

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
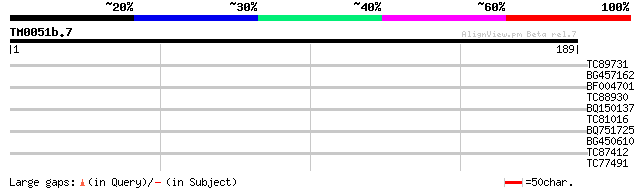
Query= TM0051b.7
(189 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC89731 similar to GP|15809986|gb|AAL06920.1 AT3g19540/T31J18_4 ... 30 0.73
BG457162 28 2.1
BF004701 similar to GP|15982777|gb| unknown protein {Arabidopsis... 27 3.6
TC88930 similar to SP|Q99MR1|PERQ_MOUSE PERQ amino acid rich wit... 27 4.7
BQ150137 similar to OMNI|NT01MC0313 transposase {Magnetococcus s... 27 4.7
TC81016 similar to SP|P25795|DHAX_PEA Turgor-responsive protein ... 27 6.2
BQ751725 homologue to GP|10047327|dbj KIAA1625 protein {Homo sap... 27 6.2
BG450610 similar to SP|P25795|DHAX Turgor-responsive protein 26G... 27 6.2
TC87412 similar to PIR|T00587|T00587 probable ubiquitin--protein... 26 8.1
TC77491 similar to GP|13937149|gb|AAK50068.1 At1g79380/T8K14_20 ... 26 8.1
>TC89731 similar to GP|15809986|gb|AAL06920.1 AT3g19540/T31J18_4
{Arabidopsis thaliana}, partial (21%)
Length = 717
Score = 29.6 bits (65), Expect = 0.73
Identities = 13/18 (72%), Positives = 13/18 (72%)
Frame = -2
Query: 97 FPSLNPPSDPSEPSGPVF 114
FPSL P S PS PSGP F
Sbjct: 440 FPSLFPTSSPSVPSGPSF 387
>BG457162
Length = 661
Score = 28.1 bits (61), Expect = 2.1
Identities = 20/65 (30%), Positives = 32/65 (48%), Gaps = 4/65 (6%)
Frame = +3
Query: 83 PNRNRPIQLI--WSRFFPSLNPP--SDPSEPSGPVFGRGPVGFVQEVMVKENVHINIFDI 138
P+R + LI +SR+ PS+NPP + + + P F + PV M + I +F
Sbjct: 261 PSRGLTLVLI*NFSRYHPSINPPQLNHRTTRAFPTFSQYPVSIFPHSM-SSSTSIQLFSS 437
Query: 139 QGAPL 143
+G L
Sbjct: 438 EGNKL 452
>BF004701 similar to GP|15982777|gb| unknown protein {Arabidopsis thaliana},
partial (47%)
Length = 505
Score = 27.3 bits (59), Expect = 3.6
Identities = 10/18 (55%), Positives = 15/18 (82%), Gaps = 1/18 (5%)
Frame = -1
Query: 132 HINIFDIQG-APLFYQWI 148
H+NIF+IQG P+F +W+
Sbjct: 466 HVNIFEIQGNCPIF*EWV 413
>TC88930 similar to SP|Q99MR1|PERQ_MOUSE PERQ amino acid rich with GYF
domain protein 1. [Mouse] {Mus musculus}, partial (1%)
Length = 910
Score = 26.9 bits (58), Expect = 4.7
Identities = 11/32 (34%), Positives = 18/32 (55%)
Frame = -1
Query: 22 HRNRLRQNEDEQAISDQRSMEEEGKKLRRCSG 53
++ R +QN+ E+ +Q E +K RRC G
Sbjct: 331 NQERNQQNDGEKVPEEQNERSPERRKKRRCGG 236
>BQ150137 similar to OMNI|NT01MC0313 transposase {Magnetococcus sp. MC-1},
partial (9%)
Length = 1056
Score = 26.9 bits (58), Expect = 4.7
Identities = 14/28 (50%), Positives = 18/28 (64%), Gaps = 1/28 (3%)
Frame = +3
Query: 16 GGSGRFHRNRLRQN-EDEQAISDQRSME 42
G SGR H +RLR+N E + I +RS E
Sbjct: 513 GASGRLHDDRLRRNVESTRKIEKKRSSE 596
>TC81016 similar to SP|P25795|DHAX_PEA Turgor-responsive protein 26G (EC
1.2.1.-). [Garden pea] {Pisum sativum}, partial (71%)
Length = 1132
Score = 26.6 bits (57), Expect = 6.2
Identities = 10/17 (58%), Positives = 12/17 (69%)
Frame = -2
Query: 96 FFPSLNPPSDPSEPSGP 112
FFPS PP+DP +P P
Sbjct: 345 FFPSQEPPTDPVDPI*P 295
>BQ751725 homologue to GP|10047327|dbj KIAA1625 protein {Homo sapiens},
partial (2%)
Length = 514
Score = 26.6 bits (57), Expect = 6.2
Identities = 15/49 (30%), Positives = 19/49 (38%)
Frame = -3
Query: 39 RSMEEEGKKLRRCSGGVARVVAAQRMAAQVMRRRVVAVVRSDWAPNRNR 87
RS + LR CS A+ + R + RR S W P R R
Sbjct: 455 RSSPPWSRSLRTCSRPRAQTASTARPRGKRTTRRTTRTASSAWPPPRRR 309
>BG450610 similar to SP|P25795|DHAX Turgor-responsive protein 26G (EC
1.2.1.-). [Garden pea] {Pisum sativum}, partial (33%)
Length = 619
Score = 26.6 bits (57), Expect = 6.2
Identities = 10/17 (58%), Positives = 12/17 (69%)
Frame = -1
Query: 96 FFPSLNPPSDPSEPSGP 112
FFPS PP+DP +P P
Sbjct: 349 FFPSQEPPTDPVDPI*P 299
>TC87412 similar to PIR|T00587|T00587 probable ubiquitin--protein ligase (EC
6.3.2.19) - Arabidopsis thaliana, partial (42%)
Length = 1628
Score = 26.2 bits (56), Expect = 8.1
Identities = 10/42 (23%), Positives = 17/42 (39%)
Frame = +3
Query: 64 MAAQVMRRRVVAVVRSDWAPNRNRPIQLIWSRFFPSLNPPSD 105
MAA ++R + DW N + +++ P P D
Sbjct: 519 MAASILRAETFGIPTPDWVKNPTKLAEVVDRMIVPDFQPKKD 644
>TC77491 similar to GP|13937149|gb|AAK50068.1 At1g79380/T8K14_20 {Arabidopsis
thaliana}, partial (81%)
Length = 1537
Score = 26.2 bits (56), Expect = 8.1
Identities = 10/18 (55%), Positives = 14/18 (77%)
Frame = -2
Query: 40 SMEEEGKKLRRCSGGVAR 57
SM++ G LR+CSGG+ R
Sbjct: 1104 SMDDVGILLRKCSGGICR 1051
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.326 0.143 0.432
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 5,575,904
Number of Sequences: 36976
Number of extensions: 71487
Number of successful extensions: 603
Number of sequences better than 10.0: 20
Number of HSP's better than 10.0 without gapping: 594
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 602
length of query: 189
length of database: 9,014,727
effective HSP length: 91
effective length of query: 98
effective length of database: 5,649,911
effective search space: 553691278
effective search space used: 553691278
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 55 (25.8 bits)
Lotus: description of TM0051b.7


