

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0052a.2
(68 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
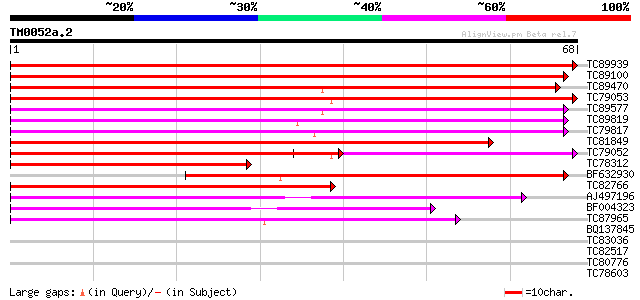
Score E
Sequences producing significant alignments: (bits) Value
TC89939 similar to GP|15289911|dbj|BAB63606. hypothetical protei... 121 6e-29
TC89100 similar to GP|21553849|gb|AAM62942.1 putative RING zinc ... 110 8e-26
TC89470 similar to PIR|T52142|T52142 RING finger protein RMA1 [v... 89 2e-19
TC79053 similar to GP|22795037|gb|AAN05420.1 putative RING prote... 86 3e-18
TC89577 similar to GP|22795037|gb|AAN05420.1 putative RING prote... 82 4e-17
TC89819 weakly similar to GP|22795037|gb|AAN05420.1 putative RIN... 82 5e-17
TC79817 similar to GP|22795037|gb|AAN05420.1 putative RING prote... 79 3e-16
TC81849 similar to GP|19698885|gb|AAL91178.1 putative protein {A... 77 1e-15
TC79052 similar to GP|22795037|gb|AAN05420.1 putative RING prote... 61 8e-14
TC78312 similar to GP|19698885|gb|AAL91178.1 putative protein {A... 56 3e-09
BF632930 weakly similar to PIR|T52142|T521 RING finger protein R... 55 5e-09
TC82766 similar to GP|3128183|gb|AAC16087.1| unknown protein {Ar... 53 2e-08
AJ497196 weakly similar to SP|O60683|PEXA_ Peroxisome assembly p... 47 1e-06
BF004323 similar to GP|4337011|gb| zinc-binding peroxisomal inte... 40 1e-04
TC87965 similar to GP|11994306|dbj|BAB01736. emb|CAA66822.1~gene... 38 9e-04
BQ137845 similar to GP|22795037|gb| putative RING protein {Popul... 35 0.007
TC83036 similar to PIR|C96637|C96637 hypothetical protein F11P17... 34 0.010
TC82517 homologue to GP|7688065|emb|CAB89694.1 constitutively ph... 33 0.016
TC80776 similar to GP|13605815|gb|AAK32893.1 At2g32700/F24L7.16 ... 33 0.021
TC78603 weakly similar to GP|19698827|gb|AAL91149.1 unknown prot... 33 0.028
>TC89939 similar to GP|15289911|dbj|BAB63606. hypothetical protein~similar
to Arabidopsis thaliana chromosome 1 F18O14.3, partial
(23%)
Length = 978
Score = 121 bits (303), Expect = 6e-29
Identities = 50/68 (73%), Positives = 59/68 (86%)
Frame = +2
Query: 1 CNICFDLAQDPVITLCGYLFRWPCLYKWLHFHSLSQQCPVCNALAEEDKLVPLHGRGNTS 60
CNICFDLAQDP+ITLCG+LF WPCLYKWLHFHS S++CPVC A+ EE+KLVPL+GRG TS
Sbjct: 146 CNICFDLAQDPIITLCGHLFCWPCLYKWLHFHSQSRECPVCKAMIEEEKLVPLYGRGKTS 325
Query: 61 SSSSSSSI 68
+ S S+
Sbjct: 326 TDPRSKSV 349
>TC89100 similar to GP|21553849|gb|AAM62942.1 putative RING zinc finger
protein {Arabidopsis thaliana}, partial (44%)
Length = 1047
Score = 110 bits (276), Expect = 8e-26
Identities = 47/67 (70%), Positives = 55/67 (81%)
Frame = +1
Query: 1 CNICFDLAQDPVITLCGYLFRWPCLYKWLHFHSLSQQCPVCNALAEEDKLVPLHGRGNTS 60
CNICFDLAQDPVITLCG+LF WPCLY+WLH HS SQ+ PVC AL +E+KLVPL+GRG T
Sbjct: 361 CNICFDLAQDPVITLCGHLFCWPCLYRWLHHHSHSQEFPVCKALVQEEKLVPLYGRGKTQ 540
Query: 61 SSSSSSS 67
+ + S
Sbjct: 541 TDPRTKS 561
>TC89470 similar to PIR|T52142|T52142 RING finger protein RMA1 [validated] -
Arabidopsis thaliana, partial (39%)
Length = 1031
Score = 89.4 bits (220), Expect = 2e-19
Identities = 37/73 (50%), Positives = 50/73 (67%), Gaps = 7/73 (9%)
Frame = +1
Query: 1 CNICFDLAQDPVITLCGYLFRWPCLYKWLHFHSLSQ-------QCPVCNALAEEDKLVPL 53
CNIC + QDPV+TLCG+L+ WPC+YKWL+FH+ +Q QCPVC + + LVPL
Sbjct: 292 CNICLECVQDPVVTLCGHLYCWPCIYKWLNFHAENQEKQKEEPQCPVCKSEISKSSLVPL 471
Query: 54 HGRGNTSSSSSSS 66
+GRG T+ S +
Sbjct: 472 YGRGQTTPPSKGN 510
>TC79053 similar to GP|22795037|gb|AAN05420.1 putative RING protein {Populus
x canescens}, partial (28%)
Length = 1139
Score = 85.9 bits (211), Expect = 3e-18
Identities = 36/74 (48%), Positives = 50/74 (66%), Gaps = 6/74 (8%)
Frame = +2
Query: 1 CNICFDLAQDPVITLCGYLFRWPCLYKWLHFHSLSQQ------CPVCNALAEEDKLVPLH 54
CNIC + A DPV+TLCG+L+ WPC+YKWL+ S S + CPVC A+ LVPL+
Sbjct: 254 CNICLESANDPVVTLCGHLYCWPCIYKWLNVQSSSVEPDTQPTCPVCKAVISHTSLVPLY 433
Query: 55 GRGNTSSSSSSSSI 68
GRG ++S + S+ +
Sbjct: 434 GRGKSNSETESNKL 475
>TC89577 similar to GP|22795037|gb|AAN05420.1 putative RING protein {Populus
x canescens}, partial (56%)
Length = 954
Score = 82.0 bits (201), Expect = 4e-17
Identities = 36/74 (48%), Positives = 44/74 (58%), Gaps = 7/74 (9%)
Frame = +1
Query: 1 CNICFDLAQDPVITLCGYLFRWPCLYKWLHFHSLSQ-------QCPVCNALAEEDKLVPL 53
CNIC D A +PV+TLCG+L+ WPC+YKWLH S QCPVC +VPL
Sbjct: 205 CNICLDFANEPVVTLCGHLYCWPCIYKWLHVQSDDSLGSDEHPQCPVCKENISHTTMVPL 384
Query: 54 HGRGNTSSSSSSSS 67
+GRG + S SS
Sbjct: 385 YGRGGHTPPSGKSS 426
>TC89819 weakly similar to GP|22795037|gb|AAN05420.1 putative RING protein
{Populus x canescens}, partial (33%)
Length = 880
Score = 81.6 bits (200), Expect = 5e-17
Identities = 35/75 (46%), Positives = 44/75 (58%), Gaps = 8/75 (10%)
Frame = +2
Query: 1 CNICFDLAQDPVITLCGYLFRWPCLYKWLHFHS--------LSQQCPVCNALAEEDKLVP 52
CNIC D QDPV+T CG+L+ WPC+YKWL S QCPVC + + LVP
Sbjct: 191 CNICLDCVQDPVVTFCGHLYCWPCIYKWLDIQSGISSENEKQKPQCPVCKSELSQSSLVP 370
Query: 53 LHGRGNTSSSSSSSS 67
L+GRG T + S +
Sbjct: 371 LYGRGQTMTLSEGKA 415
>TC79817 similar to GP|22795037|gb|AAN05420.1 putative RING protein {Populus
x canescens}, partial (32%)
Length = 1228
Score = 79.3 bits (194), Expect = 3e-16
Identities = 36/73 (49%), Positives = 43/73 (58%), Gaps = 6/73 (8%)
Frame = +1
Query: 1 CNICFDLAQDPVITLCGYLFRWPCLYKWLHFHSLS------QQCPVCNALAEEDKLVPLH 54
CNIC D A +PV+TLCG+L+ W C+YKWL S S QCPVC K+VPL+
Sbjct: 325 CNICLDFAHEPVVTLCGHLYCWSCIYKWLFVQSASLAPDEPPQCPVCKDGISHTKMVPLY 504
Query: 55 GRGNTSSSSSSSS 67
GRG T S S
Sbjct: 505 GRGQTLSRCDRDS 543
>TC81849 similar to GP|19698885|gb|AAL91178.1 putative protein {Arabidopsis
thaliana}, partial (16%)
Length = 1477
Score = 77.4 bits (189), Expect = 1e-15
Identities = 28/58 (48%), Positives = 42/58 (72%)
Frame = +1
Query: 1 CNICFDLAQDPVITLCGYLFRWPCLYKWLHFHSLSQQCPVCNALAEEDKLVPLHGRGN 58
CNIC D+A+DPV+T CG+LF WPC Y+ + +S +++CPVC E ++P++G GN
Sbjct: 160 CNICLDIARDPVLTCCGHLFCWPCFYQLSYAYSKAKECPVCKGEVTESGIIPIYGHGN 333
>TC79052 similar to GP|22795037|gb|AAN05420.1 putative RING protein {Populus
x canescens}, partial (28%)
Length = 503
Score = 61.2 bits (147), Expect(2) = 8e-14
Identities = 25/46 (54%), Positives = 32/46 (69%), Gaps = 6/46 (13%)
Frame = +3
Query: 1 CNICFDLAQDPVITLCGYLFRWPCLYKWLHFHSLSQQ------CPV 40
CNIC + A DPV+TLCG+L+ WPC+YKWL+ S S + CPV
Sbjct: 201 CNICLESANDPVVTLCGHLYCWPCIYKWLNVQSSSVEPDTQPTCPV 338
Score = 30.0 bits (66), Expect(2) = 8e-14
Identities = 12/34 (35%), Positives = 20/34 (58%)
Frame = +1
Query: 35 SQQCPVCNALAEEDKLVPLHGRGNTSSSSSSSSI 68
S+ C A+ LVPL+GRG ++S + S+ +
Sbjct: 322 SRHARFCKAVISHTSLVPLYGRGKSNSETESNKL 423
>TC78312 similar to GP|19698885|gb|AAL91178.1 putative protein {Arabidopsis
thaliana}, partial (14%)
Length = 1317
Score = 55.8 bits (133), Expect = 3e-09
Identities = 20/29 (68%), Positives = 25/29 (85%)
Frame = +3
Query: 1 CNICFDLAQDPVITLCGYLFRWPCLYKWL 29
CNIC DLA++PV+T CG+LF W CLY+WL
Sbjct: 1230 CNICLDLAKEPVLTCCGHLFCWQCLYRWL 1316
>BF632930 weakly similar to PIR|T52142|T521 RING finger protein RMA1
[validated] - Arabidopsis thaliana, partial (18%)
Length = 393
Score = 55.1 bits (131), Expect = 5e-09
Identities = 22/50 (44%), Positives = 32/50 (64%), Gaps = 4/50 (8%)
Frame = +1
Query: 22 WPCLYKWLHF----HSLSQQCPVCNALAEEDKLVPLHGRGNTSSSSSSSS 67
WPC+YKW++ H+ +CP+C + E LVPL+GRG T+SSS +
Sbjct: 4 WPCIYKWINSTSWEHNEKPECPICKSEISESTLVPLYGRGKTTSSSEGEA 153
>TC82766 similar to GP|3128183|gb|AAC16087.1| unknown protein {Arabidopsis
thaliana}, partial (10%)
Length = 690
Score = 53.1 bits (126), Expect = 2e-08
Identities = 19/39 (48%), Positives = 28/39 (71%)
Frame = +2
Query: 1 CNICFDLAQDPVITLCGYLFRWPCLYKWLHFHSLSQQCP 39
CNIC D A +PV+T CG+LF W C Y+ + +S +++CP
Sbjct: 572 CNICLDTANNPVLTCCGHLFCWECFYQLAYAYSNAKECP 688
>AJ497196 weakly similar to SP|O60683|PEXA_ Peroxisome assembly protein 10
(Peroxin-10). [Human] {Homo sapiens}, partial (11%)
Length = 582
Score = 47.4 bits (111), Expect = 1e-06
Identities = 19/62 (30%), Positives = 31/62 (49%)
Frame = +1
Query: 1 CNICFDLAQDPVITLCGYLFRWPCLYKWLHFHSLSQQCPVCNALAEEDKLVPLHGRGNTS 60
C++C + T CG++F W C+ WL + QCP+C E +L+P+ N +
Sbjct: 310 CSLCLEARTQDTATPCGHVFCWTCISGWLQTRA---QCPLCRDFVEISRLIPIQNMINQA 480
Query: 61 SS 62
S
Sbjct: 481 RS 486
>BF004323 similar to GP|4337011|gb| zinc-binding peroxisomal integral
membrane protein {Arabidopsis thaliana}, partial (32%)
Length = 513
Score = 40.4 bits (93), Expect = 1e-04
Identities = 16/51 (31%), Positives = 24/51 (46%)
Frame = -1
Query: 1 CNICFDLAQDPVITLCGYLFRWPCLYKWLHFHSLSQQCPVCNALAEEDKLV 51
C +C Q P T CG++F W C+ +W + +CP+C LV
Sbjct: 255 CTLCLSNRQHPTATSCGHVFCWNCITEWC---NEKPECPLCRTPITHSSLV 112
>TC87965 similar to GP|11994306|dbj|BAB01736.
emb|CAA66822.1~gene_id:MLJ15.12~unknown protein
{Arabidopsis thaliana}, partial (20%)
Length = 955
Score = 37.7 bits (86), Expect = 9e-04
Identities = 19/60 (31%), Positives = 29/60 (47%), Gaps = 6/60 (10%)
Frame = +2
Query: 1 CNICFDLAQDPVITLCGYLFRWPCLYKWL------HFHSLSQQCPVCNALAEEDKLVPLH 54
C IC + P IT CG++F +PC+ ++L H ++CP+C L LH
Sbjct: 752 CPICLEHPLCPQITSCGHIFCFPCILQYLMLGEEDHKGDRWKRCPLCFVTISVKDLYTLH 931
>BQ137845 similar to GP|22795037|gb| putative RING protein {Populus x
canescens}, partial (15%)
Length = 490
Score = 34.7 bits (78), Expect = 0.007
Identities = 15/26 (57%), Positives = 17/26 (64%)
Frame = +1
Query: 37 QCPVCNALAEEDKLVPLHGRGNTSSS 62
QCPVC K+VPL+GRG T SS
Sbjct: 52 QCPVCKHDISHTKMVPLYGRGQTLSS 129
>TC83036 similar to PIR|C96637|C96637 hypothetical protein F11P17.13
[imported] - Arabidopsis thaliana, partial (20%)
Length = 1256
Score = 34.3 bits (77), Expect = 0.010
Identities = 14/42 (33%), Positives = 23/42 (54%)
Frame = +2
Query: 1 CNICFDLAQDPVITLCGYLFRWPCLYKWLHFHSLSQQCPVCN 42
C IC D ++ V+++CG++F C+ + H QCP N
Sbjct: 686 CGICNDAPEEAVVSVCGHVFCNQCICE--HLTGEDNQCPATN 805
>TC82517 homologue to GP|7688065|emb|CAB89694.1 constitutively
photomorphogenic 1 protein {Pisum sativum}, partial
(22%)
Length = 671
Score = 33.5 bits (75), Expect = 0.016
Identities = 16/52 (30%), Positives = 22/52 (41%)
Frame = +3
Query: 1 CNICFDLAQDPVITLCGYLFRWPCLYKWLHFHSLSQQCPVCNALAEEDKLVP 52
C IC + +D +T CG+ F + C+ L S CP C L P
Sbjct: 159 CPICMQIIKDAFLTSCGHSFCYMCIITHLRNKS---DCPCCGHYLTNSNLFP 305
>TC80776 similar to GP|13605815|gb|AAK32893.1 At2g32700/F24L7.16
{Arabidopsis thaliana}, partial (19%)
Length = 1087
Score = 33.1 bits (74), Expect = 0.021
Identities = 13/41 (31%), Positives = 22/41 (52%)
Frame = +2
Query: 1 CNICFDLAQDPVITLCGYLFRWPCLYKWLHFHSLSQQCPVC 41
C +C DL P++ CGY+ + C++K + +CP C
Sbjct: 107 CCVCLDLFYKPIVLSCGYMCCFWCIHKSMS-GVRESKCPTC 226
>TC78603 weakly similar to GP|19698827|gb|AAL91149.1 unknown protein
{Arabidopsis thaliana}, partial (28%)
Length = 1559
Score = 32.7 bits (73), Expect = 0.028
Identities = 14/47 (29%), Positives = 23/47 (48%), Gaps = 4/47 (8%)
Frame = +3
Query: 1 CNICFDLAQDP----VITLCGYLFRWPCLYKWLHFHSLSQQCPVCNA 43
C++C Q+ ++ C + F PC+ WL H + CP+C A
Sbjct: 537 CSVCLGEFQEEESLRILPKCSHAFHIPCIDTWLSSH---KNCPLCRA 668
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.324 0.137 0.473
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 3,885,954
Number of Sequences: 36976
Number of extensions: 73019
Number of successful extensions: 729
Number of sequences better than 10.0: 129
Number of HSP's better than 10.0 without gapping: 708
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 718
length of query: 68
length of database: 9,014,727
effective HSP length: 44
effective length of query: 24
effective length of database: 7,387,783
effective search space: 177306792
effective search space used: 177306792
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 51 (24.3 bits)
Lotus: description of TM0052a.2


