLWSSIIGS 329
+ V+W S++ +
Sbjct: 911 PNDVIWRSLLSA 946
Score = 38.5 bits (88), Expect = 0.008
Identities = 20/47 (42%), Positives = 27/47 (56%)
Frame = +1
Query: 51 SVLPSVIKACSSLHSHTLGTLLHGLALTTGSHSDPVVSNSLISLYAK 97
S L +V+ AC+ L S LG +HG+ L S + VV SLI +Y K
Sbjct: 412 STLVNVLSACTHLGSPDLGKCIHGILLRNISELNVVVKTSLIDMYVK 552
>TC83158 weakly similar to PIR|H86191|H86191 hypothetical protein [imported]
- Arabidopsis thaliana, partial (42%)
Length = 1194
Score = 214 bits (545), Expect = 8e-56
Identities = 101/292 (34%), Positives = 182/292 (61%)
Frame = +2
Query: 308 AEQIFERSSFRDVVLWSSIIGSYSQRGDSYKALTLFNKMLTEETEPNYVTLLAVISACTN 367
A ++F++ ++VV W+ +IG + ++ +AL F +M P++VT++A+ISAC N
Sbjct: 17 ALKLFDKLPVKNVVSWTVVIGGFVKKECYEEALECFREMQLAGVVPDFVTVIAIISACAN 196
Query: 368 LSSLKHGCGLHGFILKFGLSFSISLSNALMNMYAKCGCLEGSLQIFLEMQNRDSISWSTM 427
L +L G +H ++K ++ + N+L++MYA+CGC+E + Q+F M R+ +SW+++
Sbjct: 197 LGALGLGLWVHRLVMKKEFRDNVKVLNSLIDMYARCGCIELARQVFDGMSQRNLVSWNSI 376
Query: 428 ISAYGLHGHGEQALKLFSEMKERGVKPDALTFLAVLSACNHAGLVTEGQQVFKQVNADYE 487
I + ++G ++AL F MK+ G++P+ +++ + L+AC+HAGL+ EG ++F + D+
Sbjct: 377 IVGFAVNGLADKALSFFRSMKKEGLEPNGVSYTSALTACSHAGLIDEGLKIFADIKRDHR 556
Query: 488 IPLTIEHYACLVDLLGRSGKVEDALEIMRTMPMKPSTRIWSSLVSACKLHGRLDIAEMLG 547
IEHY CLVDL R+G++++A ++++ MPM P+ + SL++AC+ G +++AE +
Sbjct: 557 NSPRIEHYGCLVDLYSRAGRLKEAWDVIKKMPMMPNEVVLGSLLAACRTQGDVELAEKVM 736
Query: 548 PQLIRSEPDNAANYTLLNMIYAEHGHWLHKEQVMEAMKLQRLKKCYGFSRIE 599
+ P +NY L + IYA G W +V MK + L+K FS IE
Sbjct: 737 KYQVELYPGGDSNYVLFSNIYAAVGKWDGASKVRREMKERGLQKNLAFSSIE 892
Score = 118 bits (296), Expect = 6e-27
Identities = 70/237 (29%), Positives = 123/237 (51%), Gaps = 1/237 (0%)
Frame = +2
Query: 100 DIQSARQVFDTMPHRDPISWNSMINAYLQNGCLEEALEMLKGVYLLGLVPKPELLASMVS 159
D+ A ++FD +P ++ +SW +I +++ C EEALE + + L G+VP + +++S
Sbjct: 5 DVDDALKLFDKLPVKNVVSWTVVIGGFVKKECYEEALECFREMQLAGVVPDFVTVIAIIS 184
Query: 160 LCGRKMGSRIGRQIHALVVVDGRVEHSVVLSTALVDFYFRCDDALMASRVFDGMEVKNEV 219
C +G +H L V+ +V + +L+D Y RC +A +VFDGM +N V
Sbjct: 185 ACANLGALGLGLWVHRL-VMKKEFRDNVKVLNSLIDMYARCGCIELARQVFDGMSQRNLV 361
Query: 220 SWTAMISGCIANLDYDAAFARFRAMQVEGVDPNRVTLIALLPACAKPGFVKHGKEIHGYA 279
SW ++I G N D A + FR+M+ EG++PN V+ + L AC+ G + G +I
Sbjct: 362 SWNSIIVGFAVNGLADKALSFFRSMKKEGLEPNGVSYTSALTACSHAGLIDEGLKIFADI 541
Query: 280 FRHGIESSHI-FSSALINMYCQSGESLHLAEQIFERSSFRDVVLWSSIIGSYSQRGD 335
R S I L+++Y ++G + I + + V+ S++ + +GD
Sbjct: 542 KRDHRNSPRIEHYGCLVDLYSRAGRLKEAWDVIKKMPMMPNEVVLGSLLAACRTQGD 712
Score = 114 bits (285), Expect = 1e-25
Identities = 78/251 (31%), Positives = 132/251 (52%), Gaps = 2/251 (0%)
Frame = +2
Query: 201 DDALMASRVFDGMEVKNEVSWTAMISGCIANLDYDAAFARFRAMQVEGVDPNRVTLIALL 260
DDAL ++FD + VKN VSWT +I G + Y+ A FR MQ+ GV P+ VT+IA++
Sbjct: 11 DDAL---KLFDKLPVKNVVSWTVVIGGFVKKECYEEALECFREMQLAGVVPDFVTVIAII 181
Query: 261 PACAKPGFVKHGKEIHGYAFRHGIESSHIFSSALINMYCQSGESLHLAEQIFERSSFRDV 320
ACA G + G +H + + ++LI+MY + G + LA Q+F+ S R++
Sbjct: 182 SACANLGALGLGLWVHRLVMKKEFRDNVKVLNSLIDMYARCG-CIELARQVFDGMSQRNL 358
Query: 321 VLWSSIIGSYSQRGDSYKALTLFNKMLTEETEPNYVTLLAVISACTNLSSLKHGCGLHGF 380
V W+SII ++ G + KAL+ F M E EPN V+ + ++AC++ + G +
Sbjct: 359 VSWNSIIVGFAVNGLADKALSFFRSMKKEGLEPNGVSYTSALTACSHAGLIDEGLKIFAD 538
Query: 381 ILK-FGLSFSISLSNALMNMYAKCGCLEGSLQIFLEM-QNRDSISWSTMISAYGLHGHGE 438
I + S I L+++Y++ G L+ + + +M + + ++++A G E
Sbjct: 539 IKRDHRNSPRIEHYGCLVDLYSRAGRLKEAWDVIKKMPMMPNEVVLGSLLAACRTQGDVE 718
Query: 439 QALKLFSEMKE 449
A K+ E
Sbjct: 719 LAEKVMKYQVE 751
Score = 87.0 bits (214), Expect = 2e-17
Identities = 76/279 (27%), Positives = 123/279 (43%), Gaps = 4/279 (1%)
Frame = +2
Query: 22 IKSLISQGLYHQTLELFTQLHFSAPHSISSVLPSVIKACSSLHSHTLGTLLHGLALTTGS 81
I + + Y + LE F ++ + + ++I AC++L + LG +H L +
Sbjct: 74 IGGFVKKECYEEALECFREMQLAGVVPDFVTVIAIISACANLGALGLGLWVHRLVMKKEF 253
Query: 82 HSDPVVSNSLISLYAKFSDIQSARQVFDTMPHRDPISWNSMINAYLQNGCLEEALEMLKG 141
+ V NSLI +YA+ I+ ARQVFD M R+ +SWNS+I + NG ++AL +
Sbjct: 254 RDNVKVLNSLIDMYARCGCIELARQVFDGMSQRNLVSWNSIIVGFAVNGLADKALSFFRS 433
Query: 142 VYLLGLVPKPELLASMVSLCGRKMGSRIGRQIHALVVVDGRVEHSVVLSTALVDFYFRCD 201
+ GL P S ++ C G +I A + D R + LVD Y R
Sbjct: 434 MKKEGLEPNGVSYTSALTACSHAGLIDEGLKIFADIKRDHRNSPRIEHYGCLVDLYSRAG 613
Query: 202 DALMASRVFDGME-VKNEVSWTAMISGCIANLDYDAA--FARFRAMQVEGVDPNRVTLIA 258
A V M + NEV ++++ C D + A +++ G D N V
Sbjct: 614 RLKEAWDVIKKMPMMPNEVVLGSLLAACRTQGDVELAEKVMKYQVELYPGGDSNYVLFSN 793
Query: 259 LLPACAK-PGFVKHGKEIHGYAFRHGIESSHIFSSALIN 296
+ A K G K +E+ G++ + FSS I+
Sbjct: 794 IYAAVGKWDGASKVRREMK----ERGLQKNLAFSSIEID 898
>TC90451 similar to PIR|H84508|H84508 hypothetical protein At2g13600
[imported] - Arabidopsis thaliana, partial (12%)
Length = 1362
Score = 126 bits (316), Expect(2) = 3e-48
Identities = 66/207 (31%), Positives = 123/207 (58%), Gaps = 2/207 (0%)
Frame = +2
Query: 248 GVDPNRVTLIALLPACAKPGFVKHGKEIHGYAFRHGIESSHIFSSALINMYCQSGESLHL 307
GV P+R TL +L A ++ ++ G+++H Y + G++S+ S IN+YC++G+
Sbjct: 470 GVLPDRYTLPIVLKAVSQSFAIQLGQQVHSYGIKLGLQSNEYCESGFINLYCKAGD-FDS 646
Query: 308 AEQIFERSSFRDVVLWSSIIGSYSQRGDSYKALTLFNKMLTEETEPNYVTLLAVISACTN 367
A ++F+ + + W+++I SQ G + A+ +F M EP+ +T+++V+SAC +
Sbjct: 647 AHKVFDENHEPKLGSWNALISGLSQGGLAMDAIVVFVDMKRHGFEPDGITMVSVMSACGS 826
Query: 368 LSSLKHGCGLHGFILKFGLS--FSISLSNALMNMYAKCGCLEGSLQIFLEMQNRDSISWS 425
+ L LH ++ + + I +SN+L++MY KCG ++ + ++F M++R+ SW+
Sbjct: 827 IGDLYLALQLHKYVFQAKTNEWTVILMSNSLIDMYGKCGRMDLAYEVFATMEDRNVSSWT 1006
Query: 426 TMISAYGLHGHGEQALKLFSEMKERGV 452
+MI Y +HGH ++AL F MK GV
Sbjct: 1007SMIVGYAMHGHAKEALGCFHCMKGVGV 1087
Score = 96.3 bits (238), Expect = 3e-20
Identities = 60/217 (27%), Positives = 108/217 (49%), Gaps = 1/217 (0%)
Frame = +2
Query: 323 WSSIIGSYSQRGDSYKALTLFNKMLTEETEPNYVTLLAVISACTNLSSLKHGCGLHGFIL 382
W++II SY++ AL ++ ML P+ TL V+ A + +++ G +H + +
Sbjct: 389 WNNIIRSYTRLESPQNALRIYVSMLRAGVLPDRYTLPIVLKAVSQSFAIQLGQQVHSYGI 568
Query: 383 KFGLSFSISLSNALMNMYAKCGCLEGSLQIFLEMQNRDSISWSTMISAYGLHGHGEQALK 442
K GL + + +N+Y K G + + ++F E SW+ +IS G A+
Sbjct: 569 KLGLQSNEYCESGFINLYCKAGDFDSAHKVFDENHEPKLGSWNALISGLSQGGLAMDAIV 748
Query: 443 LFSEMKERGVKPDALTFLAVLSACNHAGLVTEGQQVFKQV-NADYEIPLTIEHYACLVDL 501
+F +MK G +PD +T ++V+SAC G + Q+ K V A I L+D+
Sbjct: 749 VFVDMKRHGFEPDGITMVSVMSACGSIGDLYLALQLHKYVFQAKTNEWTVILMSNSLIDM 928
Query: 502 LGRSGKVEDALEIMRTMPMKPSTRIWSSLVSACKLHG 538
G+ G+++ A E+ TM + + W+S++ +HG
Sbjct: 929 YGKCGRMDLAYEVFATMEDR-NVSSWTSMIVGYAMHG 1036
Score = 90.1 bits (222), Expect = 2e-18
Identities = 62/198 (31%), Positives = 92/198 (46%), Gaps = 1/198 (0%)
Frame = +2
Query: 53 LPSVIKACSSLHSHTLGTLLHGLALTTGSHSDPVVSNSLISLYAKFSDIQSARQVFDTMP 112
LP V+KA S + LG +H + G S+ + I+LY K D SA +VFD
Sbjct: 494 LPIVLKAVSQSFAIQLGQQVHSYGIKLGLQSNEYCESGFINLYCKAGDFDSAHKVFDENH 673
Query: 113 HRDPISWNSMINAYLQNGCLEEALEMLKGVYLLGLVPKPELLASMVSLCGRKMGSRIGRQ 172
SWN++I+ Q G +A+ + + G P + S++S CG + Q
Sbjct: 674 EPKLGSWNALISGLSQGGLAMDAIVVFVDMKRHGFEPDGITMVSVMSACGSIGDLYLALQ 853
Query: 173 IHALVVVDGRVEHSVVL-STALVDFYFRCDDALMASRVFDGMEVKNEVSWTAMISGCIAN 231
+H V E +V+L S +L+D Y +C +A VF ME +N SWT+MI G +
Sbjct: 854 LHKYVFQAKTNEWTVILMSNSLIDMYGKCGRMDLAYEVFATMEDRNVSSWTSMIVGYAMH 1033
Query: 232 LDYDAAFARFRAMQVEGV 249
A F M+ GV
Sbjct: 1034GHAKEALGCFHCMKGVGV 1087
Score = 86.7 bits (213), Expect(2) = 2e-20
Identities = 68/273 (24%), Positives = 131/273 (47%), Gaps = 7/273 (2%)
Frame = +2
Query: 81 SHSDPV-VSNSLISLYAKFSDIQS--ARQVFDTMPHRDPIS--WNSMINAYLQNGCLEEA 135
S +DPV V +L+S + D+ A + +P S WN++I +Y + + A
Sbjct: 260 SGNDPVTVIATLLSNTTRIRDLNQIYAHILLTRFLESNPASFNWNNIIRSYTRLESPQNA 439
Query: 136 LEMLKGVYLLGLVPKPELLASMVSLCGRKMGSRIGRQIHALVVVDGRVEHSVVLSTALVD 195
L + + G++P L ++ + ++G+Q+H+ + G ++ + + ++
Sbjct: 440 LRIYVSMLRAGVLPDRYTLPIVLKAVSQSFAIQLGQQVHSYGIKLG-LQSNEYCESGFIN 616
Query: 196 FYFRCDDALMASRVFDGMEVKNEVSWTAMISGCIANLDYDAAFARFRAMQVEGVDPNRVT 255
Y + D A +VFD SW A+ISG A F M+ G +P+ +T
Sbjct: 617 LYCKAGDFDSAHKVFDENHEPKLGSWNALISGLSQGGLAMDAIVVFVDMKRHGFEPDGIT 796
Query: 256 LIALLPACAKPGFVKHGKEIHGYAFRHGIE--SSHIFSSALINMYCQSGESLHLAEQIFE 313
+++++ AC G + ++H Y F+ + + S++LI+MY + G + LA ++F
Sbjct: 797 MVSVMSACGSIGDLYLALQLHKYVFQAKTNEWTVILMSNSLIDMYGKCGR-MDLAYEVFA 973
Query: 314 RSSFRDVVLWSSIIGSYSQRGDSYKALTLFNKM 346
R+V W+S+I Y+ G + +AL F+ M
Sbjct: 974 TMEDRNVSSWTSMIVGYAMHGHAKEALGCFHCM 1072
Score = 84.3 bits (207), Expect(2) = 3e-48
Identities = 38/78 (48%), Positives = 53/78 (67%)
Frame = +1
Query: 450 RGVKPDALTFLAVLSACNHAGLVTEGQQVFKQVNADYEIPLTIEHYACLVDLLGRSGKVE 509
RGVKP+ +TF+ VLSAC H G V EG+ F + Y I ++HY C+VDLLGR+G +
Sbjct: 1081 RGVKPNYVTFIGVLSACVHGGTVQEGRFYFDMMKNIYGITPQLQHYGCMVDLLGRAGLFD 1260
Query: 510 DALEIMRTMPMKPSTRIW 527
DA ++ MPMKP++ +W
Sbjct: 1261 DARRMVEEMPMKPNSVVW 1314
Score = 30.8 bits (68), Expect(2) = 2e-20
Identities = 16/75 (21%), Positives = 40/75 (53%), Gaps = 2/75 (2%)
Frame = +1
Query: 352 EPNYVTLLAVISACTNLSSLKHGCGLHGFILK-FGLSFSISLSNALMNMYAKCGCLEGSL 410
+PNYVT + V+SAC + +++ G + +G++ + ++++ + G + +
Sbjct: 1090 KPNYVTFIGVLSACVHGGTVQEGRFYFDMMKNIYGITPQLQHYGCMVDLLGRAGLFDDAR 1269
Query: 411 QIFLEMQNR-DSISW 424
++ EM + +S+ W
Sbjct: 1270 RMVEEMPMKPNSVVW 1314
>BF633384 similar to PIR|T46179|T46 hypothetical protein T8H10.30 -
Arabidopsis thaliana, partial (22%)
Length = 595
Score = 147 bits (371), Expect = 1e-35
Identities = 73/176 (41%), Positives = 113/176 (63%), Gaps = 6/176 (3%)
Frame = +1
Query: 422 ISWSTMISAYGLHGHGEQALKLFSEMKERG-----VKPDALTFLAVLSACNHAGLVTEGQ 476
I+W+ +I AYG+HG GE+ALKLF M E G ++P+ +T++A+ ++ +H+G+V EG
Sbjct: 1 ITWNVLIMAYGMHGKGEEALKLFRRMVEEGDNNREIRPNEVTYIAIFASLSHSGMVDEGL 180
Query: 477 QVFKQVNADYEIPLTIEHYACLVDLLGRSGKVEDALEIMRTMPMK-PSTRIWSSLVSACK 535
+F + A + I T +HYACLVDLLGRSG++E+A +++TMP WSSL+ ACK
Sbjct: 181 NLFYTMKAKHGIEPTSDHYACLVDLLGRSGQIEEAYNLIKTMPSNMKKVDAWSSLLGACK 360
Query: 536 LHGRLDIAEMLGPQLIRSEPDNAANYTLLNMIYAEHGHWLHKEQVMEAMKLQRLKK 591
+H L+I E+ L +P+ A+ Y LL+ IY+ G W V + MK + ++K
Sbjct: 361 IHQNLEIGEIAAKNLFVLDPNVASYYVLLSNIYSSAGLWDQAIDVRKKMKEKGVRK 528
>BE941714 weakly similar to GP|11994734|dbj selenium-binding protein-like
{Arabidopsis thaliana}, partial (10%)
Length = 619
Score = 146 bits (368), Expect = 3e-35
Identities = 77/203 (37%), Positives = 124/203 (60%), Gaps = 5/203 (2%)
Frame = +2
Query: 389 SISLSNALMNMYAKCGCLEGSLQIFLEMQNRDSISWSTMISAYGLHGHGEQALKLFSEMK 448
S+ LS +L++MYAKCG LE + ++F M RD + W+ MIS +HG G+ ALKLF +M+
Sbjct: 11 SVRLSTSLLDMYAKCGNLELAKRLFDSMNMRDVVCWNAMISGMAMHGDGKGALKLFYDME 190
Query: 449 ERGVKPDALTFLAVLSACNHAGLVTEGQQVFKQVNADYEIPLTIEHYACLVDLLGRSGKV 508
+ GVKPD +TF+AV +AC+++G+ EG + ++ + Y I EHY CLVDLL R+G
Sbjct: 191 KVGVKPDDITFIAVFTACSYSGMAYEGLMLLDKMCSVYNIVPKSEHYGCLVDLLSRAGLF 370
Query: 509 EDALEIMRTMPM----KPSTRIWSSLVSACKLHGRLDIAEMLGPQLIRSEPD-NAANYTL 563
E+A+ ++R + T W + +SAC HG +AE+ ++++ + ++ Y L
Sbjct: 371 EEAMVMIRKITNSWNGSEETLAWRAFLSACCNHGETQLAELAAEKVLQLDNHIHSGVYVL 550
Query: 564 LNMIYAEHGHWLHKEQVMEAMKL 586
L+ +YA G +V + MK+
Sbjct: 551 LSNLYAASGKHNDARRVKDMMKI 619
Score = 67.8 bits (164), Expect = 1e-11
Identities = 35/87 (40%), Positives = 50/87 (57%)
Frame = +2
Query: 186 SVVLSTALVDFYFRCDDALMASRVFDGMEVKNEVSWTAMISGCIANLDYDAAFARFRAMQ 245
SV LST+L+D Y +C + +A R+FD M +++ V W AMISG + D A F M+
Sbjct: 11 SVRLSTSLLDMYAKCGNLELAKRLFDSMNMRDVVCWNAMISGMAMHGDGKGALKLFYDME 190
Query: 246 VEGVDPNRVTLIALLPACAKPGFVKHG 272
GV P+ +T IA+ AC+ G G
Sbjct: 191 KVGVKPDDITFIAVFTACSYSGMAYEG 271
Score = 64.3 bits (155), Expect = 1e-10
Identities = 42/162 (25%), Positives = 82/162 (49%), Gaps = 12/162 (7%)
Frame = +2
Query: 291 SSALINMYCQSGESLHLAEQIFERSSFRDVVLWSSIIGSYSQRGDSYKALTLFNKMLTEE 350
S++L++MY + G +L LA+++F+ + RDVV W+++I + GD AL LF M
Sbjct: 23 STSLLDMYAKCG-NLELAKRLFDSMNMRDVVCWNAMISGMAMHGDGKGALKLFYDMEKVG 199
Query: 351 TEPNYVTLLAVISACT-------NLSSLKHGCGLHGFILKFGLSFSISLSNALMNMYAKC 403
+P+ +T +AV +AC+ L L C ++ + K L+++ ++
Sbjct: 200 VKPDDITFIAVFTACSYSGMAYEGLMLLDKMCSVYNIVPK------SEHYGCLVDLLSRA 361
Query: 404 GCLEGSLQIFLEMQN-----RDSISWSTMISAYGLHGHGEQA 440
G E ++ + ++ N ++++W +SA HG + A
Sbjct: 362 GLFEEAMVMIRKITNSWNGSEETLAWRAFLSACCNHGETQLA 487
Score = 50.1 bits (118), Expect = 2e-06
Identities = 37/152 (24%), Positives = 75/152 (49%), Gaps = 10/152 (6%)
Frame = +2
Query: 87 VSNSLISLYAKFSDIQSARQVFDTMPHRDPISWNSMINAYLQNGCLEEALEMLKGVYLLG 146
+S SL+ +YAK +++ A+++FD+M RD + WN+MI+ +G + AL++ + +G
Sbjct: 20 LSTSLLDMYAKCGNLELAKRLFDSMNMRDVVCWNAMISGMAMHGDGKGALKLFYDMEKVG 199
Query: 147 LVPKPELLASMVSLCGRKMGSRIGRQIHALVVVDGRVE-HSVVLST----ALVDFYFRC- 200
+ P ++ + C S G L+++D +++V + LVD R
Sbjct: 200 VKPDDITFIAVFTAC-----SYSGMAYEGLMLLDKMCSVYNIVPKSEHYGCLVDLLSRAG 364
Query: 201 ---DDALMASRVFDGMEVKNE-VSWTAMISGC 228
+ +M ++ + E ++W A +S C
Sbjct: 365 LFEEAMVMIRKITNSWNGSEETLAWRAFLSAC 460
>TC83482 weakly similar to GP|9758453|dbj|BAB08982.1 selenium-binding
protein-like {Arabidopsis thaliana}, partial (10%)
Length = 895
Score = 140 bits (352), Expect = 2e-33
Identities = 68/180 (37%), Positives = 108/180 (59%)
Frame = +2
Query: 420 DSISWSTMISAYGLHGHGEQALKLFSEMKERGVKPDALTFLAVLSACNHAGLVTEGQQVF 479
D +SW+ +I Y HG +AL+ F MK G PD +TF+ VLSAC HAGLV EG++ F
Sbjct: 2 DLVSWNAIIGGYASHGLATRALEEFDRMKVVGT-PDEVTFVNVLSACVHAGLVEEGEKHF 178
Query: 480 KQVNADYEIPLTIEHYACLVDLLGRSGKVEDALEIMRTMPMKPSTRIWSSLVSACKLHGR 539
+ Y I +EHY+C+VDL GR+G+ ++A +++ MP +P +W +L++AC LH
Sbjct: 179 TDMLTKYGIQAEMEHYSCMVDLYGRAGRFDEAENLIKNMPFEPDVVLWGALLAACGLHSN 358
Query: 540 LDIAEMLGPQLIRSEPDNAANYTLLNMIYAEHGHWLHKEQVMEAMKLQRLKKCYGFSRIE 599
L++ E ++ R E + +Y++L+ I E G W ++ + MK + +KK S +E
Sbjct: 359 LELGEYAAERIRRLESSHPVSYSVLSKIQGEKGVWSSVNELRDTMKERGIKKQTAISWVE 538
Score = 57.0 bits (136), Expect = 2e-08
Identities = 33/122 (27%), Positives = 65/122 (53%), Gaps = 2/122 (1%)
Frame = +2
Query: 319 DVVLWSSIIGSYSQRGDSYKALTLFNKMLTEETEPNYVTLLAVISACTNLSSLKHG-CGL 377
D+V W++IIG Y+ G + +AL F++M T P+ VT + V+SAC + ++ G
Sbjct: 2 DLVSWNAIIGGYASHGLATRALEEFDRMKVVGT-PDEVTFVNVLSACVHAGLVEEGEKHF 178
Query: 378 HGFILKFGLSFSISLSNALMNMYAKCGCLEGSLQIFLEMQ-NRDSISWSTMISAYGLHGH 436
+ K+G+ + + ++++Y + G + + + M D + W +++A GLH +
Sbjct: 179 TDMLTKYGIQAEMEHYSCMVDLYGRAGRFDEAENLIKNMPFEPDVVLWGALLAACGLHSN 358
Query: 437 GE 438
E
Sbjct: 359 LE 364
Score = 34.7 bits (78), Expect = 0.11
Identities = 29/126 (23%), Positives = 53/126 (42%), Gaps = 12/126 (9%)
Frame = +2
Query: 115 DPISWNSMINAYLQNGCLEEALEMLKGVYLLGLVPKPELLASMVSLCGRKMGSRIGRQIH 174
D +SWN++I Y +G ALE + ++G P +++S C +H
Sbjct: 2 DLVSWNAIIGGYASHGLATRALEEFDRMKVVG-TPDEVTFVNVLSAC-----------VH 145
Query: 175 ALVVVDGRVEHSVVLS-----------TALVDFYFRCDDALMASRVFDGMEVKNE-VSWT 222
A +V +G + +L+ + +VD Y R A + M + + V W
Sbjct: 146 AGLVEEGEKHFTDMLTKYGIQAEMEHYSCMVDLYGRAGRFDEAENLIKNMPFEPDVVLWG 325
Query: 223 AMISGC 228
A+++ C
Sbjct: 326 ALLAAC 343
>BE202961 weakly similar to GP|6016735|gb|A hypothetical protein {Arabidopsis
thaliana}, partial (14%)
Length = 613
Score = 129 bits (323), Expect = 4e-30
Identities = 65/196 (33%), Positives = 114/196 (58%), Gaps = 2/196 (1%)
Frame = +2
Query: 249 VDPNRVTLIALLPACAKPGFVKHGKEIHGYAFRHGIESSHIFSSALINMY--CQSGESLH 306
+ PN VT++++L ACA ++ G +IH A +G E S+AL++MY C S E
Sbjct: 35 IKPNWVTVVSVLRACACISNLEEGMKIHELAVNYGFEMETTVSTALMDMYMKCFSPEK-- 208
Query: 307 LAEQIFERSSFRDVVLWSSIIGSYSQRGDSYKALTLFNKMLTEETEPNYVTLLAVISACT 366
A IF R +DV+ W+ + Y+ G ++++ +F ML+ T P+ + L+ +++ +
Sbjct: 209 -AVDIFNRMPKKDVIAWAVLFSGYADNGMVHESMWVFRNMLSSGTRPDAIALVKILTTVS 385
Query: 367 NLSSLKHGCGLHGFILKFGLSFSISLSNALMNMYAKCGCLEGSLQIFLEMQNRDSISWST 426
L L+ H F++K G + + +L+ +YAKC +E + ++F M +D ++WS+
Sbjct: 386 ELGILQQAVCFHAFVIKNGFENNQFIGASLIEVYAKCSSIEDANKVFKGMTYKDVVTWSS 565
Query: 427 MISAYGLHGHGEQALK 442
+I+AYG HG GE+ALK
Sbjct: 566 IIAAYGFHGQGEEALK 613
Score = 105 bits (261), Expect = 7e-23
Identities = 57/201 (28%), Positives = 109/201 (53%), Gaps = 1/201 (0%)
Frame = +2
Query: 340 LTLFNKMLTEETEPNYVTLLAVISACTNLSSLKHGCGLHGFILKFGLSFSISLSNALMNM 399
L L ++ML + +PN+VT+++V+ AC +S+L+ G +H + +G ++S ALM+M
Sbjct: 2 LDLSSEMLDKRIKPNWVTVVSVLRACACISNLEEGMKIHELAVNYGFEMETTVSTALMDM 181
Query: 400 YAKCGCLEGSLQIFLEMQNRDSISWSTMISAYGLHGHGEQALKLFSEMKERGVKPDALTF 459
Y KC E ++ IF M +D I+W+ + S Y +G +++ +F M G +PDA+
Sbjct: 182 YMKCFSPEKAVDIFNRMPKKDVIAWAVLFSGYADNGMVHESMWVFRNMLSSGTRPDAIAL 361
Query: 460 LAVLSACNHAGLVTEGQQVFK-QVNADYEIPLTIEHYACLVDLLGRSGKVEDALEIMRTM 518
+ +L+ + G++ + + +E I A L+++ + +EDA ++ + M
Sbjct: 362 VKILTTVSELGILQQAVCFHAFVIKNGFENNQFIG--ASLIEVYAKCSSIEDANKVFKGM 535
Query: 519 PMKPSTRIWSSLVSACKLHGR 539
K WSS+++A HG+
Sbjct: 536 TYK-DVVTWSSIIAAYGFHGQ 595
Score = 91.3 bits (225), Expect = 1e-18
Identities = 53/187 (28%), Positives = 93/187 (49%)
Frame = +2
Query: 154 LASMVSLCGRKMGSRIGRQIHALVVVDGRVEHSVVLSTALVDFYFRCDDALMASRVFDGM 213
+ S++ C G +IH L V G E +STAL+D Y +C A +F+ M
Sbjct: 56 VVSVLRACACISNLEEGMKIHELAVNYG-FEMETTVSTALMDMYMKCFSPEKAVDIFNRM 232
Query: 214 EVKNEVSWTAMISGCIANLDYDAAFARFRAMQVEGVDPNRVTLIALLPACAKPGFVKHGK 273
K+ ++W + SG N + FR M G P+ + L+ +L ++ G ++
Sbjct: 233 PKKDVIAWAVLFSGYADNGMVHESMWVFRNMLSSGTRPDAIALVKILTTVSELGILQQAV 412
Query: 274 EIHGYAFRHGIESSHIFSSALINMYCQSGESLHLAEQIFERSSFRDVVLWSSIIGSYSQR 333
H + ++G E++ ++LI +Y + S+ A ++F+ +++DVV WSSII +Y
Sbjct: 413 CFHAFVIKNGFENNQFIGASLIEVYAKC-SSIEDANKVFKGMTYKDVVTWSSIIAAYGFH 589
Query: 334 GDSYKAL 340
G +AL
Sbjct: 590 GQGEEAL 610
Score = 58.9 bits (141), Expect = 5e-09
Identities = 32/109 (29%), Positives = 56/109 (51%)
Frame = +2
Query: 29 GLYHQTLELFTQLHFSAPHSISSVLPSVIKACSSLHSHTLGTLLHGLALTTGSHSDPVVS 88
G+ H+++ +F + S + L ++ S L H + G ++ +
Sbjct: 287 GMVHESMWVFRNMLSSGTRPDAIALVKILTTVSELGILQQAVCFHAFVIKNGFENNQFIG 466
Query: 89 NSLISLYAKFSDIQSARQVFDTMPHRDPISWNSMINAYLQNGCLEEALE 137
SLI +YAK S I+ A +VF M ++D ++W+S+I AY +G EEAL+
Sbjct: 467 ASLIEVYAKCSSIEDANKVFKGMTYKDVVTWSSIIAAYGFHGQGEEALK 613
>AW774626 similar to PIR|T47909|T47 hypothetical protein T20K12.70 -
Arabidopsis thaliana, partial (12%)
Length = 703
Score = 124 bits (310), Expect = 1e-28
Identities = 79/236 (33%), Positives = 128/236 (53%), Gaps = 2/236 (0%)
Frame = +3
Query: 193 LVDFYFRCDDALMASRVFDGMEV--KNEVSWTAMISGCIANLDYDAAFARFRAMQVEGVD 250
LVD Y +C A +F G+E KN V WTAM++G N D A FR M +GV+
Sbjct: 3 LVDMYAKCKCVSEAEFLFKGLEFDRKNHVLWTAMVTGYAQNGDGYKAVEFFRYMHAQGVE 182
Query: 251 PNRVTLIALLPACAKPGFVKHGKEIHGYAFRHGIESSHIFSSALINMYCQSGESLHLAEQ 310
N+ T +L AC+ G+++HG+ + G S+ SAL++MY + G+ L A+
Sbjct: 183 CNQYTFPTILTACSSVLARCFGEQVHGFIVKSGFGSNVYVQSALVDMYAKCGD-LKNAKN 359
Query: 311 IFERSSFRDVVLWSSIIGSYSQRGDSYKALTLFNKMLTEETEPNYVTLLAVISACTNLSS 370
+ E DVV W+S++ + + G +AL LF M + + T +V++ C + S
Sbjct: 360 MLETMEDDDVVSWNSLMVGFVRHGLEEEALRLFKNMHGRNMKIDDYTFPSVLNCCV-VGS 536
Query: 371 LKHGCGLHGFILKFGLSFSISLSNALMNMYAKCGCLEGSLQIFLEMQNRDSISWST 426
+ + +HG I+K G +SNAL++MYAK G ++ + +F +M +D ISW++
Sbjct: 537 I-NPKSVHGLIIKTGFENYKLVSNALVDMYAKTGDMDCAYTVFEKMLEKDVISWTS 701
Score = 117 bits (293), Expect = 1e-26
Identities = 72/247 (29%), Positives = 133/247 (53%), Gaps = 11/247 (4%)
Frame = +3
Query: 294 LINMYCQSGESLHLAEQIFERSSF--RDVVLWSSIIGSYSQRGDSYKALTLFNKMLTEET 351
L++MY + + + AE +F+ F ++ VLW++++ Y+Q GD YKA+ F M +
Sbjct: 3 LVDMYAKC-KCVSEAEFLFKGLEFDRKNHVLWTAMVTGYAQNGDGYKAVEFFRYMHAQGV 179
Query: 352 EPNYVTLLAVISACTNLSSLKHGCGLHGFILKFGLSFSISLSNALMNMYAKCGCLEGSLQ 411
E N T +++AC+++ + G +HGFI+K G ++ + +AL++MYAKCG L+ +
Sbjct: 180 ECNQYTFPTILTACSSVLARCFGEQVHGFIVKSGFGSNVYVQSALVDMYAKCGDLKNAKN 359
Query: 412 IFLEMQNRDSISWSTMISAYGLHGHGEQALKLFSEMKERGVKPDALTFLAVLSAC----- 466
+ M++ D +SW++++ + HG E+AL+LF M R +K D TF +VL+ C
Sbjct: 360 MLETMEDDDVVSWNSLMVGFVRHGLEEEALRLFKNMHGRNMKIDDYTFPSVLNCCVVGSI 539
Query: 467 ----NHAGLVTEGQQVFKQVNADYEIPLTIEHYACLVDLLGRSGKVEDALEIMRTMPMKP 522
H ++ G + +K V+ LVD+ ++G ++ A + M ++
Sbjct: 540 NPKSVHGLIIKTGFENYKLVS------------NALVDMYAKTGDMDCAYTVFEKM-LEK 680
Query: 523 STRIWSS 529
W+S
Sbjct: 681 DVISWTS 701
Score = 104 bits (259), Expect = 1e-22
Identities = 56/192 (29%), Positives = 106/192 (55%)
Frame = +3
Query: 32 HQTLELFTQLHFSAPHSISSVLPSVIKACSSLHSHTLGTLLHGLALTTGSHSDPVVSNSL 91
++ +E F +H P+++ ACSS+ + G +HG + +G S+ V ++L
Sbjct: 135 YKAVEFFRYMHAQGVECNQYTFPTILTACSSVLARCFGEQVHGFIVKSGFGSNVYVQSAL 314
Query: 92 ISLYAKFSDIQSARQVFDTMPHRDPISWNSMINAYLQNGCLEEALEMLKGVYLLGLVPKP 151
+ +YAK D+++A+ + +TM D +SWNS++ ++++G EEAL + K ++ +
Sbjct: 315 VDMYAKCGDLKNAKNMLETMEDDDVVSWNSLMVGFVRHGLEEEALRLFKNMHGRNMKIDD 494
Query: 152 ELLASMVSLCGRKMGSRIGRQIHALVVVDGRVEHSVVLSTALVDFYFRCDDALMASRVFD 211
S+++ C +GS + +H L++ G E+ ++S ALVD Y + D A VF+
Sbjct: 495 YTFPSVLNCC--VVGSINPKSVHGLIIKTG-FENYKLVSNALVDMYAKTGDMDCAYTVFE 665
Query: 212 GMEVKNEVSWTA 223
M K+ +SWT+
Sbjct: 666 KMLEKDVISWTS 701
Score = 100 bits (250), Expect = 1e-21
Identities = 60/237 (25%), Positives = 118/237 (49%), Gaps = 2/237 (0%)
Frame = +3
Query: 91 LISLYAKFSDIQSARQVFDTMP--HRDPISWNSMINAYLQNGCLEEALEMLKGVYLLGLV 148
L+ +YAK + A +F + ++ + W +M+ Y QNG +A+E + ++ G+
Sbjct: 3 LVDMYAKCKCVSEAEFLFKGLEFDRKNHVLWTAMVTGYAQNGDGYKAVEFFRYMHAQGVE 182
Query: 149 PKPELLASMVSLCGRKMGSRIGRQIHALVVVDGRVEHSVVLSTALVDFYFRCDDALMASR 208
++++ C + G Q+H +V G +V + +ALVD Y +C D A
Sbjct: 183 CNQYTFPTILTACSSVLARCFGEQVHGFIVKSG-FGSNVYVQSALVDMYAKCGDLKNAKN 359
Query: 209 VFDGMEVKNEVSWTAMISGCIANLDYDAAFARFRAMQVEGVDPNRVTLIALLPACAKPGF 268
+ + ME + VSW +++ G + + + A F+ M + + T ++L C
Sbjct: 360 MLETMEDDDVVSWNSLMVGFVRHGLEEEALRLFKNMHGRNMKIDDYTFPSVLNCCVVGSI 539
Query: 269 VKHGKEIHGYAFRHGIESSHIFSSALINMYCQSGESLHLAEQIFERSSFRDVVLWSS 325
+ K +HG + G E+ + S+AL++MY ++G+ + A +FE+ +DV+ W+S
Sbjct: 540 --NPKSVHGLIIKTGFENYKLVSNALVDMYAKTGD-MDCAYTVFEKMLEKDVISWTS 701
>AW776322 weakly similar to PIR|H84859|H84 hypothetical protein At2g42920
[imported] - Arabidopsis thaliana, partial (27%)
Length = 633
Score = 122 bits (305), Expect = 5e-28
Identities = 58/177 (32%), Positives = 107/177 (59%), Gaps = 1/177 (0%)
Frame = +1
Query: 424 WSTMISAYGLHGHGEQALKLFSEMKE-RGVKPDALTFLAVLSACNHAGLVTEGQQVFKQV 482
W+++I ++GH +A + FS+++ + +KPD+++F+ VL+AC H G + + + F+ +
Sbjct: 31 WNSIIIGLAMNGHEREAFEFFSKLESSKLLKPDSVSFIGVLTACKHLGAINKARDYFELM 210
Query: 483 NADYEIPLTIEHYACLVDLLGRSGKVEDALEIMRTMPMKPSTRIWSSLVSACKLHGRLDI 542
YEI +I+HY C+VD+LG++G +E+A E+++ MP+KP IW SL+S+C+ H + I
Sbjct: 211 MNKYEIEPSIKHYTCIVDVLGQAGLLEEAEELIKGMPLKPDAIIWGSLLSSCRKHRNVQI 390
Query: 543 AEMLGPQLIRSEPDNAANYTLLNMIYAEHGHWLHKEQVMEAMKLQRLKKCYGFSRIE 599
A ++ P +A+ Y L++ ++A + + MK +K G S IE
Sbjct: 391 ARRAAQRVYELNPSDASGYVLMSNVHAASNKFEEAIEQRLLMKENLTEKEPGCSSIE 561
Score = 30.8 bits (68), Expect = 1.6
Identities = 22/77 (28%), Positives = 38/77 (48%), Gaps = 5/77 (6%)
Frame = +1
Query: 101 IQSARQVFDTMPHRDPIS-----WNSMINAYLQNGCLEEALEMLKGVYLLGLVPKPELLA 155
I AR F+ M ++ I + +++ Q G LEEA E++KG + L P +
Sbjct: 178 INKARDYFELMMNKYEIEPSIKHYTCIVDVLGQAGLLEEAEELIKG---MPLKPDAIIWG 348
Query: 156 SMVSLCGRKMGSRIGRQ 172
S++S C + +I R+
Sbjct: 349 SLLSSCRKHRNVQIARR 399
>BF640739 weakly similar to GP|7363286|dbj Similar to Arabidopsis thaliana
DNA chromosome 4 BAC clone F4I10; vegetative storage
protein., partial (14%)
Length = 444
Score = 118 bits (295), Expect = 7e-27
Identities = 58/144 (40%), Positives = 94/144 (65%), Gaps = 1/144 (0%)
Frame = +1
Query: 338 KALTLFNKMLTEETEPNYVTLLAVISACTNLSSLKHGCGLHGFILKFGLSFSISLSNALM 397
+AL LF M ++ +P+ T ++VI+A +LS + +HG ++ + ++ ++ AL+
Sbjct: 7 EALNLFCTMQSQGIKPDSFTFVSVITALADLSVTRQAKWIHGLAIRTNMDTNVFVATALV 186
Query: 398 NMYAKCGCLEGSLQIFLEMQNRDSISWSTMISAYGLHGHGEQALKLFSEMK-ERGVKPDA 456
+MYAKCG +E + ++F MQ R I+W+ MI YG HG G+ AL LF +M+ E +KP+
Sbjct: 187 DMYAKCGAIETARELFDMMQERHVITWNAMIDGYGTHGLGKAALDLFDDMQNEASLKPND 366
Query: 457 LTFLAVLSACNHAGLVTEGQQVFK 480
+TFL+V+SAC+H+G V EG FK
Sbjct: 367 ITFLSVISACSHSGFVEEGLYYFK 438
Score = 82.4 bits (202), Expect = 5e-16
Identities = 43/135 (31%), Positives = 78/135 (56%), Gaps = 1/135 (0%)
Frame = +1
Query: 241 FRAMQVEGVDPNRVTLIALLPACAKPGFVKHGKEIHGYAFRHGIESSHIFSSALINMYCQ 300
F MQ +G+ P+ T ++++ A A + K IHG A R ++++ ++AL++MY +
Sbjct: 22 FCTMQSQGIKPDSFTFVSVITALADLSVTRQAKWIHGLAIRTNMDTNVFVATALVDMYAK 201
Query: 301 SGESLHLAEQIFERSSFRDVVLWSSIIGSYSQRGDSYKALTLFNKMLTEET-EPNYVTLL 359
G ++ A ++F+ R V+ W+++I Y G AL LF+ M E + +PN +T L
Sbjct: 202 CG-AIETARELFDMMQERHVITWNAMIDGYGTHGLGKAALDLFDDMQNEASLKPNDITFL 378
Query: 360 AVISACTNLSSLKHG 374
+VISAC++ ++ G
Sbjct: 379 SVISACSHSGFVEEG 423
Score = 65.9 bits (159), Expect = 4e-11
Identities = 40/131 (30%), Positives = 66/131 (49%), Gaps = 1/131 (0%)
Frame = +1
Query: 32 HQTLELFTQLHFSAPHSISSVLPSVIKACSSLHSHTLGTLLHGLALTTGSHSDPVVSNSL 91
++ L LF + S SVI A + L +HGLA+ T ++ V+ +L
Sbjct: 4 NEALNLFCTMQSQGIKPDSFTFVSVITALADLSVTRQAKWIHGLAIRTNMDTNVFVATAL 183
Query: 92 ISLYAKFSDIQSARQVFDTMPHRDPISWNSMINAYLQNGCLEEALEMLKGVY-LLGLVPK 150
+ +YAK I++AR++FD M R I+WN+MI+ Y +G + AL++ + L P
Sbjct: 184 VDMYAKCGAIETARELFDMMQERHVITWNAMIDGYGTHGLGKAALDLFDDMQNEASLKPN 363
Query: 151 PELLASMVSLC 161
S++S C
Sbjct: 364 DITFLSVISAC 396
Score = 65.5 bits (158), Expect = 6e-11
Identities = 42/142 (29%), Positives = 73/142 (50%), Gaps = 1/142 (0%)
Frame = +1
Query: 132 LEEALEMLKGVYLLGLVPKPELLASMVSLCGRKMGSRIGRQIHALVVVDGRVEHSVVLST 191
+ EAL + + G+ P S+++ +R + IH L + ++ +V ++T
Sbjct: 1 VNEALNLFCTMQSQGIKPDSFTFVSVITALADLSVTRQAKWIHGLAIRTN-MDTNVFVAT 177
Query: 192 ALVDFYFRCDDALMASRVFDGMEVKNEVSWTAMISGCIANLDYDAAFARFRAMQVE-GVD 250
ALVD Y +C A +FD M+ ++ ++W AMI G + AA F MQ E +
Sbjct: 178 ALVDMYAKCGAIETARELFDMMQERHVITWNAMIDGYGTHGLGKAALDLFDDMQNEASLK 357
Query: 251 PNRVTLIALLPACAKPGFVKHG 272
PN +T ++++ AC+ GFV+ G
Sbjct: 358 PNDITFLSVISACSHSGFVEEG 423
>TC80535 weakly similar to GP|14335136|gb|AAK59848.1 At2g15690/F9O13.24
{Arabidopsis thaliana}, partial (55%)
Length = 1727
Score = 113 bits (282), Expect = 2e-25
Identities = 63/229 (27%), Positives = 117/229 (50%)
Frame = +2
Query: 330 YSQRGDSYKALTLFNKMLTEETEPNYVTLLAVISACTNLSSLKHGCGLHGFILKFGLSFS 389
+ Q G +AL L K + + + N +L C S++ +H + L+
Sbjct: 218 FCQEGKVKEALELMEKGI--KADANCFEIL--FDLCGKSKSVEDAKKVHDYFLQSTFRSD 385
Query: 390 ISLSNALMNMYAKCGCLEGSLQIFLEMQNRDSISWSTMISAYGLHGHGEQALKLFSEMKE 449
+ N ++ MY C + + ++F M NR+ SW MI Y G++ L+LF +M E
Sbjct: 386 FKMHNKVIEMYGNCKSMTDARRVFDHMPNRNMDSWHMMIRGYANSTMGDEGLQLFEQMNE 565
Query: 450 RGVKPDALTFLAVLSACNHAGLVTEGQQVFKQVNADYEIPLTIEHYACLVDLLGRSGKVE 509
G++ + T LAVLSAC A V + + + + Y I +EHY L+D+LG+SG ++
Sbjct: 566 LGLEITSETMLAVLSACGSAEAVEDAYIYLESMKSKYGIEPGVEHYMGLLDVLGQSGYLK 745
Query: 510 DALEIMRTMPMKPSTRIWSSLVSACKLHGRLDIAEMLGPQLIRSEPDNA 558
+A E + +P +P+ ++ +L + ++HG +D+ + + ++ +P A
Sbjct: 746 EAEEFIEQLPFEPTVTVFETLKNYARIHGDVDLEDHVEELIVSLDPSKA 892
Score = 58.9 bits (141), Expect = 5e-09
Identities = 49/211 (23%), Positives = 87/211 (41%), Gaps = 1/211 (0%)
Frame = +2
Query: 123 INAYLQNGCLEEALEMLKGVYLLGLVPKPELLASMVSLCGRKMGSRIGRQIHALVVVDGR 182
+ + Q G ++EALE+++ G+ + LCG+ +++H +
Sbjct: 209 LTRFCQEGKVKEALELMEK----GIKADANCFEILFDLCGKSKSVEDAKKVHDYFL-QST 373
Query: 183 VEHSVVLSTALVDFYFRCDDALMASRVFDGMEVKNEVSWTAMISGCIANLDYDAAFARFR 242
+ +++ Y C A RVFD M +N SW MI G + D F
Sbjct: 374 FRSDFKMHNKVIEMYGNCKSMTDARRVFDHMPNRNMDSWHMMIRGYANSTMGDEGLQLFE 553
Query: 243 AMQVEGVDPNRVTLIALLPACAKPGFVKHGK-EIHGYAFRHGIESSHIFSSALINMYCQS 301
M G++ T++A+L AC V+ + ++GIE L+++ QS
Sbjct: 554 QMNELGLEITSETMLAVLSACGSAEAVEDAYIYLESMKSKYGIEPGVEHYMGLLDVLGQS 733
Query: 302 GESLHLAEQIFERSSFRDVVLWSSIIGSYSQ 332
G L AE+ E+ F V + +Y++
Sbjct: 734 G-YLKEAEEFIEQLPFEPTVTVFETLKNYAR 823
Score = 55.8 bits (133), Expect = 5e-08
Identities = 37/150 (24%), Positives = 66/150 (43%)
Frame = +2
Query: 13 SPTPSPSQQIKSLISQGLYHQTLELFTQLHFSAPHSISSVLPSVIKACSSLHSHTLGTLL 72
+P P + +G + LEL + + ++ + C S +
Sbjct: 182 APPPPSIVDLTRFCQEGKVKEALELMEK----GIKADANCFEILFDLCGKSKSVEDAKKV 349
Query: 73 HGLALTTGSHSDPVVSNSLISLYAKFSDIQSARQVFDTMPHRDPISWNSMINAYLQNGCL 132
H L + SD + N +I +Y + AR+VFD MP+R+ SW+ MI Y +
Sbjct: 350 HDYFLQSTFRSDFKMHNKVIEMYGNCKSMTDARRVFDHMPNRNMDSWHMMIRGYANSTMG 529
Query: 133 EEALEMLKGVYLLGLVPKPELLASMVSLCG 162
+E L++ + + LGL E + +++S CG
Sbjct: 530 DEGLQLFEQMNELGLEITSETMLAVLSACG 619
Score = 48.9 bits (115), Expect = 6e-06
Identities = 39/179 (21%), Positives = 81/179 (44%), Gaps = 2/179 (1%)
Frame = +2
Query: 259 LLPACAKPGFVKHGKEIHGYAFRHGIESSHIFSSALINMYCQSGESLHLAEQIFERSSFR 318
L C K V+ K++H Y + S + +I MY +S+ A ++F+ R
Sbjct: 299 LFDLCGKSKSVEDAKKVHDYFLQSTFRSDFKMHNKVIEMYGNC-KSMTDARRVFDHMPNR 475
Query: 319 DVVLWSSIIGSYSQRGDSYKALTLFNKMLTEETEPNYVTLLAVISACTNLSSLKHG-CGL 377
++ W +I Y+ + L LF +M E T+LAV+SAC + +++ L
Sbjct: 476 NMDSWHMMIRGYANSTMGDEGLQLFEQMNELGLEITSETMLAVLSACGSAEAVEDAYIYL 655
Query: 378 HGFILKFGLSFSISLSNALMNMYAKCGCLEGSLQIFLEMQNRDSIS-WSTMISAYGLHG 435
K+G+ + L+++ + G L+ + + ++ +++ + T+ + +HG
Sbjct: 656 ESMKSKYGIEPGVEHYMGLLDVLGQSGYLKEAEEFIEQLPFEPTVTVFETLKNYARIHG 832
>BG645370 weakly similar to PIR|T00405|T004 hypothetical protein At2g44880
[imported] - Arabidopsis thaliana, partial (11%)
Length = 784
Score = 111 bits (278), Expect = 7e-25
Identities = 55/133 (41%), Positives = 82/133 (61%)
Frame = -2
Query: 459 FLAVLSACNHAGLVTEGQQVFKQVNADYEIPLTIEHYACLVDLLGRSGKVEDALEIMRTM 518
F AVL+ACNHAG + +G++ F ++ +Y + I+ YACL+DL R+G + A ++M M
Sbjct: 783 FTAVLTACNHAGFIDKGEEYFNKMITNYGLSPDIDIYACLIDLYARNGNLRKARDLMEEM 604
Query: 519 PMKPSTRIWSSLVSACKLHGRLDIAEMLGPQLIRSEPDNAANYTLLNMIYAEHGHWLHKE 578
P P+ IWSS +SACK++G +++ QLI+ EP NAA Y L IY G W
Sbjct: 603 PYDPNCIIWSSFLSACKIYGDVELGREAAIQLIKMEPCNAAPYLTLAHIYTTKGLWNEAS 424
Query: 579 QVMEAMKLQRLKK 591
+V M+ QR+K+
Sbjct: 423 EVRSLMQ-QRVKR 388
Score = 35.0 bits (79), Expect = 0.083
Identities = 24/81 (29%), Positives = 43/81 (52%), Gaps = 2/81 (2%)
Frame = -2
Query: 360 AVISACTNLSSLKHGCG-LHGFILKFGLSFSISLSNALMNMYAKCGCLEGSLQIFLEM-Q 417
AV++AC + + G + I +GLS I + L+++YA+ G L + + EM
Sbjct: 777 AVLTACNHAGFIDKGEEYFNKMITNYGLSPDIDIYACLIDLYARNGNLRKARDLMEEMPY 598
Query: 418 NRDSISWSTMISAYGLHGHGE 438
+ + I WS+ +SA ++G E
Sbjct: 597 DPNCIIWSSFLSACKIYGDVE 535
Score = 29.6 bits (65), Expect = 3.5
Identities = 19/56 (33%), Positives = 34/56 (59%), Gaps = 6/56 (10%)
Frame = -2
Query: 91 LISLYAKFSDIQSARQVFDTMPHRDP--ISWNSMINAYLQNGCL----EEALEMLK 140
LI LYA+ +++ AR + + MP+ DP I W+S ++A G + E A++++K
Sbjct: 666 LIDLYARNGNLRKARDLMEEMPY-DPNCIIWSSFLSACKIYGDVELGREAAIQLIK 502
>BQ147515 weakly similar to PIR|T05355|T05 hypothetical protein F8B4.150 -
Arabidopsis thaliana, partial (23%)
Length = 427
Score = 104 bits (260), Expect = 9e-23
Identities = 48/142 (33%), Positives = 81/142 (56%)
Frame = +2
Query: 380 FILKFGLSFSISLSNALMNMYAKCGCLEGSLQIFLEMQNRDSISWSTMISAYGLHGHGEQ 439
++L+ +S N ++ MY +CG ++ ++ +F M RD + MI +G E
Sbjct: 2 YVLQHLSPLKVSTCNGILEMYFQCGSVDDAVNVFKNMTERDLTTICIMIKQLAKNGFAED 181
Query: 440 ALKLFSEMKERGVKPDALTFLAVLSACNHAGLVTEGQQVFKQVNADYEIPLTIEHYACLV 499
++ LF++ K G+KPD F+ V AC G + EG F+ ++ DY+I T+EHY LV
Sbjct: 182 SIDLFTQFKRSGLKPDGQMFIGVFGACXMLGDIVEGMLHFESMSRDYDIVPTMEHYVSLV 361
Query: 500 DLLGRSGKVEDALEIMRTMPMK 521
D++G G +++ALE + MPM+
Sbjct: 362 DMIGSIGHLDEALEFIEKMPME 427
Score = 28.5 bits (62), Expect = 7.8
Identities = 22/93 (23%), Positives = 37/93 (39%), Gaps = 2/93 (2%)
Frame = +2
Query: 22 IKSLISQGLYHQTLELFTQLHFSAPHSISSVLPSVIKACSSLHSHTLGTLLHGLALTTGS 81
IK L G +++LFTQ S + V AC L G +LH +++
Sbjct: 146 IKQLAKNGFAEDSIDLFTQFKRSGLKPDGQMFIGVFGACXMLGDIVEG-MLHFESMSRDY 322
Query: 82 HSDPVVSN--SLISLYAKFSDIQSARQVFDTMP 112
P + + SL+ + + A + + MP
Sbjct: 323 DIVPTMEHYVSLVDMIGSIGHLDEALEFIEKMP 421
>BF642497 weakly similar to PIR|H96713|H96 hypothetical protein T6L1.11
[imported] - Arabidopsis thaliana, partial (20%)
Length = 461
Score = 102 bits (254), Expect = 4e-22
Identities = 45/146 (30%), Positives = 93/146 (62%)
Frame = +3
Query: 321 VLWSSIIGSYSQRGDSYKALTLFNKMLTEETEPNYVTLLAVISACTNLSSLKHGCGLHGF 380
+ W+S+I ++Q G A+ +F +M E + + T +V++AC + +L+ G +H +
Sbjct: 6 ISWTSMITGFTQNGLDRDAIDIFREMKLENLQMDQYTFGSVLTACGGVMALQEGKQVHAY 185
Query: 381 ILKFGLSFSISLSNALMNMYAKCGCLEGSLQIFLEMQNRDSISWSTMISAYGLHGHGEQA 440
I++ +I +++AL++MY KC ++ + +F +M ++ +SW+ M+ YG +G+ E+A
Sbjct: 186 IIRTDYKDNIFVASALVDMYCKCKNIKSAEAVFKKMTCKNVVSWTAMLVGYGQNGYSEEA 365
Query: 441 LKLFSEMKERGVKPDALTFLAVLSAC 466
+K FS+M++ G++PD T +V+S+C
Sbjct: 366 VKTFSDMQKYGIEPDDFTLGSVISSC 443
Score = 97.4 bits (241), Expect = 1e-20
Identities = 55/154 (35%), Positives = 95/154 (60%), Gaps = 1/154 (0%)
Frame = +3
Query: 219 VSWTAMISGCIAN-LDYDAAFARFRAMQVEGVDPNRVTLIALLPACAKPGFVKHGKEIHG 277
+SWT+MI+G N LD DA FR M++E + ++ T ++L AC ++ GK++H
Sbjct: 6 ISWTSMITGFTQNGLDRDAIDI-FREMKLENLQMDQYTFGSVLTACGGVMALQEGKQVHA 182
Query: 278 YAFRHGIESSHIFSSALINMYCQSGESLHLAEQIFERSSFRDVVLWSSIIGSYSQRGDSY 337
Y R + + +SAL++MYC+ +++ AE +F++ + ++VV W++++ Y Q G S
Sbjct: 183 YIIRTDYKDNIFVASALVDMYCKC-KNIKSAEAVFKKMTCKNVVSWTAMLVGYGQNGYSE 359
Query: 338 KALTLFNKMLTEETEPNYVTLLAVISACTNLSSL 371
+A+ F+ M EP+ TL +VIS+C NL+SL
Sbjct: 360 EAVKTFSDMQKYGIEPDDFTLGSVISSCANLASL 461
Score = 89.7 bits (221), Expect = 3e-18
Identities = 50/148 (33%), Positives = 82/148 (54%)
Frame = +3
Query: 117 ISWNSMINAYLQNGCLEEALEMLKGVYLLGLVPKPELLASMVSLCGRKMGSRIGRQIHAL 176
ISW SMI + QNG +A+++ + + L L S+++ CG M + G+Q+HA
Sbjct: 6 ISWTSMITGFTQNGLDRDAIDIFREMKLENLQMDQYTFGSVLTACGGVMALQEGKQVHAY 185
Query: 177 VVVDGRVEHSVVLSTALVDFYFRCDDALMASRVFDGMEVKNEVSWTAMISGCIANLDYDA 236
++ + ++ +++ALVD Y +C + A VF M KN VSWTAM+ G N +
Sbjct: 186 IIRTD-YKDNIFVASALVDMYCKCKNIKSAEAVFKKMTCKNVVSWTAMLVGYGQNGYSEE 362
Query: 237 AFARFRAMQVEGVDPNRVTLIALLPACA 264
A F MQ G++P+ TL +++ +CA
Sbjct: 363 AVKTFSDMQKYGIEPDDFTLGSVISSCA 446
Score = 67.8 bits (164), Expect = 1e-11
Identities = 37/143 (25%), Positives = 67/143 (45%)
Frame = +3
Query: 19 SQQIKSLISQGLYHQTLELFTQLHFSAPHSISSVLPSVIKACSSLHSHTLGTLLHGLALT 78
+ I GL +++F ++ SV+ AC + + G +H +
Sbjct: 15 TSMITGFTQNGLDRDAIDIFREMKLENLQMDQYTFGSVLTACGGVMALQEGKQVHAYIIR 194
Query: 79 TGSHSDPVVSNSLISLYAKFSDIQSARQVFDTMPHRDPISWNSMINAYLQNGCLEEALEM 138
T + V+++L+ +Y K +I+SA VF M ++ +SW +M+ Y QNG EEA++
Sbjct: 195 TDYKDNIFVASALVDMYCKCKNIKSAEAVFKKMTCKNVVSWTAMLVGYGQNGYSEEAVKT 374
Query: 139 LKGVYLLGLVPKPELLASMVSLC 161
+ G+ P L S++S C
Sbjct: 375 FSDMQKYGIEPDDFTLGSVISSC 443
Score = 52.4 bits (124), Expect = 5e-07
Identities = 31/102 (30%), Positives = 55/102 (53%), Gaps = 1/102 (0%)
Frame = +3
Query: 421 SISWSTMISAYGLHGHGEQALKLFSEMKERGVKPDALTFLAVLSACNHAGLVTEGQQVFK 480
SISW++MI+ + +G A+ +F EMK ++ D TF +VL+AC + EG+QV
Sbjct: 3 SISWTSMITGFTQNGLDRDAIDIFREMKLENLQMDQYTFGSVLTACGGVMALQEGKQVHA 182
Query: 481 Q-VNADYEIPLTIEHYACLVDLLGRSGKVEDALEIMRTMPMK 521
+ DY+ + + + LVD+ + ++ A + + M K
Sbjct: 183 YIIRTDYKDNIFVA--SALVDMYCKCKNIKSAEAVFKKMTCK 302
>AW687696 weakly similar to PIR|T49965|T499 hypothetical protein F8M21.190 -
Arabidopsis thaliana, partial (16%)
Length = 289
Score = 101 bits (251), Expect = 9e-22
Identities = 43/92 (46%), Positives = 69/92 (74%)
Frame = +2
Query: 453 KPDALTFLAVLSACNHAGLVTEGQQVFKQVNADYEIPLTIEHYACLVDLLGRSGKVEDAL 512
+PD +TFL VL AC+H GLV EG++ F+ +N DY I TI+HY C+VDLLGR+G +A
Sbjct: 14 RPDEITFLCVLCACSHGGLVDEGRRYFEIMNRDYNIKPTIKHYGCMVDLLGRAGLFVEAY 193
Query: 513 EIMRTMPMKPSTRIWSSLVSACKLHGRLDIAE 544
E++++MP++ + IW +L++AC+ +G +++ E
Sbjct: 194 ELIKSMPVECNAIIWRTLLAACRNYGNVELGE 289
>TC83657 weakly similar to GP|11994382|dbj|BAB02341.
gb|AAB82628.1~gene_id:MIL23.2~similar to unknown protein
{Arabidopsis thaliana}, partial (10%)
Length = 1299
Score = 94.0 bits (232), Expect = 2e-19
Identities = 53/147 (36%), Positives = 85/147 (57%), Gaps = 4/147 (2%)
Frame = +3
Query: 460 LAVLSACNHAGLVTEGQQVFKQVNADYEIPLTIEHYACLVDLLGRSGKVEDALEIMRTMP 519
L VLSAC H GL++E +V ++ +Y I + I HY C+VDLLGR+GK+++A E+++ MP
Sbjct: 78 LXVLSACAHGGLMSEALEVISKME-EYGIEMGIRHYGCMVDLLGRAGKLKEAYELIKRMP 254
Query: 520 MKPSTRIWSSLVSACKLHGRLDIAEMLGPQLIRSEPDNAA----NYTLLNMIYAEHGHWL 575
MKP+ + +++ AC +H + +AE + + D+AA + LL+ IYA W
Sbjct: 255 MKPNETVLGAMIGACWIHSDMKMAEQVMKMI---GADSAACVNSHNVLLSNIYAASEKWE 425
Query: 576 HKEQVMEAMKLQRLKKCYGFSRIETGN 602
E + +M +K G+S I N
Sbjct: 426 KAEMIRSSMVDGGSEKIPGYSSIILSN 506
>AW257410 weakly similar to GP|10177533|db selenium-binding protein-like
{Arabidopsis thaliana}, partial (21%)
Length = 724
Score = 92.8 bits (229), Expect = 3e-19
Identities = 56/181 (30%), Positives = 95/181 (51%), Gaps = 1/181 (0%)
Frame = +1
Query: 273 KEIHGYAFRHGIESSHIFSSALINMYCQSGES-LHLAEQIFERSSFRDVVLWSSIIGSYS 331
K+I+ + G + S L+ Y S L A +F+R S + V+W+++I +YS
Sbjct: 28 KQIYVQFLKKGTIRHKLTVSRLLTTYASMEFSNLTYARMVFDRISSPNTVMWNTMIRAYS 207
Query: 332 QRGDSYKALTLFNKMLTEETEPNYVTLLAVISACTNLSSLKHGCGLHGFILKFGLSFSIS 391
D +AL L+++ML N T ++ AC+ LS+L +H I+K G +
Sbjct: 208 NSNDPEEALLLYHQMLHHSIPHNAYTFPFLLKACSALSALAETHQIHVQIIKRGFGSEVY 387
Query: 392 LSNALMNMYAKCGCLEGSLQIFLEMQNRDSISWSTMISAYGLHGHGEQALKLFSEMKERG 451
+N+L+ +YA G ++ + +F + +RD +SW+TMI Y G+ E A K+F M E+
Sbjct: 388 ATNSLLRVYAISGSIKSAHVLFDLLPSRDIVSWNTMIDGYIKCGNVEMAYKIFQAMPEKN 567
Query: 452 V 452
V
Sbjct: 568 V 570
Score = 71.6 bits (174), Expect = 8e-13
Identities = 40/113 (35%), Positives = 65/113 (57%)
Frame = +1
Query: 30 LYHQTLELFTQLHFSAPHSISSVLPSVIKACSSLHSHTLGTLLHGLALTTGSHSDPVVSN 89
LYHQ LH S PH+ + P ++KACS+L + +H + G S+ +N
Sbjct: 238 LYHQ------MLHHSIPHN-AYTFPFLLKACSALSALAETHQIHVQIIKRGFGSEVYATN 396
Query: 90 SLISLYAKFSDIQSARQVFDTMPHRDPISWNSMINAYLQNGCLEEALEMLKGV 142
SL+ +YA I+SA +FD +P RD +SWN+MI+ Y++ G +E A ++ + +
Sbjct: 397 SLLRVYAISGSIKSAHVLFDLLPSRDIVSWNTMIDGYIKCGNVEMAYKIFQAM 555
Score = 65.5 bits (158), Expect = 6e-11
Identities = 48/183 (26%), Positives = 80/183 (43%), Gaps = 3/183 (1%)
Frame = +1
Query: 167 SRIG--RQIHALVVVDGRVEHSVVLSTALVDFY-FRCDDALMASRVFDGMEVKNEVSWTA 223
S IG +QI+ + G + H + +S L + + A VFD + N V W
Sbjct: 10 SNIGELKQIYVQFLKKGTIRHKLTVSRLLTTYASMEFSNLTYARMVFDRISSPNTVMWNT 189
Query: 224 MISGCIANLDYDAAFARFRAMQVEGVDPNRVTLIALLPACAKPGFVKHGKEIHGYAFRHG 283
MI + D + A + M + N T LL AC+ + +IH + G
Sbjct: 190 MIRAYSNSNDPEEALLLYHQMLHHSIPHNAYTFPFLLKACSALSALAETHQIHVQIIKRG 369
Query: 284 IESSHIFSSALINMYCQSGESLHLAEQIFERSSFRDVVLWSSIIGSYSQRGDSYKALTLF 343
S +++L+ +Y SG S+ A +F+ RD+V W+++I Y + G+ A +F
Sbjct: 370 FGSEVYATNSLLRVYAISG-SIKSAHVLFDLLPSRDIVSWNTMIDGYIKCGNVEMAYKIF 546
Query: 344 NKM 346
M
Sbjct: 547 QAM 555
Score = 63.5 bits (153), Expect = 2e-10
Identities = 55/209 (26%), Positives = 89/209 (42%), Gaps = 32/209 (15%)
Frame = +1
Query: 77 LTTGSHSDPVVSNSLISLYA--KFSDIQSARQVFDTMPHRDPISWNSMINAYLQNGCLEE 134
L G+ + + L++ YA +FS++ AR VFD + + + WN+MI AY + EE
Sbjct: 49 LKKGTIRHKLTVSRLLTTYASMEFSNLTYARMVFDRISSPNTVMWNTMIRAYSNSNDPEE 228
Query: 135 AL----EMLKGVYLLGLVPKPELLASM-------------VSLCGRKMGSRI-------- 169
AL +ML P LL + V + R GS +
Sbjct: 229 ALLLYHQMLHHSIPHNAYTFPFLLKACSALSALAETHQIHVQIIKRGFGSEVYATNSLLR 408
Query: 170 -----GRQIHALVVVDGRVEHSVVLSTALVDFYFRCDDALMASRVFDGMEVKNEVSWTAM 224
G A V+ D +V ++D Y +C + MA ++F M KN +SWT+M
Sbjct: 409 VYAISGSIKSAHVLFDLLPSRDIVSWNTMIDGYIKCGNVEMAYKIFQAMPEKNVISWTSM 588
Query: 225 ISGCIANLDYDAAFARFRAMQVEGVDPNR 253
I G + + A + + M V + P++
Sbjct: 589 IVGFVRTGMHKKALSLLQQMLVARIKPDK 675
Score = 50.8 bits (120), Expect = 1e-06
Identities = 52/233 (22%), Positives = 108/233 (46%), Gaps = 7/233 (3%)
Frame = +1
Query: 365 CTNLSSLKHGCGLHGFILKFGLSFSISLSNALMNMYAKC--GCLEGSLQIFLEMQNRDSI 422
C+N+ LK ++ LK G + L+ YA L + +F + + +++
Sbjct: 7 CSNIGELKQ---IYVQFLKKGTIRHKLTVSRLLTTYASMEFSNLTYARMVFDRISSPNTV 177
Query: 423 SWSTMISAYGLHGHGEQALKLFSEMKERGVKPDALTFLAVLSACNHAGLVTEGQQVFKQV 482
W+TMI AY E+AL L+ +M + +A TF +L AC+ + E Q+ Q+
Sbjct: 178 MWNTMIRAYSNSNDPEEALLLYHQMLHHSIPHNAYTFPFLLKACSALSALAETHQIHVQI 357
Query: 483 NADYEIPLTIEHYA--CLVDLLGRSGKVEDALEIMRTMPMKPSTRIWSSLVSACKLHGRL 540
+ E YA L+ + SG ++ A + +P + W++++ G +
Sbjct: 358 ---IKRGFGSEVYATNSLLRVYAISGSIKSAHVLFDLLPSRDIVS-WNTMIDGYIKCGNV 525
Query: 541 DIAEMLGPQLIRSEPD-NAANYTLLNMIYAEHGHWLHKE--QVMEAMKLQRLK 590
++A ++ ++ P+ N ++T + + + G +HK+ +++ M + R+K
Sbjct: 526 EMAY----KIFQAMPEKNVISWTSMIVGFVRTG--MHKKALSLLQQMLVARIK 666
>TC84835 weakly similar to PIR|T02328|T02328 hypothetical protein At2g34400
[imported] - Arabidopsis thaliana, partial (5%)
Length = 647
Score = 84.7 bits (208), Expect = 9e-17
Identities = 48/185 (25%), Positives = 96/185 (50%), Gaps = 1/185 (0%)
Frame = +1
Query: 295 INMYCQSGESLHLAEQIFERSSFRDVVLWSSIIGSYSQRGDSYKALTLFNKMLTEETEPN 354
++++ + H + IF + + L++ +I S+ +R ++LFN++ P+
Sbjct: 25 VSIHLNNNNYFHYSLSIFNHTLHPSLFLYNLLIKSFFKRNSFQTLISLFNQLRLNGLYPD 204
Query: 355 YVTLLAVISACTNLSSLKHGCGLHGFILKFGLSFSISLSNALMNMYAKCGCLEGSLQIFL 414
T V+ A ++ + G +H F+ K GL +SN+ M+MYA+ G ++ ++F
Sbjct: 205 NYTYPFVLKAVAFIADFRQGTKIHAFVFKTGLDSDYYVSNSFMDMYAELGRIDFVRKLFD 384
Query: 415 EMQNRDSISWSTMISAYGLHGHGEQALKLFSEMK-ERGVKPDALTFLAVLSACNHAGLVT 473
E+ RDS+SW+ MIS E+A+++F M+ + K T ++ L+AC + V
Sbjct: 385 EISERDSVSWNVMISGCVKCRRFEEAVEVFQRMRVDSNEKISEATVVSSLTACAASRNVE 564
Query: 474 EGQQV 478
G+++
Sbjct: 565 VGKEI 579
Score = 80.9 bits (198), Expect = 1e-15
Identities = 45/158 (28%), Positives = 78/158 (48%), Gaps = 1/158 (0%)
Frame = +1
Query: 22 IKSLISQGLYHQTLELFTQLHFSAPHSISSVLPSVIKACSSLHSHTLGTLLHGLALTTGS 81
IKS + + + LF QL + + + P V+KA + + GT +H TG
Sbjct: 121 IKSFFKRNSFQTLISLFNQLRLNGLYPDNYTYPFVLKAVAFIADFRQGTKIHAFVFKTGL 300
Query: 82 HSDPVVSNSLISLYAKFSDIQSARQVFDTMPHRDPISWNSMINAYLQNGCLEEALEMLKG 141
SD VSNS + +YA+ I R++FD + RD +SWN MI+ ++ EEA+E+ +
Sbjct: 301 DSDYYVSNSFMDMYAELGRIDFVRKLFDEISERDSVSWNVMISGCVKCRRFEEAVEVFQR 480
Query: 142 VYLLGLVPKPE-LLASMVSLCGRKMGSRIGRQIHALVV 178
+ + E + S ++ C +G++IH ++
Sbjct: 481 MRVDSNEKISEATVVSSLTACAASRNVEVGKEIHGFII 594
Score = 70.1 bits (170), Expect = 2e-12
Identities = 44/173 (25%), Positives = 81/173 (46%), Gaps = 1/173 (0%)
Frame = +1
Query: 107 VFDTMPHRDPISWNSMINAYLQNGCLEEALEMLKGVYLLGLVPKPELLASMVSLCGRKMG 166
+F+ H +N +I ++ + + + + + L GL P ++
Sbjct: 73 IFNHTLHPSLFLYNLLIKSFFKRNSFQTLISLFNQLRLNGLYPDNYTYPFVLKAVAFIAD 252
Query: 167 SRIGRQIHALVVVDGRVEHSVVLSTALVDFYFRCDDALMASRVFDGMEVKNEVSWTAMIS 226
R G +IHA V G ++ +S + +D Y ++FD + ++ VSW MIS
Sbjct: 253 FRQGTKIHAFVFKTG-LDSDYYVSNSFMDMYAELGRIDFVRKLFDEISERDSVSWNVMIS 429
Query: 227 GCIANLDYDAAFARFRAMQVEGVDP-NRVTLIALLPACAKPGFVKHGKEIHGY 278
GC+ ++ A F+ M+V+ + + T+++ L ACA V+ GKEIHG+
Sbjct: 430 GCVKCRRFEEAVEVFQRMRVDSNEKISEATVVSSLTACAASRNVEVGKEIHGF 588
Score = 65.9 bits (159), Expect = 4e-11
Identities = 37/164 (22%), Positives = 83/164 (50%), Gaps = 1/164 (0%)
Frame = +1
Query: 221 WTAMISGCIANLDYDAAFARFRAMQVEGVDPNRVTLIALLPACAKPGFVKHGKEIHGYAF 280
+ +I + + F +++ G+ P+ T +L A A + G +IH + F
Sbjct: 109 YNLLIKSFFKRNSFQTLISLFNQLRLNGLYPDNYTYPFVLKAVAFIADFRQGTKIHAFVF 288
Query: 281 RHGIESSHIFSSALINMYCQSGESLHLAEQIFERSSFRDVVLWSSIIGSYSQRGDSYKAL 340
+ G++S + S++ ++MY + G + ++F+ S RD V W+ +I + +A+
Sbjct: 289 KTGLDSDYYVSNSFMDMYAELGR-IDFVRKLFDEISERDSVSWNVMISGCVKCRRFEEAV 465
Query: 341 TLFNKMLTEETEP-NYVTLLAVISACTNLSSLKHGCGLHGFILK 383
+F +M + E + T+++ ++AC +++ G +HGFI++
Sbjct: 466 EVFQRMRVDSNEKISEATVVSSLTACAASRNVEVGKEIHGFIIE 597
>BF635667 weakly similar to PIR|C85359|C853 hypothetical protein AT4g30700
[imported] - Arabidopsis thaliana, partial (6%)
Length = 603
Score = 83.6 bits (205), Expect = 2e-16
Identities = 46/137 (33%), Positives = 75/137 (54%), Gaps = 1/137 (0%)
Frame = +3
Query: 459 FLAVLSACNHAGLVTEGQQVFKQVNADYEIPLTIEHYACLVDLLGRSGKVEDALEIMRTM 518
F V++ C GL+ +G+++F + +DY + T EHYAC+VDL R G++E+AL+++ M
Sbjct: 6 F*LVVATC---GLLDQGKKIFGSMISDYGLEPTAEHYACMVDLFSRCGRLEEALQLLERM 176
Query: 519 PMKPST-RIWSSLVSACKLHGRLDIAEMLGPQLIRSEPDNAANYTLLNMIYAEHGHWLHK 577
T +W +L++ C +H + I E+ L + EP+N +NY L IY G L
Sbjct: 177 KSSSVTGSMWGALLAGCVMHQNVKIGEVAAHHLFQLEPNNTSNYVALWGIYQSRGMVLGV 356
Query: 578 EQVMEAMKLQRLKKCYG 594
+ M+ L K G
Sbjct: 357 STIXGKMRDLGLVKTPG 407
>AW585413 weakly similar to PIR|B85153|B851 hypothetical protein AT4g14050
[imported] - Arabidopsis thaliana, partial (5%)
Length = 577
Score = 77.4 bits (189), Expect = 1e-14
Identities = 51/160 (31%), Positives = 87/160 (53%), Gaps = 3/160 (1%)
Frame = +1
Query: 120 NSMINAYLQNGCLEEALEMLKGVYLLGLVPKPE--LLASMVSLCGRKMGSRIGRQIHALV 177
N I + ++ +ALE L + GL+ KP +L + +S C + + +G QIHA +
Sbjct: 94 NVYIRKHSKSASTCQALESLSRMN--GLIEKPTKYVLCNALSSCAKTLNWHLGIQIHAYM 267
Query: 178 VVDGRVEHSVVLSTALVDFYFRCDDALMASRVFDGMEVKNEVSWTAMISGCIANLDYDAA 237
+ G E ++ L +ALVDFY +C + A+++F M+ ++VSWT++I+G AN A
Sbjct: 268 IRSG-YEDNLFLCSALVDFYAKCFAIVDANKIFRAMKQHDQVSWTSLIAGFSANKQGRDA 444
Query: 238 FARFRAMQVEGVDPNRVTLIALLPAC-AKPGFVKHGKEIH 276
F+ M + PN TL +++ AC G ++H +H
Sbjct: 445 LLLFKEMLGTQIRPNCFTLTSVINACVXXNGVLEHCPTLH 564
Score = 77.4 bits (189), Expect = 1e-14
Identities = 38/140 (27%), Positives = 75/140 (53%)
Frame = +1
Query: 327 IGSYSQRGDSYKALTLFNKMLTEETEPNYVTLLAVISACTNLSSLKHGCGLHGFILKFGL 386
I +S+ + +AL ++M +P L +S+C + G +H ++++ G
Sbjct: 103 IRKHSKSASTCQALESLSRMNGLIEKPTKYVLCNALSSCAKTLNWHLGIQIHAYMIRSGY 282
Query: 387 SFSISLSNALMNMYAKCGCLEGSLQIFLEMQNRDSISWSTMISAYGLHGHGEQALKLFSE 446
++ L +AL++ YAKC + + +IF M+ D +SW+++I+ + + G AL LF E
Sbjct: 283 EDNLFLCSALVDFYAKCFAIVDANKIFRAMKQHDQVSWTSLIAGFSANKQGRDALLLFKE 462
Query: 447 MKERGVKPDALTFLAVLSAC 466
M ++P+ T +V++AC
Sbjct: 463 MLGTQIRPNCFTLTSVINAC 522
Score = 67.8 bits (164), Expect = 1e-11
Identities = 41/125 (32%), Positives = 59/125 (46%)
Frame = +1
Query: 251 PNRVTLIALLPACAKPGFVKHGKEIHGYAFRHGIESSHIFSSALINMYCQSGESLHLAEQ 310
P + L L +CAK G +IH Y R G E + SAL++ Y + + A +
Sbjct: 181 PTKYVLCNALSSCAKTLNWHLGIQIHAYMIRSGYEDNLFLCSALVDFYAKCFAIVD-ANK 357
Query: 311 IFERSSFRDVVLWSSIIGSYSQRGDSYKALTLFNKMLTEETEPNYVTLLAVISACTNLSS 370
IF D V W+S+I +S AL LF +ML + PN TL +VI+AC +
Sbjct: 358 IFRAMKQHDQVSWTSLIAGFSANKQGRDALLLFKEMLGTQIRPNCFTLTSVINACVXXNG 537
Query: 371 LKHGC 375
+ C
Sbjct: 538 VLEHC 552
Score = 52.8 bits (125), Expect = 4e-07
Identities = 31/129 (24%), Positives = 60/129 (46%)
Frame = +1
Query: 33 QTLELFTQLHFSAPHSISSVLPSVIKACSSLHSHTLGTLLHGLALTTGSHSDPVVSNSLI 92
Q LE ++++ VL + + +C+ + LG +H + +G + + ++L+
Sbjct: 136 QALESLSRMNGLIEKPTKYVLCNALSSCAKTLNWHLGIQIHAYMIRSGYEDNLFLCSALV 315
Query: 93 SLYAKFSDIQSARQVFDTMPHRDPISWNSMINAYLQNGCLEEALEMLKGVYLLGLVPKPE 152
YAK I A ++F M D +SW S+I + N +AL + K + + P
Sbjct: 316 DFYAKCFAIVDANKIFRAMKQHDQVSWTSLIAGFSANKQGRDALLLFKEMLGTQIRPNCF 495
Query: 153 LLASMVSLC 161
L S+++ C
Sbjct: 496 TLTSVINAC 522
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.322 0.135 0.401
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 19,298,569
Number of Sequences: 36976
Number of extensions: 278135
Number of successful extensions: 1719
Number of sequences better than 10.0: 123
Number of HSP's better than 10.0 without gapping: 1491
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1650
length of query: 605
length of database: 9,014,727
effective HSP length: 102
effective length of query: 503
effective length of database: 5,243,175
effective search space: 2637317025
effective search space used: 2637317025
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 61 (28.1 bits)
Lotus: description of TM0055.11


