

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0052a.1
(387 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
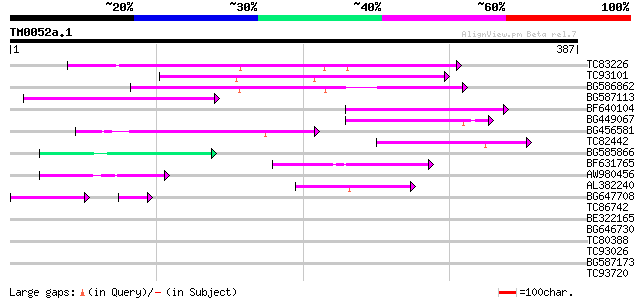
Score E
Sequences producing significant alignments: (bits) Value
TC83226 weakly similar to PIR|G86419|G86419 probable reverse tra... 120 9e-28
TC93101 91 1e-18
BG586862 85 4e-17
BG587113 weakly similar to PIR|A84888|A8 hypothetical protein At... 70 1e-12
BF640104 55 4e-08
BG449067 54 8e-08
BG456581 53 2e-07
TC82442 48 7e-06
BG585866 47 1e-05
BF631765 46 2e-05
AW980456 44 8e-05
AL382240 42 3e-04
BG647708 weakly similar to GP|13786450|gb| putative reverse tran... 38 8e-04
TC86742 similar to PIR|T01541|T01541 hypothetical protein A_IG00... 40 0.001
BE322165 40 0.002
BG646730 35 0.064
TC80388 34 0.084
TC93026 similar to PIR|T01856|T01856 hypothetical protein F9D12.... 34 0.11
BG587173 weakly similar to GP|4512630|gb| Contains reverse trans... 34 0.11
TC93720 similar to GP|21743320|dbj|BAC03314. putative nematode r... 32 0.32
>TC83226 weakly similar to PIR|G86419|G86419 probable reverse transcriptase
100033-105622 [imported] - Arabidopsis thaliana, partial
(2%)
Length = 885
Score = 120 bits (301), Expect = 9e-28
Identities = 84/285 (29%), Positives = 133/285 (46%), Gaps = 16/285 (5%)
Frame = +3
Query: 40 WPESSNGHYTSKAGYVFLRKLASIEAPSSSTAVGFSPRFWKGFWATSALPRCKETCWRAI 99
W + G Y+ K+GY LR + + ++ST+ WK W+ +PR K WR +
Sbjct: 6 WMHNPTGIYSVKSGYNTLRTWQTQQINNTSTSSD-ETLIWKKIWSLHTIPRHKVLLWRIL 182
Query: 100 SGYLPLRDRLFRRHVDVDPGCCLCSGENETAEHLFMLCPAAKQV*YASSLSLRVDSF--I 157
+ LP+R L +R + P C C + ET HLFM CP +K+V + S+L + D+
Sbjct: 183 NDSLPVRSSLRKRGIQCYPLCPRCHSKTETITHLFMSCPLSKRVWFGSNLCINFDNLPNP 362
Query: 158 AFADFWAWVVEQGDSEFISTVQTIIYAIWEARNQVQFQQRTFSVTGVLRRVAAMNS---- 213
F + + D + IIY +W ARN + +T +++R + S
Sbjct: 363 NFIN*LYEAIL*KDECITI*IAAIIYNLWHARNLSVLEDQTILEMDIIQRASNCISDYKQ 542
Query: 214 -------ASQKSDGGPQTGSRPA---RWRRPQPGTIKCNFDASFRGGGTAGLGMVARNSE 263
+ ++ P++ RPA +W+RP G +K N DA+ + G GLG++ R+
Sbjct: 543 ANTQAPPSMARTGYDPRSQHRPAKNTKWKRPNLGLVKVNTDANLQNHGKWGLGIIIRDEV 722
Query: 264 GEAMAAACSFPVPISSPLLAEAMAMRWTMQLAIDLGFH*IVLETD 308
G MAA+ L AEA A+ M+ A D GF + E D
Sbjct: 723 GLVMAASTWETDGNDRALEAEAYALLTGMRFAKDCGFXKVXFEGD 857
>TC93101
Length = 675
Score = 90.5 bits (223), Expect = 1e-18
Identities = 64/220 (29%), Positives = 100/220 (45%), Gaps = 22/220 (10%)
Frame = -1
Query: 103 LPLRDRLFRRHVDVDPGCCLCSGENETAEHLFMLCPAAKQV*YASSLSLRV--DSFIAFA 160
LP+R L +R V+ P C C ET H+FM C ++V + S LS+R +S I F+
Sbjct: 663 LPVRXELNKRGVNCPPLCPRCYFNLETTNHIFMSCERTQRVWFGSQLSIRFPDNSTINFS 484
Query: 161 DFWAWVVEQGDSEFISTVQTIIYAIWEARNQVQFQQRTFSVTGVLR-------------- 206
D+ + E I + I Y+IW ARN+ F+ + S +++
Sbjct: 483 DWLFDAISNQTEEIIIKISAITYSIWHARNKAIFENQFVSEDTIIQ*AQNSILAYEQATI 304
Query: 207 ------RVAAMNSASQKSDGGPQTGSRPARWRRPQPGTIKCNFDASFRGGGTAGLGMVAR 260
V + SA+ ++ + + +RW++P +K N DA+ + G GLG + R
Sbjct: 303 KPQNPNIVLSSLSATSNTNTTRRRSNVRSRWQKPLNNILKANCDANLQVQGRWGLGCIIR 124
Query: 261 NSEGEAMAAACSFPVPISSPLLAEAMAMRWTMQLAIDLGF 300
N++GEA A AE A+ M LA D GF
Sbjct: 123 NADGEAKVTATWCINGFDCAATAETYAILAAMYLAKDYGF 4
>BG586862
Length = 804
Score = 85.1 bits (209), Expect = 4e-17
Identities = 65/235 (27%), Positives = 101/235 (42%), Gaps = 5/235 (2%)
Frame = -1
Query: 83 WATSALPRCKETCWRAISGYLPLRDRLFRRHVDVDPGCCLCSGENETAEHLFMLCPAAKQ 142
W +PR K WR + LP++D L +R + C C + ET +HLF+ C ++
Sbjct: 651 WGIKTIPRHKSFLWRLLHNALPVKDELHKRGIRCSLLCPRCESKIETVQHLFLNCEVTQK 472
Query: 143 V*YASSLSLRVDS--FIAFADFWAWVVEQGDSEFISTVQTIIYAIWEARNQVQFQQRTFS 200
+ S L + S + F D+ + + D E I + ++Y+IW ARNQ F+
Sbjct: 471 EWFGSQLGINFHSSGVLHFHDWITNFILKNDEETIIALTALLYSIWHARNQKVFENIDVP 292
Query: 201 VTGVLRRVAAMNSA---SQKSDGGPQTGSRPARWRRPQPGTIKCNFDASFRGGGTAGLGM 257
V++R ++ + +Q SD + + P+ G+G+
Sbjct: 291 GDVVIQRASSSLHSFKMAQVSDSVLPSNAIPSY--------------------SLWGIGV 172
Query: 258 VARNSEGEAMAAACSFPVPISSPLLAEAMAMRWTMQLAIDLGFH*IVLETDCLNL 312
VARN EG AMA+ I AEA + M A D GF E+D NL
Sbjct: 171 VARNCEGLAMASGTWLRHGIPCATTAEAWGIYQAMVFAGDCGFSKFEFESDNGNL 7
>BG587113 weakly similar to PIR|A84888|A8 hypothetical protein At2g45230
[imported] - Arabidopsis thaliana, partial (10%)
Length = 767
Score = 70.1 bits (170), Expect = 1e-12
Identities = 38/134 (28%), Positives = 57/134 (42%)
Frame = -3
Query: 10 DLIEFVFSPSTAEIILSIPLSLNGGRDILFWPESSNGHYTSKAGYVFLRKLASIEAPSSS 69
+LI +F T ILSI G D W S +GHY+ K+GY + + +
Sbjct: 504 NLINSIFPEGTRRKILSIHPQGPIGEDSYSWEYSKSGHYSVKSGYYVQTNIIAAANQRGT 325
Query: 70 TAVGFSPRFWKGFWATSALPRCKETCWRAISGYLPLRDRLFRRHVDVDPGCCLCSGENET 129
++ W + P+ + WR IS LP + RH+ D C C E+ET
Sbjct: 324 VDQPSLDDLYQRVWKYNTSPKVRHFLWRCISNSLPTAANMRSRHISKDGSCSRCGMESET 145
Query: 130 AEHLFMLCPAAKQV 143
H+ CP A+ +
Sbjct: 144 VNHILFQCPYARLI 103
>BF640104
Length = 344
Score = 55.5 bits (132), Expect = 4e-08
Identities = 35/112 (31%), Positives = 51/112 (45%), Gaps = 1/112 (0%)
Frame = -1
Query: 230 RWRRPQPGTIKCNFDASFRGGGTAGLGMVARNSEGEAMAAACSFPVPISSPLLAEAMAMR 289
+W +P G IK N DA+ G+G++ RN G MA+ P+ AEA +
Sbjct: 341 KWIKPHQGVIKINCDANLTSEDVWGIGVITRNDNGIVMASGTWNRPGFMCPITAEAWGVY 162
Query: 290 WTMQLAIDLGFH*IVLETDCLNLFNAWRTIEE-DRSYLSILIRDCRLLISAF 340
A+D GF ++ E D L + EE RSYL +I+ L+ F
Sbjct: 161 QAALFALDQGFQNVLFENDNEKLISMLSREEEGHRSYLGSIIQSIISLVPRF 6
>BG449067
Length = 578
Score = 54.3 bits (129), Expect = 8e-08
Identities = 36/104 (34%), Positives = 52/104 (49%), Gaps = 3/104 (2%)
Frame = -2
Query: 230 RWRRPQPGTIKCNFDASFRGGGTAGLGMVARNSEGEAMAAACSFPVPISSPLLAEAMAMR 289
+W++P+ IK N DA+ G+G+VARN EG AMA+ F S AEA +
Sbjct: 310 KWKKPEKDIIKLNSDANLSSTDLWGIGVVARNDEGFAMASGTWFRFGFPSATTAEAWGIY 131
Query: 290 WTMQLAIDLGFH*IVLETD---CLNLFNAWRTIEEDRSYLSILI 330
M A + GF + E+D + + N T E +R YL +I
Sbjct: 130 QAMIFAGEYGFSKVQFESDNERVIQMLNG--TEEVNRLYLGSII 5
>BG456581
Length = 683
Score = 52.8 bits (125), Expect = 2e-07
Identities = 48/171 (28%), Positives = 69/171 (40%), Gaps = 5/171 (2%)
Frame = +2
Query: 46 GHYTSKAGYVFLRKLASIEAPSSSTAVGFSPRFWKGFWATSALPRCKETCWRAISGYLPL 105
G T K Y FL S AP +ST W + P CWR LP
Sbjct: 8 GKLTIKLVYSFLTSHTSC-APWAST-----------IWNSCIPPSHSFICWRLAHDRLPT 151
Query: 106 RDRLFRRHVDVDPGCCLCSGENETAEHLFMLCPAAKQV*YASSLSLRVD-SFIAFADFWA 164
D L R + C C + ET++HLF+ C + LRV F +F +
Sbjct: 152 DDNLSSRGCALVSMCSFCLEQVETSDHLFLRCKFVVTLWSWLCSQLRVGLDFSSFKALLS 331
Query: 165 WVVEQGDSE----FISTVQTIIYAIWEARNQVQFQQRTFSVTGVLRRVAAM 211
+ S+ +++ V ++++IW ARN V+F S V RV A+
Sbjct: 332 SLPRHCSSQVRDLYVAAVVHMVHSIWWARNNVRFSSAKVSAHAVQVRVHAL 484
>TC82442
Length = 578
Score = 47.8 bits (112), Expect = 7e-06
Identities = 35/109 (32%), Positives = 53/109 (48%), Gaps = 3/109 (2%)
Frame = -3
Query: 251 GTAGLGMVARNSEGEAMAAACSFPVPISSPLLAEAMAMRWTMQLAIDLGFH*IVLETDCL 310
G GLG + NS G+AM +A + AEA A+ M LA+ ++ E+DC
Sbjct: 330 GRWGLGCIIHNSAGQAMTSATWSRNGFNCAATAEAYAILAAMHLALTSDLKHMIFESDCE 151
Query: 311 NLFNAW-RTIEEDR--SYLSILIRDCRLLISAFEFVSLCFVRRTGNAVA 356
+ R IE+D SYL ++ + + F+ S F+ R+GN VA
Sbjct: 150 AVIRKLNRGIEKDIDISYLGSVLEEIWKISEKFDMCSFRFIPRSGNRVA 4
>BG585866
Length = 828
Score = 47.0 bits (110), Expect = 1e-05
Identities = 32/122 (26%), Positives = 49/122 (39%), Gaps = 1/122 (0%)
Frame = +3
Query: 21 AEIILSIPLSLNGG-RDILFWPESSNGHYTSKAGYVFLRKLASIEAPSSSTAVGFSPRFW 79
AE+I + L+ N D WP +SNG YT+K+GY ++ S + V ++ W
Sbjct: 300 AEVINNTHLNFNASIGDAYIWPHNSNGVYTAKSGYSWIL--------SQTETVNYNNSSW 455
Query: 80 KGFWATSALPRCKETCWRAISGYLPLRDRLFRRHVDVDPGCCLCSGENETAEHLFMLCPA 139
W + K W A +P L R++ C C E+ H C
Sbjct: 456 SWIWRLKIPEKYKFFLWLACHNAVPTLSLLNHRNMVNSAICSRCGEHEESFFHCVRDCRF 635
Query: 140 AK 141
+K
Sbjct: 636 SK 641
>BF631765
Length = 498
Score = 46.2 bits (108), Expect = 2e-05
Identities = 37/110 (33%), Positives = 52/110 (46%)
Frame = +1
Query: 180 TIIYAIWEARNQVQFQQRTFSVTGVLRRVAAMNSASQKSDGGPQTGSRPARWRRPQPGTI 239
T ++ IW+ RNQ F F V V + A+ G Q P R + P G+I
Sbjct: 124 TSLWLIWKHRNQKVFNNVDFDPPLVSALVEEFSIANLPLKRG-QVCVVP-R*QPPPFGSI 297
Query: 240 KCNFDASFRGGGTAGLGMVARNSEGEAMAAACSFPVPISSPLLAEAMAMR 289
K N A G G+ G G +ARN+ G A++A SPLLAE + ++
Sbjct: 298 KVNVGAGCFGDGSTGWGFIARNAAGLALSAETKSDGISYSPLLAECLLLK 447
>AW980456
Length = 779
Score = 44.3 bits (103), Expect = 8e-05
Identities = 28/89 (31%), Positives = 39/89 (43%)
Frame = -2
Query: 21 AEIILSIPLSLNGGRDILFWPESSNGHYTSKAGYVFLRKLASIEAPSSSTAVGFSPRFWK 80
A+ I+S PL + D L W + +G Y K+ Y F +E S+ + P W
Sbjct: 253 ADKIMSTPLISHA*LDRLIWKDEKHGKYYVKSAYRFC-----VEELFDSSYL-HRPGNWS 92
Query: 81 GFWATSALPRCKETCWRAISGYLPLRDRL 109
G W P+ + WR G LP R RL
Sbjct: 91 GIWKLKVPPKVQNLVWRMCRGCLPTRIRL 5
>AL382240
Length = 320
Score = 42.4 bits (98), Expect = 3e-04
Identities = 29/86 (33%), Positives = 38/86 (43%), Gaps = 4/86 (4%)
Frame = +3
Query: 196 QRTFSVTGVLRRVAAMNSASQKSDGGPQTG-SRPAR--WRRPQPGTIKCNFDASFRGG-G 251
QR V R + S+S PQ SRP W++P PG KCN DASF
Sbjct: 15 QRATHVFEEWRTTQRIRSSSNTQSTTPQVQPSRPEEDVWKKPAPGXYKCNIDASFSTSFN 194
Query: 252 TAGLGMVARNSEGEAMAAACSFPVPI 277
GLGM R+ + + A + P+
Sbjct: 195 IVGLGMCLRDEDDAFVLARTEWFAPL 272
>BG647708 weakly similar to GP|13786450|gb| putative reverse transcriptase
{Oryza sativa}, partial (9%)
Length = 708
Score = 37.7 bits (86), Expect(2) = 8e-04
Identities = 18/54 (33%), Positives = 28/54 (51%)
Frame = +3
Query: 1 DHDRHCWRRDLIEFVFSPSTAEIILSIPLSLNGGRDILFWPESSNGHYTSKAGY 54
D+D W RDLI F+ A I+ IPLS+ D + W +G Y+ ++ +
Sbjct: 417 DYDTKQWNRDLIFHSFNNYAAHQIIKIPLSMRQPEDKIIWHWEKDGIYSVRSAH 578
Score = 22.3 bits (46), Expect(2) = 8e-04
Identities = 8/23 (34%), Positives = 10/23 (42%)
Frame = +2
Query: 75 SPRFWKGFWATSALPRCKETCWR 97
+P WK W T A + WR
Sbjct: 632 NPVIWKAIWNTPASRSVQNFLWR 700
>TC86742 similar to PIR|T01541|T01541 hypothetical protein A_IG005I10.16 -
Arabidopsis thaliana, partial (19%)
Length = 2073
Score = 40.4 bits (93), Expect = 0.001
Identities = 34/107 (31%), Positives = 56/107 (51%), Gaps = 1/107 (0%)
Frame = -1
Query: 257 MVARNSE-GEAMAAACSFPVPISSPLLAEAMAMRWTMQLAIDLGFH*IVLETDCLNLFNA 315
+V R+S+ G A +C+ + ++PL AE A ++ A++LG + I LETD L + NA
Sbjct: 471 LVIRDSQFGFLGALSCN--IGHATPLEAEFCACMIAIEKAMELGLNNICLETDSLKVVNA 298
Query: 316 WRTIEEDRSYLSILIRDCRLLISAFEFVSLCFVRRTGNAVADFLARN 362
+ I + + +C + V + + R GN VAD LAR+
Sbjct: 297 FHKIVGIPWQMRVRWHNCIRFCHSIACVCV-HIPREGNLVADALARH 160
>BE322165
Length = 650
Score = 39.7 bits (91), Expect = 0.002
Identities = 29/80 (36%), Positives = 40/80 (49%), Gaps = 3/80 (3%)
Frame = -3
Query: 254 GLGMVARNSEGEAMAAACSFPVPISSPLLAEAMAMRWTMQLAIDLGFH*IVLETD---CL 310
G+G+VARN EG AMA+ F S AEA + M A + GF + E+D +
Sbjct: 246 GIGVVARNDEGFAMASGTWFRFGFPSATTAEAWGIYQAMIFAGEYGFSKVQFESDNERVI 67
Query: 311 NLFNAWRTIEEDRSYLSILI 330
+ N T E +R YL +I
Sbjct: 66 QMLNG--TEEVNRLYLGSII 13
>BG646730
Length = 799
Score = 34.7 bits (78), Expect = 0.064
Identities = 25/66 (37%), Positives = 40/66 (59%), Gaps = 2/66 (3%)
Frame = +1
Query: 8 RRDLIEFVFSPSTAEIILSIPLS--LNGGRDILFWPESSNGHYTSKAGYVFLRKLASIEA 65
+RDL+ VFS A+ IL+IPLS L R IL+W N Y+ ++ + L K+A+ ++
Sbjct: 604 KRDLVRNVFSAFEAKQILNIPLSWRLWPDRRILYWERDRN--YSVRSAH-HLIKVATKDS 774
Query: 66 PSSSTA 71
P S++
Sbjct: 775 PEVSSS 792
>TC80388
Length = 1112
Score = 34.3 bits (77), Expect = 0.084
Identities = 17/55 (30%), Positives = 28/55 (50%)
Frame = +2
Query: 256 GMVARNSEGEAMAAACSFPVPISSPLLAEAMAMRWTMQLAIDLGFH*IVLETDCL 310
G V +N+ G+ C + P + EA+ +R +QLA D G + ++TD L
Sbjct: 644 GCVFKNNSGDISMIVCKKESIATDPCMVEALGVR*CLQLARDQGLQKLEIQTDAL 808
>TC93026 similar to PIR|T01856|T01856 hypothetical protein F9D12.19 -
Arabidopsis thaliana, partial (16%)
Length = 831
Score = 33.9 bits (76), Expect = 0.11
Identities = 32/131 (24%), Positives = 54/131 (40%)
Frame = +3
Query: 237 GTIKCNFDASFRGGGTAGLGMVARNSEGEAMAAACSFPVPISSPLLAEAMAMRWTMQLAI 296
G +KC DA F + GLGM + + A +S + + +
Sbjct: 18 GFLKCKVDAGFSTSVSTGLGMYCCDWNKAFVGAKD*EGSFLSCYSRMGNLQSFESSPMGS 197
Query: 297 DLGFH*IVLETDCLNLFNAWRTIEEDRSYLSILIRDCRLLISAFEFVSLCFVRRTGNAVA 356
+L FH ++ + D L + D S ++I+ C LIS F + F+ R N+V
Sbjct: 198 ELDFHDVLFDVD-YKLVADSNNPKRDISSFGLIIKKCINLISFFLNFRVSFIGRQANSVV 374
Query: 357 DFLARNANTYA 367
+A+ A + A
Sbjct: 375 HKIAKEALSIA 407
>BG587173 weakly similar to GP|4512630|gb| Contains reverse transcriptase
domain (rvt) PF|00078. {Arabidopsis thaliana}, partial
(5%)
Length = 292
Score = 33.9 bits (76), Expect = 0.11
Identities = 16/61 (26%), Positives = 27/61 (44%)
Frame = +1
Query: 83 WATSALPRCKETCWRAISGYLPLRDRLFRRHVDVDPGCCLCSGENETAEHLFMLCPAAKQ 142
W + +P+ W A LP R R + C LC +E+ +HLF+ C ++
Sbjct: 103 WFSGHIPKHAFNMWVANYDRLPTRCRFAAWDTSIVKTCLLCGHYDESRDHLFLRCSFSEH 282
Query: 143 V 143
+
Sbjct: 283 I 285
>TC93720 similar to GP|21743320|dbj|BAC03314. putative nematode
resistance-like protein {Oryza sativa (japonica
cultivar-group)}, partial (3%)
Length = 457
Score = 32.3 bits (72), Expect = 0.32
Identities = 31/142 (21%), Positives = 55/142 (37%), Gaps = 14/142 (9%)
Frame = -2
Query: 166 VVEQGDSEFISTVQTIIYAIWEARNQVQFQQRTFSVTGVLRRV--------AAMNSASQK 217
+++Q S+ + T+++++W++ +QQ S + V+ AA S
Sbjct: 429 LLQQFSSDQCELMATVMWSLWKSCRMKLWQQNNESNSQVVHHATHLLNDWRAAQTI*SHN 250
Query: 218 SDGGPQTGSRPAR-----WRRPQPGTIKCNFDAS-FRGGGTAGLGMVARNSEGEAMAAAC 271
D P + W P KCN DAS F GLGM + + + A
Sbjct: 249 HDQSNMVYPCPMKQDEEAWMTLTPDMYKCNIDASFFTSLNKVGLGMCLADDADDFVLAKT 70
Query: 272 SFPVPISSPLLAEAMAMRWTMQ 293
+ P EA+ + T++
Sbjct: 69 IWFAPSCDIDAGEAVRLHTTLE 4
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.327 0.138 0.447
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 13,569,980
Number of Sequences: 36976
Number of extensions: 199370
Number of successful extensions: 1343
Number of sequences better than 10.0: 54
Number of HSP's better than 10.0 without gapping: 1323
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1335
length of query: 387
length of database: 9,014,727
effective HSP length: 98
effective length of query: 289
effective length of database: 5,391,079
effective search space: 1558021831
effective search space used: 1558021831
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.7 bits)
S2: 59 (27.3 bits)
Lotus: description of TM0052a.1


