

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0051b.2
(276 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
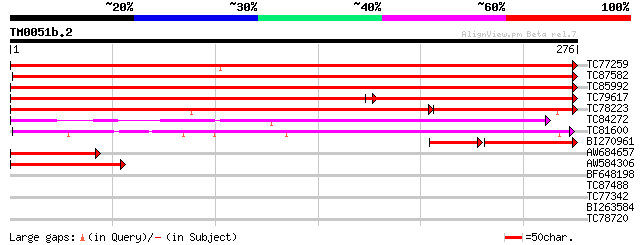
Score E
Sequences producing significant alignments: (bits) Value
TC77259 similar to PIR|T12558|T12558 porin - common ice plant, p... 448 e-127
TC87582 similar to PIR|S36454|S36454 porin por1 - garden pea, co... 442 e-125
TC85992 similar to GP|5031279|gb|AAD38145.1| porin {Prunus armen... 395 e-111
TC79617 weakly similar to GP|9757871|dbj|BAB08458.1 porin-like p... 175 8e-74
TC78223 weakly similar to GP|9758310|dbj|BAB08784.1 porin-like p... 189 9e-63
TC84272 weakly similar to PIR|S46959|S46959 porin I 36K - potat... 152 2e-37
TC81600 weakly similar to SP|P07144|PORI_NEUCR Outer mitochondri... 77 9e-15
BI270961 similar to GP|9757871|dbj| porin-like protein {Arabidop... 51 4e-12
AW684657 similar to GP|9757871|dbj| porin-like protein {Arabidop... 66 1e-11
AW584306 weakly similar to PIR|T09116|T091 voltage-dependent ani... 58 3e-09
BF648198 29 2.2
TC87488 28 2.9
TC77342 similar to GP|13785211|emb|CAC37357. putative membrane p... 28 3.8
BI263584 similar to SP|Q9P6C8|ADH1 Alcohol dehydrogenase I (EC 1... 27 6.5
TC78720 annexin; annexin [Medicago truncatula] 27 6.5
>TC77259 similar to PIR|T12558|T12558 porin - common ice plant, partial
(98%)
Length = 1113
Score = 448 bits (1153), Expect = e-127
Identities = 216/278 (77%), Positives = 253/278 (90%), Gaps = 2/278 (0%)
Frame = +1
Query: 1 MAKGPGLYTDIGKKARDLLFKDYHSDQKFTITTFSPTGVAITSSGTKKGELFVADVNTQL 60
M+KGPGLY+DIGKKARDLL+KDY++DQKFT+TT+SPTG+A+T+S TKKGELF+AD+NTQL
Sbjct: 61 MSKGPGLYSDIGKKARDLLYKDYNTDQKFTLTTYSPTGIALTASSTKKGELFLADINTQL 240
Query: 61 KNKNITTDIKVDTDSNLFTTIAVNDPAPGLKAIFSFKVPDQ--RSGKVELQYLHDYAGIS 118
K+KN+TTDIKVDTDSNLFTTI V +PAPGLK I SF++P+ +SGK E+QYLHDYAGIS
Sbjct: 241 KHKNVTTDIKVDTDSNLFTTITVTEPAPGLKTILSFRIPEPELKSGKAEVQYLHDYAGIS 420
Query: 119 SSIGLTANPIVNFSGVIGTNVLALGADLTFDTKIGELTKSNAGLSFTKDDLIAALTLNDK 178
+SIGLTANP+ NFSGVIG NV+ALG DL++DTK G T+ NAGL+FTKDDLIA+LTLN+K
Sbjct: 421 ASIGLTANPVANFSGVIGNNVVALGGDLSYDTKTGVFTRLNAGLNFTKDDLIASLTLNEK 600
Query: 179 GDALNASYYHVVSPLTNTAVGAEVTHRFSTNENTLTLGTQHALDPLTTLKARVNNFGKAS 238
+ALNASYYHVV+P+T TAVGAEV+HRFST ENTLTLGTQHA+D LTT K R NNFG AS
Sbjct: 601 ANALNASYYHVVNPVTKTAVGAEVSHRFSTKENTLTLGTQHAIDSLTTAKLRFNNFGLAS 780
Query: 239 ALIQHEWRPKSFFTISGEVDTKAIEKSAKVGLALALKP 276
LIQHEWRPKSFFTISGEVDTKAIEKSA++G+ALALKP
Sbjct: 781 GLIQHEWRPKSFFTISGEVDTKAIEKSARIGIALALKP 894
>TC87582 similar to PIR|S36454|S36454 porin por1 - garden pea, complete
Length = 1307
Score = 442 bits (1136), Expect = e-125
Identities = 216/275 (78%), Positives = 240/275 (86%)
Frame = +1
Query: 2 AKGPGLYTDIGKKARDLLFKDYHSDQKFTITTFSPTGVAITSSGTKKGELFVADVNTQLK 61
AKGPGLYTDIGKKARDLLFKDY+SDQKFT++T+SPTGVAITSSGTKKGELFV DVNTQLK
Sbjct: 64 AKGPGLYTDIGKKARDLLFKDYNSDQKFTVSTYSPTGVAITSSGTKKGELFVGDVNTQLK 243
Query: 62 NKNITTDIKVDTDSNLFTTIAVNDPAPGLKAIFSFKVPDQRSGKVELQYLHDYAGISSSI 121
NKN T DIKVDT+SNLFTTI V +PAPG+KAI S KVPD SGKVELQYLHDYAGIS +I
Sbjct: 244 NKNYTADIKVDTESNLFTTITVTEPAPGVKAIISCKVPDPASGKVELQYLHDYAGISGNI 423
Query: 122 GLTANPIVNFSGVIGTNVLALGADLTFDTKIGELTKSNAGLSFTKDDLIAALTLNDKGDA 181
GL ANP+VN SGV GTN LA G D+++DTK+ ELTKSN G+SF KDDL+ ALTLN+KGD
Sbjct: 424 GLKANPVVNLSGVFGTNALAFGGDISYDTKLRELTKSNVGVSFIKDDLLGALTLNEKGDV 603
Query: 182 LNASYYHVVSPLTNTAVGAEVTHRFSTNENTLTLGTQHALDPLTTLKARVNNFGKASALI 241
LNASYYHV++P+TNTAVG ++ HRFST EN TLG QHALDPLTTLK R+ N KASALI
Sbjct: 604 LNASYYHVINPVTNTAVGVDMGHRFSTRENNFTLGVQHALDPLTTLKGRITNSAKASALI 783
Query: 242 QHEWRPKSFFTISGEVDTKAIEKSAKVGLALALKP 276
QHEWRPKS TIS E+DTKAIEKSAKVGL+L LKP
Sbjct: 784 QHEWRPKSLITISTELDTKAIEKSAKVGLSLVLKP 888
>TC85992 similar to GP|5031279|gb|AAD38145.1| porin {Prunus armeniaca},
partial (88%)
Length = 1293
Score = 395 bits (1015), Expect = e-111
Identities = 192/276 (69%), Positives = 237/276 (85%)
Frame = +1
Query: 1 MAKGPGLYTDIGKKARDLLFKDYHSDQKFTITTFSPTGVAITSSGTKKGELFVADVNTQL 60
M KGPGLY+DIGK+ARDLLFKDY +D KFTITT + GV ITS+G +KGE F+ADV+T++
Sbjct: 67 MVKGPGLYSDIGKRARDLLFKDYQNDHKFTITTQTLAGVEITSAGVRKGEAFLADVSTKV 246
Query: 61 KNKNITTDIKVDTDSNLFTTIAVNDPAPGLKAIFSFKVPDQRSGKVELQYLHDYAGISSS 120
KNKN+TTDIKVDT+SNL TTI V++ PGLK IFSF P+Q+SGKVELQYLHDYAGIS+S
Sbjct: 247 KNKNVTTDIKVDTNSNLRTTITVDETVPGLKTIFSFIYPEQKSGKVELQYLHDYAGISTS 426
Query: 121 IGLTANPIVNFSGVIGTNVLALGADLTFDTKIGELTKSNAGLSFTKDDLIAALTLNDKGD 180
IGLTA P VN S V+G N++++G+D++F+T G L K N GL+ T DLIA+LT+ND+GD
Sbjct: 427 IGLTATPTVNISAVLGNNLVSVGSDVSFETSSGSLNKCNFGLNVTHADLIASLTVNDRGD 606
Query: 181 ALNASYYHVVSPLTNTAVGAEVTHRFSTNENTLTLGTQHALDPLTTLKARVNNFGKASAL 240
+LNASYYHVVSPLTNTAVGAE++H FS+NEN LT+GTQHALDP+T LKA+VNN+GKASAL
Sbjct: 607 SLNASYYHVVSPLTNTAVGAELSHSFSSNENVLTIGTQHALDPVTLLKAKVNNYGKASAL 786
Query: 241 IQHEWRPKSFFTISGEVDTKAIEKSAKVGLALALKP 276
IQH+W ++ F++ GEVDT AI+K+AKVGLA ALKP
Sbjct: 787 IQHDWSRQARFSLVGEVDTAAIQKTAKVGLACALKP 894
>TC79617 weakly similar to GP|9757871|dbj|BAB08458.1 porin-like protein
{Arabidopsis thaliana}, partial (97%)
Length = 1208
Score = 175 bits (443), Expect(2) = 8e-74
Identities = 89/179 (49%), Positives = 121/179 (66%)
Frame = +2
Query: 1 MAKGPGLYTDIGKKARDLLFKDYHSDQKFTITTFSPTGVAITSSGTKKGELFVADVNTQL 60
M+KGPGLY+DIGKKA+DLL KDY+SDQK T++++S TGVA+TS+ KKG L DV Q
Sbjct: 68 MSKGPGLYSDIGKKAKDLLTKDYNSDQKLTVSSYSTTGVALTSTALKKGGLSTGDVAGQY 247
Query: 61 KNKNITTDIKVDTDSNLFTTIAVNDPAPGLKAIFSFKVPDQRSGKVELQYLHDYAGISSS 120
K KN D+K++T S + TT+ D P K I S K+PD SGK+E+QY HD+A ++S
Sbjct: 248 KYKNTLIDVKIETASIINTTLTFTDLFPSTKTIASIKLPDYSSGKLEVQYFHDHATLTSV 427
Query: 121 IGLTANPIVNFSGVIGTNVLALGADLTFDTKIGELTKSNAGLSFTKDDLIAALTLNDKG 179
L +PI + S +G+ LA GA+ +DTK G TK N G+SFTK D A++ + +G
Sbjct: 428 ATLKQSPIFDVSATVGSPTLAFGAEAGYDTKSGSFTKYNTGISFTKLDSSASVIIWRQG 604
Score = 119 bits (299), Expect(2) = 8e-74
Identities = 57/103 (55%), Positives = 73/103 (70%)
Frame = +3
Query: 174 TLNDKGDALNASYYHVVSPLTNTAVGAEVTHRFSTNENTLTLGTQHALDPLTTLKARVNN 233
+ DKGD++ SY H + + +A ++T RFSTN+NT+T+G A+DPLT +KAR NN
Sbjct: 588 SFGDKGDSIKVSYLHHLDQIKKSAAVVDITRRFSTNQNTITVGGSFAVDPLTQIKARFNN 767
Query: 234 FGKASALIQHEWRPKSFFTISGEVDTKAIEKSAKVGLALALKP 276
GK L+QHE PKS TISGEVDTKA+EK K GLA+ALKP
Sbjct: 768 QGKVGGLLQHELFPKSVLTISGEVDTKALEKKPKFGLAIALKP 896
>TC78223 weakly similar to GP|9758310|dbj|BAB08784.1 porin-like protein
{Arabidopsis thaliana}, partial (82%)
Length = 1503
Score = 189 bits (481), Expect(2) = 9e-63
Identities = 93/208 (44%), Positives = 136/208 (64%), Gaps = 2/208 (0%)
Frame = +2
Query: 1 MAKGPGLYTDIGKKARDLLFKDYHSDQKFTITTFSPTGVAITSSGTKKGELFVADVNTQL 60
MA GP +++IGK+ARDLL+KDY+ D KF+++ S TG+ +T++G K+ + FV D+NT
Sbjct: 110 MANGPAPFSEIGKRARDLLYKDYNFDHKFSLSIPSSTGLGLTATGLKRDKFFVGDLNTLY 289
Query: 61 KNKNITTDIKVDTDSNLFTTIAVNDPA--PGLKAIFSFKVPDQRSGKVELQYLHDYAGIS 118
K+ N+T D+KV+TDSN+ T + +ND + K SF +PD +SGK+++QYLH +A I
Sbjct: 290 KSGNVTVDVKVNTDSNVSTKVTLNDVSLSHSKKVALSFNIPDHKSGKLDVQYLHPHAAID 469
Query: 119 SSIGLTANPIVNFSGVIGTNVLALGADLTFDTKIGELTKSNAGLSFTKDDLIAALTLNDK 178
SSIGL P + S IG+ +++GA++ FDT + NAG++F K D A L L DK
Sbjct: 470 SSIGLNPAPKLELSAAIGSKDISMGAEVGFDTTSASFSTYNAGIAFNKPDFSATLMLADK 649
Query: 179 GDALNASYYHVVSPLTNTAVGAEVTHRF 206
G +L ASY H V V AE+ H+F
Sbjct: 650 GQSLKASYIHYVDRPDGLTVAAELAHKF 733
Score = 68.2 bits (165), Expect(2) = 9e-63
Identities = 36/71 (50%), Positives = 46/71 (64%), Gaps = 1/71 (1%)
Frame = +3
Query: 207 STNENTLTLGTQHALDPLTTLKARVNNFGKASALIQHEWRPKSFFTISGEVDTKAIEKS- 265
S++EN T GT ++DP T LK R ++ GKA+ Q WRP S T+S E D+K I S
Sbjct: 735 SSSENRFTFGTSQSIDPKTVLKTRFSDDGKAAFQCQRAWRPNSLITLSAEYDSKKIIGSP 914
Query: 266 AKVGLALALKP 276
AK GLAL+LKP
Sbjct: 915 AKFGLALSLKP 947
>TC84272 weakly similar to PIR|S46959|S46959 porin I 36K - potato, partial
(7%)
Length = 759
Score = 152 bits (383), Expect = 2e-37
Identities = 95/269 (35%), Positives = 137/269 (50%), Gaps = 6/269 (2%)
Frame = +2
Query: 1 MAKGPGLYTDIGKKARDLLFKDYHSDQKFTITTFSPTGVAITSSGTKKGELFVADVNTQL 60
M+K PG+Y+++G A+D+L+KDY + S F+
Sbjct: 62 MSKSPGVYSNLGDNAKDVLYKDY-----------------VNQSPIHFHYQFM------- 169
Query: 61 KNKNITTDIKVDTDSNLFTTIAVNDPAPGLKAIFSFKVPDQRSGKVELQYLHDYAGISSS 120
D N + V + PG + +F VPD SGKVELQYL+ + GI+
Sbjct: 170 -------------DWNAGFSCKVQEILPGFRTVFKCTVPD--SGKVELQYLNRFTGITGC 304
Query: 121 IGLTAN------PIVNFSGVIGTNVLALGADLTFDTKIGELTKSNAGLSFTKDDLIAALT 174
IGL N P+V SG++GT++L+LGA++ +TK AG L A+LT
Sbjct: 305 IGLLGNEEGAYDPVVKLSGLLGTSILSLGANVALHIPTRSITKLTAGFGLNTAFLEASLT 484
Query: 175 LNDKGDALNASYYHVVSPLTNTAVGAEVTHRFSTNENTLTLGTQHALDPLTTLKARVNNF 234
L+D D L A+ YH V+PLT TA+ +V H S E +T+G QHA P T LKAR ++
Sbjct: 485 LHDSFDTLKATIYHEVNPLTQTAIATQVKHSLSLKETGVTIGAQHAFFPETLLKARFDSS 664
Query: 235 GKASALIQHEWRPKSFFTISGEVDTKAIE 263
GK ALIQ + + F T++ E+D A +
Sbjct: 665 GKVGALIQQGFWQRFFVTMASEIDFGATD 751
>TC81600 weakly similar to SP|P07144|PORI_NEUCR Outer mitochondrial membrane
protein porin. {Neurospora crassa}, partial (10%)
Length = 963
Score = 76.6 bits (187), Expect = 9e-15
Identities = 73/287 (25%), Positives = 122/287 (42%), Gaps = 13/287 (4%)
Frame = +3
Query: 2 AKGPGLYTDIGKKARDLLFKDYHSDQ-KFTITTFSPTGVAITSSGTKKGELFVADVNTQL 60
A P Y+DIGK A DL KD+ K T +P GV+ T +G K + ++N +L
Sbjct: 24 APTPATYSDIGKSANDLFSKDFPVGSFKLESKTTAPNGVSFTLTGLKDTKS--GNINGEL 197
Query: 61 KNKNITTDIKVDTDSNLFTTIAV--------NDPAPGLKAIFSFKV-PDQRSGKVELQYL 111
K+K + T + +TT + N A GLK + P + V +
Sbjct: 198 KSKYYYPK-RGGTFTGSWTTSNILNGQVELENSLAKGLKLDLVANIQPASGNRNVRGGFT 374
Query: 112 HDYAGISSSIGLTANPIVNFSG--VIGTNVLALGADLTFDTKIGELTKSNAGLSFTKDDL 169
+ + I L A F G V+G +G++ +D G+LT+ +T +
Sbjct: 375 FKQPALHTKIYLDALKGPTFQGDIVVGHEGFLIGSEGCYDVLEGKLTRYGVTAGYTHPEY 554
Query: 170 IAALTLNDKGDALNASYYHVVSPLTNTAVGAEVTHRFSTNENTLTLGTQHALDPLTTLKA 229
AL + + A YYH VSP A + + L G ++ LD +K
Sbjct: 555 NVALRVTNSLSAYTLLYYHRVSPEIEAGAKAHWDSQTNNPSVNLEFGAKYFLDRDAFIKG 734
Query: 230 RVNNFGKASALIQHEWRPKSFFTISGEVDTKAIEKSA-KVGLALALK 275
+V++ GK RP ++ G +DT + ++A + G+++ L+
Sbjct: 735 KVDSTGKVGIGYTQALRPGVKISLGGSIDTSRLNENAHRFGISINLE 875
>BI270961 similar to GP|9757871|dbj| porin-like protein {Arabidopsis
thaliana}, partial (6%)
Length = 494
Score = 51.2 bits (121), Expect(2) = 4e-12
Identities = 24/45 (53%), Positives = 30/45 (66%)
Frame = +2
Query: 232 NNFGKASALIQHEWRPKSFFTISGEVDTKAIEKSAKVGLALALKP 276
NN G A++QH+ KSF TISG +TKA+E S K G +L LKP
Sbjct: 86 NNHGNLEAVLQHDLTHKSFLTISGAFETKALENSPKFGFSLLLKP 220
Score = 36.6 bits (83), Expect(2) = 4e-12
Identities = 15/26 (57%), Positives = 20/26 (76%)
Frame = +3
Query: 205 RFSTNENTLTLGTQHALDPLTTLKAR 230
RFSTNENTLT+G + +DP TT + +
Sbjct: 3 RFSTNENTLTVGCSYVVDPQTTCEGK 80
>AW684657 similar to GP|9757871|dbj| porin-like protein {Arabidopsis
thaliana}, partial (14%)
Length = 548
Score = 66.2 bits (160), Expect = 1e-11
Identities = 29/44 (65%), Positives = 39/44 (87%)
Frame = +2
Query: 1 MAKGPGLYTDIGKKARDLLFKDYHSDQKFTITTFSPTGVAITSS 44
M+KGPGLY+DIGKKA+DLL KDY+SDQK T++++S TGV +S+
Sbjct: 86 MSKGPGLYSDIGKKAKDLLTKDYNSDQKLTVSSYSTTGVVSSSN 217
>AW584306 weakly similar to PIR|T09116|T091 voltage-dependent anion channel
protein - spinach, partial (19%)
Length = 185
Score = 58.2 bits (139), Expect = 3e-09
Identities = 28/56 (50%), Positives = 41/56 (73%)
Frame = +3
Query: 1 MAKGPGLYTDIGKKARDLLFKDYHSDQKFTITTFSPTGVAITSSGTKKGELFVADV 56
+ +GPGL++ I ++AR LLF+DY S FT+T+ S +GV I+SSG +G+ F ADV
Sbjct: 18 VVRGPGLFSYIRERARYLLFRDYQSAHVFTLTSHSLSGVDISSSGLGRGDAFFADV 185
>BF648198
Length = 663
Score = 28.9 bits (63), Expect = 2.2
Identities = 21/76 (27%), Positives = 34/76 (44%)
Frame = +1
Query: 128 IVNFSGVIGTNVLALGADLTFDTKIGELTKSNAGLSFTKDDLIAALTLNDKGDALNASYY 187
+V+ G+I +L L +TK + SFT D + T+ D LNAS
Sbjct: 277 LVSLLGIIVALILVLQXSELPNTKFFSSLTARIK-SFTLDTPLVNSTIGDNNVKLNASNS 453
Query: 188 HVVSPLTNTAVGAEVT 203
+ +PL N A+ + +
Sbjct: 454 NSTNPLQNIAISPQAS 501
>TC87488
Length = 1591
Score = 28.5 bits (62), Expect = 2.9
Identities = 15/41 (36%), Positives = 20/41 (48%)
Frame = +3
Query: 3 KGPGLYTDIGKKARDLLFKDYHSDQKFTITTFSPTGVAITS 43
K PGLY + K RD F D+ + QKF S G + +
Sbjct: 333 KPPGLYVENIKSGRDSYFLDHVNGQKFRFGMPSKMGTLLNT 455
>TC77342 similar to GP|13785211|emb|CAC37357. putative membrane protein
{Solanum tuberosum}, partial (47%)
Length = 1581
Score = 28.1 bits (61), Expect = 3.8
Identities = 42/169 (24%), Positives = 69/169 (39%), Gaps = 28/169 (16%)
Frame = +2
Query: 100 DQRSGKVELQYLHDYAGISSS---IGLTANPIVNFS-GVIGTNVL-----ALGADLTFDT 150
D+ SGK+++ Y+H GI+++ + NP N +IG L A G+ + +
Sbjct: 272 DETSGKLDIAYIH--TGITATNRWVAWAINPSRNLDPAMIGAQALVAIPQASGSPKAYTS 445
Query: 151 KI---------GELTKSNAGLSFTKDD----LIAALTLNDKGDALNASYYHV------VS 191
I G ++ +GLS T + + A LTL + S+ HV S
Sbjct: 446 NIIDTSTRLQEGTISYPVSGLSATYQNNEVTIFATLTLPNG----TTSFVHVWQDGVLSS 613
Query: 192 PLTNTAVGAEVTHRFSTNENTLTLGTQHALDPLTTLKARVNNFGKASAL 240
T E +H+ S L GT A + + + R N G +A+
Sbjct: 614 DSTPQEHSHESSHQNSKEVLDLVSGTSQAASGIGSRQRRRNTHGVLNAI 760
>BI263584 similar to SP|Q9P6C8|ADH1 Alcohol dehydrogenase I (EC 1.1.1.1).
{Neurospora crassa}, partial (57%)
Length = 682
Score = 27.3 bits (59), Expect = 6.5
Identities = 20/66 (30%), Positives = 31/66 (46%), Gaps = 8/66 (12%)
Frame = +2
Query: 129 VNFSGVIGTNVLALGADLTFDTKIGEL-TKSNAGLSFTKDDLIAALTLNDK-------GD 180
V FSGV T++ A D DTK+ + AG+ + DL+ + + +K G
Sbjct: 182 VKFSGVCHTDLHAWQGDWPLDTKLPLVGGHEGAGVVVARGDLVKDVKIGEKVGIKWLNGS 361
Query: 181 ALNASY 186
L+ SY
Sbjct: 362 CLSCSY 379
>TC78720 annexin; annexin [Medicago truncatula]
Length = 1202
Score = 27.3 bits (59), Expect = 6.5
Identities = 19/63 (30%), Positives = 29/63 (45%)
Frame = +2
Query: 151 KIGELTKSNAGLSFTKDDLIAALTLNDKGDALNASYYHVVSPLTNTAVGAEVTHRFSTNE 210
+I ++ K+ G+ +DLI L KGD A Y ++ P AV A V + N
Sbjct: 161 QIQQIRKAYEGIY--NEDLIKRLESEIKGDFEKAVYRWILEPAERDAVLANVAIKSGKNY 334
Query: 211 NTL 213
N +
Sbjct: 335 NVI 343
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.314 0.132 0.367
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 6,235,617
Number of Sequences: 36976
Number of extensions: 67641
Number of successful extensions: 237
Number of sequences better than 10.0: 30
Number of HSP's better than 10.0 without gapping: 230
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 231
length of query: 276
length of database: 9,014,727
effective HSP length: 95
effective length of query: 181
effective length of database: 5,502,007
effective search space: 995863267
effective search space used: 995863267
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (22.0 bits)
S2: 58 (26.9 bits)
Lotus: description of TM0051b.2


