

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0052a.5
(638 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
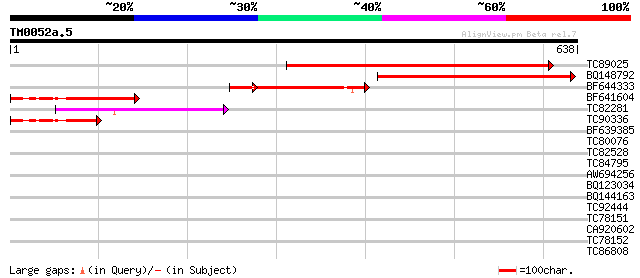
Score E
Sequences producing significant alignments: (bits) Value
TC89025 similar to GP|2462911|emb|CAB06081.1 UDP-glucose:sterol ... 401 e-112
BQ148792 similar to PIR|D96499|D96 probable UDP-glucose sterol g... 400 e-112
BF644333 similar to GP|2462911|emb UDP-glucose:sterol glucosyltr... 179 9e-50
BF641604 142 4e-34
TC82281 similar to GP|2462931|emb|CAB06082.1 UDP-glucose:sterol ... 129 3e-30
TC90336 79 5e-15
BF639385 weakly similar to GP|22830963|dbj putative UDP-glucose:... 41 0.002
TC80076 similar to GP|10177459|dbj|BAB10850. gene_id:MQB2.13~unk... 33 0.34
TC82528 weakly similar to PIR|C86356|C86356 hypothetical protein... 33 0.34
TC84795 weakly similar to GP|14532546|gb|AAK64001.1 At1g22370/T1... 32 0.75
AW694256 similar to PIR|E84908|E84 hypothetical protein At2g4689... 30 2.8
BQ123034 weakly similar to GP|14334982|gb putative glucosyltrans... 30 3.7
BQ144163 29 4.9
TC92444 homologue to GP|18254413|gb|AAL66754.1 putative copia-li... 29 4.9
TC78151 similar to GP|22655006|gb|AAM98094.1 At1g76400/F15M4_10 ... 29 6.3
CA920602 similar to GP|19911211|db glucosyltransferase-14 {Vigna... 29 6.3
TC78152 similar to PIR|T04482|T04482 ribophorin I homolog - barl... 29 6.3
TC86808 homologue to SP|Q9ZTR1|SPD1_PEA Spermidine synthase 1 (E... 28 8.3
>TC89025 similar to GP|2462911|emb|CAB06081.1 UDP-glucose:sterol
glucosyltransferase {Avena sativa}, partial (47%)
Length = 1157
Score = 401 bits (1031), Expect = e-112
Identities = 177/300 (59%), Positives = 226/300 (75%)
Frame = +1
Query: 312 VPLHIFFTMPWTPTYDFPHPLARVPQSAGYWMSYIIVDLMTWWGMRGIINDFRKKKLKLP 371
+P+HIFFTMPWTPT DFPHPL+RV Q AGY +SY IVD + W G+R +IND RKKKLKL
Sbjct: 4 IPIHIFFTMPWTPTADFPHPLSRVKQQAGYRLSYQIVDSLIWLGIRDMINDLRKKKLKLR 183
Query: 372 PIAYFSMYRGSISHLPTGYMWSPHVVPKPSDWGPLVDVVGYCFLSLASKYKPREDFVQWI 431
P+ Y S +G + +P Y+WSPH+VPKP DWGP +DVVG+CFL LAS Y+P E V+W+
Sbjct: 184 PVTYLSGSQGFENDIPHAYIWSPHLVPKPKDWGPKIDVVGFCFLDLASNYEPPESLVKWL 363
Query: 432 KKGPPPLYFGFGSMPLEDPTRTTDVILEALKDTEQRGIIDRGWGNLGSLAEVPDNVFLLE 491
+ G P+Y GFGS+P++DP + T +I+EAL+ T QRGII++GWG LG L E D+++LL+
Sbjct: 364 EDGDKPIYIGFGSLPVQDPKKMTQIIVEALETTGQRGIINKGWGGLGDLTEPKDSIYLLD 543
Query: 492 ECPHDWLFPQCSAVVHHGGAGTTATGLKAGCPTTIVPFFGDQFFWGDRIHEKELGPAPIP 551
PHDWLF QC AVVHHGGAGTTA GLKA CPTTIVPFFGDQ FWG+R+H++ +GP PIP
Sbjct: 544 NVPHDWLFLQCKAVVHHGGAGTTAAGLKAACPTTIVPFFGDQPFWGERVHDRGVGPPPIP 723
Query: 552 ISQLNVENLSNAIKFMLQPEVKSRVMLIAKLVENEDGVAAAVDAFHRHLPDELPLPTPSP 611
+ + ++ L +AI FML P+VK + +AK +ENEDGV AV AF + LP + P P
Sbjct: 724 VDEFSLPKLIDAINFMLDPKVKEHAIELAKAMENEDGVTGAVKAFFKQLPQKKPETNTEP 903
>BQ148792 similar to PIR|D96499|D96 probable UDP-glucose sterol
glucosyltransferase [imported] - Arabidopsis thaliana,
partial (31%)
Length = 701
Score = 400 bits (1028), Expect = e-112
Identities = 187/224 (83%), Positives = 202/224 (89%), Gaps = 1/224 (0%)
Frame = +3
Query: 414 FLSLASKYKPREDFVQWIKKGPPPLYFGFGSMPLEDPTRTTDVILEALKDTEQRGIIDRG 473
FL S Y+PREDF+ WIKKGPPPLYFGFGSMPLEDP TTDVIL+ALK+TEQRGIIDRG
Sbjct: 3 FLRHESNYQPREDFLHWIKKGPPPLYFGFGSMPLEDPKITTDVILKALKETEQRGIIDRG 182
Query: 474 WGNLGSLAEVPDNVFLLEECPHDWLFPQCSAVVHHGGAGTTATGLKAGCPTTIVPFFGDQ 533
WGNLG+L EV DNVFLLEECPHDWLFPQCSAVVHHGGAGTTATGLK+GCPTTIVPFFGDQ
Sbjct: 183 WGNLGNLTEVSDNVFLLEECPHDWLFPQCSAVVHHGGAGTTATGLKSGCPTTIVPFFGDQ 362
Query: 534 FFWGDRIHEKELGPAPIPISQLNVENLSNAIKFMLQPEVKSRVMLIAKLVENEDGVAAAV 593
FFWGDRIH+KELGPAPIPIS+LNVENLSNAIKFMLQPEVKSR M +AKL+E+EDGVAA V
Sbjct: 363 FFWGDRIHQKELGPAPIPISELNVENLSNAIKFMLQPEVKSRTMEVAKLIESEDGVAAXV 542
Query: 594 DAFHRHLPDELPLPTPSPQEE-DQINPLQWFFLQIGKWCCAPCG 636
DAFHRHLPDELPLPTPS E+ D ++PL WFF Q+ KWC P G
Sbjct: 543 DAFHRHLPDELPLPTPSHVEDXDHLSPLNWFFDQLAKWCXLPWG 674
>BF644333 similar to GP|2462911|emb UDP-glucose:sterol glucosyltransferase
{Avena sativa}, partial (23%)
Length = 473
Score = 179 bits (453), Expect(2) = 9e-50
Identities = 85/133 (63%), Positives = 99/133 (73%), Gaps = 3/133 (2%)
Frame = +3
Query: 275 LPACTAPDLETGVPFRAQAIIANPPAYGHAHVAEALGVPLHIFFTMPWTPTYDFPHPLAR 334
LPAC + E+ PF+A AIIANPPAYGH HVAE L VPLHIFFTMPWTPT DFPHPL+R
Sbjct: 84 LPACNSRYPESNEPFKADAIIANPPAYGHTHVAEYLNVPLHIFFTMPWTPTSDFPHPLSR 263
Query: 335 VPQSAGYWMSYIIVDLMTWWGMRGIINDFRKKKLKLPPIAYFSMYRGSIS---HLPTGYM 391
V Q GY +SY IVD + W G+R +IN+FRKKKLKL + Y RGS + +P GY+
Sbjct: 264 VRQPIGYRLSYQIVDALIWLGIRDLINEFRKKKLKLRAVTYL---RGSYTFPPDMPYGYI 434
Query: 392 WSPHVVPKPSDWG 404
WSPH+VPKP DWG
Sbjct: 435 WSPHLVPKPKDWG 473
Score = 37.0 bits (84), Expect(2) = 9e-50
Identities = 16/31 (51%), Positives = 23/31 (73%)
Frame = +2
Query: 248 RNKGLIPSGPGEVSIQLKQLKDIIDSLLPAC 278
+NKG +PSGP E+ +Q Q++ II S LP+C
Sbjct: 2 KNKGFLPSGPSEIHLQRSQIRAIIHS-LPSC 91
>BF641604
Length = 629
Score = 142 bits (358), Expect = 4e-34
Identities = 86/148 (58%), Positives = 105/148 (70%), Gaps = 2/148 (1%)
Frame = +3
Query: 1 MGSDGIDYLCKEVLEGGVDSQQKKGLWQMDGGDDKLVRV--DIKSQDGKFDLASSKEVED 58
MG +GI++LC EV+E +KGL +MD DDKLV V +I SQ+ SSKE
Sbjct: 237 MGCNGIEHLCTEVVE------DEKGLRKMD--DDKLVSVRGEIMSQEDDGS-PSSKE--- 380
Query: 59 WREKLGHKSSSLVTSSPRKGLGHCITAPVGTERNPLIADNEIILSRSMTDKRGLPRHDLI 118
SS VTS P +GL HCITAP+GTERNPLIA++EI++SRSMT+KRGL RHDL+
Sbjct: 381 -----RGSQSSEVTSLPHRGLEHCITAPIGTERNPLIAEDEIMISRSMTEKRGLRRHDLL 545
Query: 119 LDRLSEHEKEQLIAELVKIQNDGTVEVD 146
LDRLSE EK++LIA LV IQ DGTV+VD
Sbjct: 546 LDRLSEDEKQKLIANLVXIQKDGTVQVD 629
>TC82281 similar to GP|2462931|emb|CAB06082.1 UDP-glucose:sterol
glucosyltransferase {Arabidopsis thaliana}, partial
(18%)
Length = 799
Score = 129 bits (324), Expect = 3e-30
Identities = 77/201 (38%), Positives = 112/201 (55%), Gaps = 6/201 (2%)
Frame = +1
Query: 52 SSKEVEDWREKLGHKSSSLVTSSPRKGLGHCITAPVGTERNPLI--ADNEIILSRSMTDK 109
S+ V + + +SSS + P GL T PV I + ++ L RS T++
Sbjct: 199 STDSVSNGVSAINGESSSSTSGIPSTGLSKVTTLPVDISHGDKIESSPSKFKLERSKTER 378
Query: 110 RGLPRHD----LILDRLSEHEKEQLIAELVKIQNDGTVEVDMERSASVPSELLELQSFGE 165
+ R + + +++ EK +L+ + +++DGTVE D+ P L
Sbjct: 379 QRHLRPEDAAQIFNNKIPVQEKLRLLNRIATVKDDGTVEFDVPIDVE-PDALGAGSKHVN 555
Query: 166 PTVSGSLSSALKRSVPRLQIAILVVGTRGDVQPFLAIAKRLQEYGHRVRLATHSNFNTFV 225
+ S + +P L I +L+VGTRGDVQPF+AI KRLQ+YGHRVRL THSNF FV
Sbjct: 556 NVIDDSHGATDLDYIPPLNIVMLIVGTRGDVQPFVAIGKRLQDYGHRVRLXTHSNFKEFV 735
Query: 226 KSAGIDFYPLGGDPRILAGYM 246
+AG++FYPLGG P++LAGYM
Sbjct: 736 LTAGLEFYPLGGXPKVLAGYM 798
>TC90336
Length = 545
Score = 79.0 bits (193), Expect = 5e-15
Identities = 53/105 (50%), Positives = 67/105 (63%), Gaps = 2/105 (1%)
Frame = +1
Query: 1 MGSDGIDYLCKEVLEGGVDSQQKKGLWQMDGGDDKLVRV--DIKSQDGKFDLASSKEVED 58
MG +GI++LC EV+E +KGL +MD DDKLV V +I SQ+ SSKE
Sbjct: 280 MGCNGIEHLCTEVVE------DEKGLRKMD--DDKLVSVRGEIMSQEDDGS-PSSKE--- 423
Query: 59 WREKLGHKSSSLVTSSPRKGLGHCITAPVGTERNPLIADNEIILS 103
SS VTS P +GL HCITAP+GTERNPLIA++EI++S
Sbjct: 424 -----RGSQSSEVTSLPHRGLEHCITAPIGTERNPLIAEDEIMIS 543
>BF639385 weakly similar to GP|22830963|dbj putative UDP-glucose:sterol
glucosyltransferase {Oryza sativa (japonica
cultivar-group)}, partial (5%)
Length = 554
Score = 40.8 bits (94), Expect = 0.002
Identities = 35/136 (25%), Positives = 67/136 (48%), Gaps = 15/136 (11%)
Frame = +2
Query: 27 WQMDGGD----DKLVRVDIKSQDGKFDLASSKEVEDWREKLGHKSSSLVTSS-----PRK 77
W+ G D+ +RV+ + G + ++ +E+ D + + SS +SS PRK
Sbjct: 65 WRYSSGSSSSSDQSIRVE--HEIGTGNESTGQEIVDSTDAVSFGVSSCESSSSKSGIPRK 238
Query: 78 GLGHCITAPVGTERNPLI--ADNEIILSRSMTDKRGLPRHD----LILDRLSEHEKEQLI 131
GL T PV + + ++ L RS +D++ R + + D++ EK +L+
Sbjct: 239 GLHKATTMPVDISHGDKLESSPSKFKLERSKSDRQRHLRPEDAAQIFNDKIPVQEKLRLL 418
Query: 132 AELVKIQNDGTVEVDM 147
++ +++DGTVE D+
Sbjct: 419 NKIATVKDDGTVEFDV 466
>TC80076 similar to GP|10177459|dbj|BAB10850. gene_id:MQB2.13~unknown
protein {Arabidopsis thaliana}, partial (21%)
Length = 910
Score = 33.1 bits (74), Expect = 0.34
Identities = 20/62 (32%), Positives = 29/62 (46%), Gaps = 1/62 (1%)
Frame = +1
Query: 555 LNVENLSNAIKFMLQPEVK-SRVMLIAKLVENEDGVAAAVDAFHRHLPDELPLPTPSPQE 613
L E + NA+KF+ P+VK S VM +E + +D R +PD P S
Sbjct: 322 LREEQVQNAVKFLSHPKVKGSPVMYRRSFLEKKGLTKEEIDEAFRRVPDSAPTVQTSGVT 501
Query: 614 ED 615
+D
Sbjct: 502 QD 507
>TC82528 weakly similar to PIR|C86356|C86356 hypothetical protein AAF87257.1
[imported] - Arabidopsis thaliana, partial (35%)
Length = 1376
Score = 33.1 bits (74), Expect = 0.34
Identities = 17/54 (31%), Positives = 24/54 (43%), Gaps = 2/54 (3%)
Frame = +1
Query: 482 EVPDNVFLLEECPHDWLF--PQCSAVVHHGGAGTTATGLKAGCPTTIVPFFGDQ 533
E+ D + CP + + P + H G +T + AG P PFFGDQ
Sbjct: 871 EISDRGLITSWCPQEQVLIHPSIGGFLTHCGWNSTTESICAGVPMLCWPFFGDQ 1032
>TC84795 weakly similar to GP|14532546|gb|AAK64001.1 At1g22370/T16E15_3
{Arabidopsis thaliana}, partial (33%)
Length = 605
Score = 32.0 bits (71), Expect = 0.75
Identities = 16/54 (29%), Positives = 24/54 (43%), Gaps = 2/54 (3%)
Frame = +1
Query: 482 EVPDNVFLLEECPHDWLF--PQCSAVVHHGGAGTTATGLKAGCPTTIVPFFGDQ 533
E+ D + CP + + P + H G +T + AG P PF+GDQ
Sbjct: 43 EISDRGLIASWCPQEQVLNHPSIGGFLTHCGWNSTVESVLAGVPMLCWPFYGDQ 204
>AW694256 similar to PIR|E84908|E84 hypothetical protein At2g46890 [imported]
- Arabidopsis thaliana, partial (34%)
Length = 435
Score = 30.0 bits (66), Expect = 2.8
Identities = 18/43 (41%), Positives = 25/43 (57%), Gaps = 1/43 (2%)
Frame = +1
Query: 326 YDFPHPLARVPQSAGYWMSYIIVDLMTW-WGMRGIINDFRKKK 367
Y HPLA +W S I++ L+TW W +R I N FR++K
Sbjct: 247 YYAAHPLAHYD----FWRSRIVI-LLTWVWSIRLIHNYFRREK 360
>BQ123034 weakly similar to GP|14334982|gb putative glucosyltransferase
{Arabidopsis thaliana}, partial (24%)
Length = 765
Score = 29.6 bits (65), Expect = 3.7
Identities = 13/36 (36%), Positives = 17/36 (47%)
Frame = +2
Query: 500 PQCSAVVHHGGAGTTATGLKAGCPTTIVPFFGDQFF 535
P +V H G +T L AG P P F +QF+
Sbjct: 341 PATGGIVTHCGWNSTLESLNAGLPMITWPIFAEQFY 448
>BQ144163
Length = 198
Score = 29.3 bits (64), Expect = 4.9
Identities = 14/36 (38%), Positives = 20/36 (54%), Gaps = 1/36 (2%)
Frame = -3
Query: 416 SLASKYKPREDFVQW-IKKGPPPLYFGFGSMPLEDP 450
S + P+ ++ W +KGP PL FG +PLE P
Sbjct: 115 SYSGHLAPKNTWLGWGFRKGPFPLKTVFGPLPLESP 8
>TC92444 homologue to GP|18254413|gb|AAL66754.1 putative copia-like
retrotransposon Hopscotch polyprotein {Zea mays},
partial (1%)
Length = 917
Score = 29.3 bits (64), Expect = 4.9
Identities = 24/77 (31%), Positives = 36/77 (46%), Gaps = 1/77 (1%)
Frame = +2
Query: 93 PLIADNEIILSRSMTDKRGLPRHDLILDRLSEHEKEQLIAELVKIQN-DGTVEVDMERSA 151
P +A + +SR +TD RGL E + + E+VK Q + VE + E
Sbjct: 536 PWLAATSLSMSRELTDLRGLT------------EPREAVTEMVKDQEAEAAVETEEE--- 670
Query: 152 SVPSELLELQSFGEPTV 168
VP E++ LQ + TV
Sbjct: 671 EVPQEMV-LQEMAQETV 718
>TC78151 similar to GP|22655006|gb|AAM98094.1 At1g76400/F15M4_10
{Arabidopsis thaliana}, partial (88%)
Length = 2171
Score = 28.9 bits (63), Expect = 6.3
Identities = 12/30 (40%), Positives = 17/30 (56%)
Frame = +1
Query: 363 FRKKKLKLPPIAYFSMYRGSISHLPTGYMW 392
FR+ KLPP A+ YR I ++ T +W
Sbjct: 901 FRRLTAKLPPRAHSVYYRDEIGNISTSSLW 990
>CA920602 similar to GP|19911211|db glucosyltransferase-14 {Vigna angularis},
partial (35%)
Length = 817
Score = 28.9 bits (63), Expect = 6.3
Identities = 13/28 (46%), Positives = 15/28 (53%)
Frame = -3
Query: 508 HGGAGTTATGLKAGCPTTIVPFFGDQFF 535
H G +T + AG P P FGDQFF
Sbjct: 599 HCGWNSTLEAICAGVPMITWPLFGDQFF 516
>TC78152 similar to PIR|T04482|T04482 ribophorin I homolog - barley
(fragment), partial (31%)
Length = 680
Score = 28.9 bits (63), Expect = 6.3
Identities = 12/30 (40%), Positives = 17/30 (56%)
Frame = +3
Query: 363 FRKKKLKLPPIAYFSMYRGSISHLPTGYMW 392
FR+ KLPP A+ YR I ++ T +W
Sbjct: 171 FRRLTAKLPPRAHSVYYRDEIGNISTSSLW 260
>TC86808 homologue to SP|Q9ZTR1|SPD1_PEA Spermidine synthase 1 (EC 2.5.1.16)
(Putrescine aminopropyltransferase 1) (SPDSY 1). [Garden
pea], partial (93%)
Length = 1350
Score = 28.5 bits (62), Expect = 8.3
Identities = 11/17 (64%), Positives = 13/17 (75%)
Frame = +3
Query: 65 HKSSSLVTSSPRKGLGH 81
H SSSL+ S P+KG GH
Sbjct: 417 HSSSSLLNSKPQKGFGH 467
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.320 0.139 0.436
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 21,728,794
Number of Sequences: 36976
Number of extensions: 342476
Number of successful extensions: 1610
Number of sequences better than 10.0: 36
Number of HSP's better than 10.0 without gapping: 1570
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1600
length of query: 638
length of database: 9,014,727
effective HSP length: 102
effective length of query: 536
effective length of database: 5,243,175
effective search space: 2810341800
effective search space used: 2810341800
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 61 (28.1 bits)
Lotus: description of TM0052a.5


