

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0047c.1
(619 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
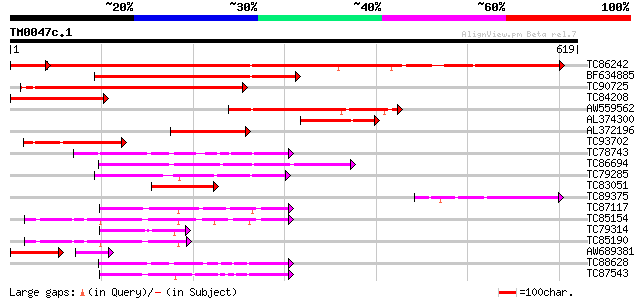
Score E
Sequences producing significant alignments: (bits) Value
TC86242 similar to GP|14486705|gb|AAK63247.1 phosphatidylinosito... 456 e-136
BF634885 similar to GP|11994666|db phosphatidylinositol/phosphat... 305 3e-83
TC90725 similar to GP|14335006|gb|AAK59767.1 AT4g39170/T22F8_70 ... 265 4e-71
TC84208 similar to GP|8778498|gb|AAF79506.1| F20N2.11 {Arabidops... 180 2e-45
AW559562 similar to GP|14486705|gb phosphatidylinositol transfer... 164 1e-40
AL374300 weakly similar to GP|16612283|gb| At2g16380/F16F14.12 {... 135 4e-32
AL372196 homologue to GP|14486705|gb| phosphatidylinositol trans... 122 3e-28
TC93702 similar to GP|16209642|gb|AAL14382.1 At2g21540/F2G1.19 {... 105 5e-23
TC78743 similar to PIR|T05953|T05953 polyphosphoinositide bindin... 82 6e-16
TC86694 similar to PIR|T05949|T05949 phosphatidylinositol-phosph... 78 1e-14
TC79285 similar to PIR|T05949|T05949 phosphatidylinositol-phosph... 76 4e-14
TC83051 similar to GP|21537211|gb|AAM61552.1 thylakoid lumen pro... 75 7e-14
TC89375 74 1e-13
TC87117 similar to GP|8894548|emb|CAB95829.1 hypothetical protei... 69 4e-12
TC85154 similar to GP|8894548|emb|CAB95829.1 hypothetical protei... 67 3e-11
TC79314 similar to GP|16930483|gb|AAL31927.1 AT3g51670/T18N14_50... 66 5e-11
TC85190 similar to GP|8894548|emb|CAB95829.1 hypothetical protei... 62 9e-10
AW689381 weakly similar to GP|11994666|dbj phosphatidylinositol/... 53 1e-09
TC88628 similar to PIR|B86282|B86282 protein F10B6.22 [imported]... 57 3e-08
TC87543 similar to PIR|A96745|A96745 probable cytosolic factor T... 56 5e-08
>TC86242 similar to GP|14486705|gb|AAK63247.1 phosphatidylinositol
transfer-like protein III {Lotus japonicus}, partial
(79%)
Length = 2271
Score = 456 bits (1174), Expect(2) = e-136
Identities = 253/575 (44%), Positives = 358/575 (62%), Gaps = 11/575 (1%)
Frame = +2
Query: 42 KFTHSLKKRG-KRKIDYRVPAVSIEDVRDEGEETVVHELRQRLIERGLLPPRHDDYYTLL 100
KF HSL+K+ +RK R +++IEDVRD E V R LI LLP RHDDY+ LL
Sbjct: 314 KFRHSLRKKSSRRKNPSRSNSLAIEDVRDVKELQAVDAFRHSLISDCLLPSRHDDYHMLL 493
Query: 101 RFLKARDFNIEKTIQMWEEMLLWRKEYGTDTILEEFEFEELEEVLQYYPQGYHGVDKEGR 160
RFLKAR F+IEK MW M+ WRKEYGTDTI+E+FEF EL+EVLQYYP GYHGVDKEGR
Sbjct: 494 RFLKARKFDIEKAKLMWANMIQWRKEYGTDTIMEDFEFSELDEVLQYYPHGYHGVDKEGR 673
Query: 161 PVYIERLGKAHPSRLMRITTIDRYLKYHVQEFERALQEKFPACSIAAKRRIFSTTTILDV 220
PVYIERLGK P++LM++TT++RYLKYHVQ FE+ KFPACSIAAKR I S+TTILDV
Sbjct: 674 PVYIERLGKVDPNKLMQVTTMERYLKYHVQGFEKTFAVKFPACSIAAKRHIDSSTTILDV 853
Query: 221 QGLGMKNFTRTAANLLAAMTKIDTNYYPETLHQMYIVNAGSGFKKMLWPAAQKFCDPKTI 280
G+G KNF++ A L+ + KID +YYPETL +M+I+NAG GF K+LW + F DPKT
Sbjct: 854 HGVGFKNFSKIARELIQRLQKIDGDYYPETLCRMFIINAGPGF-KLLWNTVKTFLDPKTS 1030
Query: 281 AKIQILDSKSLYKLLEVIDSSQLPDFLGGSCTCPTEGGCLKTSKGPWNDPDIIKLVHSAE 340
+KI +L +K KL E+ID S+LP+FLGGSCTC +GGC+++ KGPW DP+I+K+V + E
Sbjct: 1031SKINVLGNKFQSKLFEIIDVSELPEFLGGSCTCIDQGGCMRSDKGPWQDPNILKMVLNGE 1210
Query: 341 ATLVRQLSRASNEQQNF---DSFQVHSLKGRCSDSSTAESGSDINDYSSPTRQRSSTHLR 397
RQ+ SN++ D ++ R SD+STAESGS++ D +SP + T+ R
Sbjct: 1211IQCSRQIVTISNDEGTVIECDKASFPMVQIRSSDTSTAESGSEVEDITSPKASGNYTNPR 1390
Query: 398 LAPVDEEVRVPDLNGYYS----CDDSAPTTEKVVENGQFHLSQKQSLQTNDMENVTHRTI 453
L PV EE R+ G + D+ P +K V+ +++ + T + T + +
Sbjct: 1391LTPVHEEARLAGRAGLSAGLPEYDEYVPMVDKTVD----VTWKEKQVSTKNSHGSTEKYL 1558
Query: 454 SEGTSVNNLFRIIKEIVEKTNHLYVTRVLASFMERLITFIRSLRFEFWRTQNNVHPSNTT 513
S + N +Y+ ++ F + TF RS+ F + + + S++
Sbjct: 1559SRPGGSDG------------NRVYIWAIIIGFFVAIFTFARSIAFRMTKRIKD-NESDSV 1699
Query: 514 EHNINNHSAAVEAA--FERDHILPCA-QRLQRLEKVFEELNNKPYGMPLEKEKMLMDSMD 570
+ I S + A ++ + P A +RL LE+ L +KP MP EKE++L ++
Sbjct: 1700QPIIKEESLPLSPAPILTKEELAPSALERLCELEEKVVMLQSKPNVMPCEKEELLNAAVY 1879
Query: 571 RIKSVEFDLEKTKRVLHSAVKQQLEIADLVENLQK 605
R+ ++E +L TK+ L+ A+ +Q E+ ++ ++
Sbjct: 1880RVDALEAELITTKKALYEALIRQEELMAYIDGQER 1984
Score = 47.8 bits (112), Expect(2) = e-136
Identities = 24/45 (53%), Positives = 33/45 (73%), Gaps = 1/45 (2%)
Frame = +1
Query: 1 YEGQCSTDEIRE-RRSDVENSEDERRSSKIGTLRKKAMNASSKFT 44
+EG DE RE R+SD ENSED+ R ++ G+L+KK +NASSK +
Sbjct: 187 FEGFSGNDERREQRKSDFENSEDDNRRTRNGSLKKKVINASSKIS 321
>BF634885 similar to GP|11994666|db phosphatidylinositol/phosphatidylcholine
transfer protein-like {Arabidopsis thaliana}, partial
(35%)
Length = 684
Score = 305 bits (782), Expect = 3e-83
Identities = 145/225 (64%), Positives = 180/225 (79%)
Frame = +3
Query: 93 HDDYYTLLRFLKARDFNIEKTIQMWEEMLLWRKEYGTDTILEEFEFEELEEVLQYYPQGY 152
HD+Y+T+LRFLK R F+++KT+QMW +ML WRKEYG D+IL++F ++E EEV +YYP G
Sbjct: 9 HDNYHTMLRFLKGRKFDLDKTVQMWADMLHWRKEYGADSILQDFVYKEFEEVQRYYPHGM 188
Query: 153 HGVDKEGRPVYIERLGKAHPSRLMRITTIDRYLKYHVQEFERALQEKFPACSIAAKRRIF 212
HGVDKEGRPVYIERLGK P +LM +TTIDR+L+YHVQ FE+ QEKFPACSIAAKR I
Sbjct: 189 HGVDKEGRPVYIERLGKVDPIKLMNVTTIDRFLRYHVQGFEKLFQEKFPACSIAAKRHID 368
Query: 213 STTTILDVQGLGMKNFTRTAANLLAAMTKIDTNYYPETLHQMYIVNAGSGFKKMLWPAAQ 272
TTTI+DV G+ +F++ A L+ M KID + YPETL+QM+IVNAGSGF K++W A+
Sbjct: 369 KTTTIIDVHGVNWLSFSKIAHELVMRMQKIDGDNYPETLNQMFIVNAGSGF-KLVWNTAK 545
Query: 273 KFCDPKTIAKIQILDSKSLYKLLEVIDSSQLPDFLGGSCTCPTEG 317
F DPKT AKI +L SK +LLEVIDS QLPDFLGGSC+CP +G
Sbjct: 546 GFLDPKTTAKIIVLGSKFQSRLLEVIDSCQLPDFLGGSCSCPNDG 680
>TC90725 similar to GP|14335006|gb|AAK59767.1 AT4g39170/T22F8_70
{Arabidopsis thaliana}, partial (39%)
Length = 831
Score = 265 bits (677), Expect = 4e-71
Identities = 141/250 (56%), Positives = 181/250 (72%), Gaps = 2/250 (0%)
Frame = +1
Query: 12 ERRSDVENSEDERRSSKIGTLRKKAMNASSKFTHSLKK-RGKRKIDYRVPAVSIEDVRDE 70
ERR E+SED++ +IG+L+KKA+NAS K HSLKK R + R ++SIEDVR
Sbjct: 94 ERR---ESSEDDKWK-RIGSLKKKAINASCKLRHSLKKKRSSKSSGSRSNSLSIEDVRQV 261
Query: 71 GEETVVHELRQRLIERGLLPPRHDDYYTLLRFLKARDFNIEKTIQMWEEMLLWRKEYGTD 130
E V RQ L+ LLP HDDY+ LLRFLKAR F+IEK MW M+ WRK+YGTD
Sbjct: 262 EELNAVDAFRQTLLLDNLLPSIHDDYHMLLRFLKARKFDIEKAKLMWANMIQWRKDYGTD 441
Query: 131 TILEEFEFEELEEVLQYYPQGYHGVDKEGRPV-YIERLGKAHPSRLMRITTIDRYLKYHV 189
TI+E+F+F+EL+EV +Y G+ + YIERLGK P+RLM++TT++RYL+YHV
Sbjct: 442 TIIEDFDFKELQEVQKYLSTWLSWCG*GGKTLFYIERLGKVDPNRLMQVTTMERYLRYHV 621
Query: 190 QEFERALQEKFPACSIAAKRRIFSTTTILDVQGLGMKNFTRTAANLLAAMTKIDTNYYPE 249
Q FE+ KFPACSIAAKRRI S+TTILDVQG+G KNFT++A L+ + KID++YYPE
Sbjct: 622 QGFEKTFSVKFPACSIAAKRRIGSSTTILDVQGVGFKNFTKSARELIIQLQKIDSDYYPE 801
Query: 250 TLHQMYIVNA 259
TLHQM+I+NA
Sbjct: 802 TLHQMFIINA 831
>TC84208 similar to GP|8778498|gb|AAF79506.1| F20N2.11 {Arabidopsis
thaliana}, partial (13%)
Length = 664
Score = 180 bits (456), Expect = 2e-45
Identities = 92/109 (84%), Positives = 98/109 (89%), Gaps = 1/109 (0%)
Frame = +1
Query: 1 YEGQCSTDEIRERRSDVENSEDERRS-SKIGTLRKKAMNASSKFTHSLKKRGKRKIDYRV 59
+EGQCS DEIRERR DVE SED+RR SKIGTLRKKAMNASSKFTHSLKKRGKRKIDYRV
Sbjct: 337 FEGQCSNDEIRERRLDVEYSEDDRRQYSKIGTLRKKAMNASSKFTHSLKKRGKRKIDYRV 516
Query: 60 PAVSIEDVRDEGEETVVHELRQRLIERGLLPPRHDDYYTLLRFLKARDF 108
P+V+IEDVRD EET V ELRQRL+ERG LP RHDDY+TLLRFLKARDF
Sbjct: 517 PSVAIEDVRDAREETAVLELRQRLVERGSLPSRHDDYHTLLRFLKARDF 663
>AW559562 similar to GP|14486705|gb phosphatidylinositol transfer-like
protein III {Lotus japonicus}, partial (34%)
Length = 668
Score = 164 bits (414), Expect = 1e-40
Identities = 86/199 (43%), Positives = 129/199 (64%), Gaps = 9/199 (4%)
Frame = +3
Query: 239 MTKIDTNYYPETLHQMYIVNAGSGFKKMLWPAAQKFCDPKTIAKIQILDSKSLYKLLEVI 298
+ K+D + Y ET QM+I+NAG GF+ +LW + F DPKT +KI +L +K KLLEVI
Sbjct: 12 LQKVDCDNYFETFCQMFIINAGPGFR-LLWSTVKSFLDPKTTSKIHVLGNKYQSKLLEVI 188
Query: 299 DSSQLPDFLGGSCTCPTEGGCLKTSKGPWNDPDIIKLVHSAEATLVRQLSRASNEQQNFD 358
D+S+LP+FLGG+C+C EGGCL++ KGPW +P+I+K+V + E RQ+ + N +
Sbjct: 189 DASELPEFLGGTCSCADEGGCLRSDKGPWKNPEILKMVLNGEPRRARQVVKVLNSEGKVI 368
Query: 359 SF---QVHSLKGRCSDSSTAESGSDINDYSSPTRQRSSTHLRLAPVDEEVRV------PD 409
++ + +KG SD+STAESGS+ D +SP +S +HLRL PV EE ++ +
Sbjct: 369 AYAKPRYPMVKG--SDTSTAESGSEAEDIASPKAMKSYSHLRLTPVREEAKIAGKASYAN 542
Query: 410 LNGYYSCDDSAPTTEKVVE 428
++GY D+ P +K V+
Sbjct: 543 MSGY---DEYVPMVDKPVD 590
>AL374300 weakly similar to GP|16612283|gb| At2g16380/F16F14.12 {Arabidopsis
thaliana}, partial (4%)
Length = 258
Score = 135 bits (341), Expect = 4e-32
Identities = 66/86 (76%), Positives = 76/86 (87%)
Frame = +2
Query: 318 GCLKTSKGPWNDPDIIKLVHSAEATLVRQLSRASNEQQNFDSFQVHSLKGRCSDSSTAES 377
GCL+++KGPWNDPDI+KL +AEAT VRQ++RASNEQ NFDSFQ+HSLKGRCSDSS AES
Sbjct: 2 GCLRSNKGPWNDPDIMKLSXNAEATFVRQITRASNEQNNFDSFQLHSLKGRCSDSS-AES 178
Query: 378 GSDINDYSSPTRQRSSTHLRLAPVDE 403
GSD NDYSSPTRQR ++ RLAPV E
Sbjct: 179 GSDFNDYSSPTRQRRCSYPRLAPVCE 256
>AL372196 homologue to GP|14486705|gb| phosphatidylinositol transfer-like
protein III {Lotus japonicus}, partial (14%)
Length = 264
Score = 122 bits (307), Expect = 3e-28
Identities = 56/88 (63%), Positives = 74/88 (83%)
Frame = +1
Query: 176 MRITTIDRYLKYHVQEFERALQEKFPACSIAAKRRIFSTTTILDVQGLGMKNFTRTAANL 235
M++TT+DRY+KYHVQEFE++ KFPAC+IAAKR I S+TTILDVQG+G+KNFT++A L
Sbjct: 1 MQVTTMDRYVKYHVQEFEKSFAIKFPACTIAAKRHIDSSTTILDVQGVGLKNFTKSAREL 180
Query: 236 LAAMTKIDTNYYPETLHQMYIVNAGSGF 263
+ + K+D + YPETL QM+I+NAG GF
Sbjct: 181 IQRLQKVDGDNYPETLCQMFIINAGPGF 264
>TC93702 similar to GP|16209642|gb|AAL14382.1 At2g21540/F2G1.19 {Arabidopsis
thaliana}, partial (19%)
Length = 473
Score = 105 bits (262), Expect = 5e-23
Identities = 55/112 (49%), Positives = 80/112 (71%)
Frame = +1
Query: 16 DVENSEDERRSSKIGTLRKKAMNASSKFTHSLKKRGKRKIDYRVPAVSIEDVRDEGEETV 75
++E+SEDE++ K+G+++K A++ASSKF +S K+G++ RV ++ IED D E
Sbjct: 145 EMEHSEDEKKK-KVGSIKKVALSASSKFKNSFTKKGRKHS--RVMSICIEDSFDAEELQA 315
Query: 76 VHELRQRLIERGLLPPRHDDYYTLLRFLKARDFNIEKTIQMWEEMLLWRKEY 127
V LRQ LI LLP +HDD + +LRFL+AR ++IEKT QMW +ML WRKE+
Sbjct: 316 VDALRQTLILEELLPSKHDDPHMMLRFLRARKYDIEKTKQMWTDMLKWRKEF 471
>TC78743 similar to PIR|T05953|T05953 polyphosphoinositide binding protein
Ssh2 - soybean, partial (94%)
Length = 1232
Score = 82.0 bits (201), Expect = 6e-16
Identities = 71/241 (29%), Positives = 112/241 (46%), Gaps = 1/241 (0%)
Frame = +1
Query: 70 EGEETVVHELRQRLIERGLLPPRHDDYYTLLRFLKARDFNIEKTIQMWEEMLLWRKEYGT 129
E E T ++ +R + R DD + RFL+ARD +++K M+ + + WRK +
Sbjct: 322 EAELTKINLMRTLVESRDPSSKEVDDLM-IRRFLRARDLDVDKASAMFLKYMKWRKSFVP 498
Query: 130 DTILEEFEFEELEEVLQYYPQGYHGVDKEGRPVYIERLGKAHPSRLMRITTIDRYLKYHV 189
+ E + + Y Q G+DK+GRP+ + K ++ +D + +Y V
Sbjct: 499 SGSVSPSEIADDLAQEKIYVQ---GLDKKGRPIIVAFAAKHFQNK----NGLDAFKRYVV 657
Query: 190 QEFERALQEKFPACSIAAKRRIFSTTTILDVQGLGMKNFTRTAANLLAAMTKIDTNYYPE 249
E+ + P +I D++G G N L A+T I +YYPE
Sbjct: 658 FALEKLISRMPPGEE--------KFVSIADIKGWGYAN--SDIRGYLGALT-ILQDYYPE 804
Query: 250 TLHQMYIVNAGSGFKKMLWPAAQKFCDPKTIAKIQILDSKSL-YKLLEVIDSSQLPDFLG 308
L +++IV+A F K +W F D T KI +++K L LLE ID SQLP+ G
Sbjct: 805 RLGKLFIVHAPYMFMK-VWKIIYPFIDDNTKKKIVFVENKKLKATLLEEIDESQLPEIYG 981
Query: 309 G 309
G
Sbjct: 982 G 984
>TC86694 similar to PIR|T05949|T05949
phosphatidylinositol-phosphatidylcholine transfer
protein SEC14 homolog Ssh1 - soybean, partial (98%)
Length = 1433
Score = 77.8 bits (190), Expect = 1e-14
Identities = 72/284 (25%), Positives = 122/284 (42%), Gaps = 4/284 (1%)
Frame = +3
Query: 98 TLLRFLKARDFNIEKTIQMWEEMLLWRKEYGTDTILEE--FEFEELEEVLQYYPQGYHGV 155
TL+RFLKAR++N K +M + L WR + D IL + + + G G
Sbjct: 288 TLIRFLKAREWNASKAHKMLIDSLNWRVQNEIDKILSKPIIPQDLYRGLRDSQLIGLSGY 467
Query: 156 DKEGRPVYIERLGKAHPSRLMRITTIDRYLKYHVQEFERALQEKFPACSIAAKRRIFSTT 215
+EG PV+ +G + + ++ Y++ H+Q E + P+ S R I +
Sbjct: 468 SREGLPVFAIGVGLSTFDK----ASVHYYVQSHIQINEYRDRVILPSASKKHGRPITTCV 635
Query: 216 TILDVQGLGMKNFTRTAANLLAAMTKIDTNYYPETLHQMYIVNAGSGFKKMLWPAAQKFC 275
+LD+ GL + + LL ++ ID YPE + YIVNA F W +
Sbjct: 636 KVLDMTGLKLSALNQ--IKLLTIISSIDDLNYPEKTNTYYIVNAPYIFSG-CWKVVKPLL 806
Query: 276 DPKTIAKIQILDSKSLYKLLEVIDSSQLPDFLGGSCTCPTEGGCLKTSKGPWNDPDIIKL 335
+T K+Q+L +LL+++D + LP F C G + G N +
Sbjct: 807 QERTRKKVQVLQGCGRDELLKIMDYACLPHF----CKKEGSGSSKHSGSGSENCYSLDHP 974
Query: 336 VHSAEATLVRQLSRASNEQQ--NFDSFQVHSLKGRCSDSSTAES 377
H +++ SR + +++ SF V + D A++
Sbjct: 975 FHQELYNYIKEQSRMNEDRKPIKHGSFHVEFPEPSADDGEIAKT 1106
>TC79285 similar to PIR|T05949|T05949
phosphatidylinositol-phosphatidylcholine transfer
protein SEC14 homolog Ssh1 - soybean, partial (68%)
Length = 842
Score = 75.9 bits (185), Expect = 4e-14
Identities = 70/222 (31%), Positives = 98/222 (43%), Gaps = 8/222 (3%)
Frame = +3
Query: 93 HDDYYT--LLRFLKARDFNIEKTIQMWEEMLLWRKEYGTDTIL-EEFEFEELEEVLQYYP 149
H Y T L RFLKARD N+ K +M + L WR E D +L + + + V
Sbjct: 210 HQGYPTEMLARFLKARDGNVAKAQKMLIDCLHWRVENEIDKVLAKPIPADLYKPVRDSQL 389
Query: 150 QGYHGVDKEGRPVYIERLGKAHPSRLMRITTIDR-----YLKYHVQEFERALQEKFPACS 204
G G KEG PV +G ++T D+ Y++ H+Q E + P +
Sbjct: 390 IGMSGYTKEGLPVIAVGVG---------LSTYDKASDKYYIQSHIQVNEYRDRVILPTAT 542
Query: 205 IAAKRRIFSTTTILDVQGLGMKNFTRTAANLLAAMTKIDTNYYPETLHQMYIVNAGSGFK 264
R I + +LD+ GL K LL A++ ID YPE YIVNA F
Sbjct: 543 KKHGRYIGTCVKVLDMTGL--KFSALNQLRLLTAISTIDDLNYPEKTDIYYIVNAPYVF- 713
Query: 265 KMLWPAAQKFCDPKTIAKIQILDSKSLYKLLEVIDSSQLPDF 306
W + +T KIQ+L +LL+V+D + LP F
Sbjct: 714 SACWKVVKPLLQERTRKKIQVLQGCGKEELLKVMDYAFLPHF 839
>TC83051 similar to GP|21537211|gb|AAM61552.1 thylakoid lumen protein
chloroplast precursor {Arabidopsis thaliana}, partial
(56%)
Length = 915
Score = 75.1 bits (183), Expect = 7e-14
Identities = 36/74 (48%), Positives = 53/74 (70%)
Frame = +2
Query: 155 VDKEGRPVYIERLGKAHPSRLMRITTIDRYLKYHVQEFERALQEKFPACSIAAKRRIFST 214
+ ++ R VYIER+G ++L +ITT +R +K+HV E E+ L+ ++PACS+AAKR I ST
Sbjct: 659 IQRQDRSVYIERIGMIDVNKLWQITTQERLIKHHVSEQEKTLRVRYPACSLAAKRHIAST 838
Query: 215 TTILDVQGLGMKNF 228
T+ILD G+ K F
Sbjct: 839 TSILDGNGVVSKFF 880
>TC89375
Length = 1660
Score = 74.3 bits (181), Expect = 1e-13
Identities = 55/165 (33%), Positives = 88/165 (53%), Gaps = 3/165 (1%)
Frame = +2
Query: 443 NDMENVTHRTISEGTSVNNLFRIIKEI---VEKTNHLYVTRVLASFMERLITFIRSLRFE 499
ND + V + G+S NN+ R K++ V T + +VLA + F F
Sbjct: 134 NDAKEVGDVDLPRGSS-NNISRQPKKLISYVRSTLDQIIVKVLACIY---VEFAALGNFF 301
Query: 500 FWRTQNNVHPSNTTEHNINNHSAAVEAAFERDHILPCAQRLQRLEKVFEELNNKPYGMPL 559
R+ NN S+ ++ S+++E D P QRL+ LE V E+ NKP +P
Sbjct: 302 VVRSVNNQPRSHQKIQHVE--SSSLEPLTTPDIKEPIWQRLRDLEAVVTEMANKPRTIPR 475
Query: 560 EKEKMLMDSMDRIKSVEFDLEKTKRVLHSAVKQQLEIADLVENLQ 604
EKE +L +S+ RIK +EFDL+KT++ L + +Q E+A+ +E+L+
Sbjct: 476 EKEDILQESLSRIKGIEFDLQKTRKALLATASKQAELAESLESLK 610
>TC87117 similar to GP|8894548|emb|CAB95829.1 hypothetical protein {Cicer
arietinum}, partial (72%)
Length = 1662
Score = 69.3 bits (168), Expect = 4e-12
Identities = 62/221 (28%), Positives = 99/221 (44%), Gaps = 10/221 (4%)
Frame = +3
Query: 99 LLRFLKARDFNIEKTIQMWEEMLLWRKEYGTDTILEEFEFEELEEVLQYYPQGYHGVDKE 158
LL+FL+ARDF +++ M + +LWRKE+G + +++E +ELE+V+ HG DKE
Sbjct: 534 LLKFLRARDFKVKEAFTMIKNTILWRKEFGIEELMDEKLGDELEKVVY-----MHGFDKE 698
Query: 159 GRPVYIERLGKAHPSRLMRITTID-----RYLKYHVQEFERALQE-KFPACSIAAKRRIF 212
G PV G+ L T D +LK+ +Q E++++ F + I
Sbjct: 699 GHPVCYNIYGEFQNKELYNKTFSDEEKRHNFLKWRIQFLEKSIRNLDFNHGGVCT---IV 869
Query: 213 STTTILDVQGLGMKNFTRTAANLLAAMTKIDTNYYPETLHQMYIVNAGSGF---KKMLWP 269
+ D G G + L ++ + YPE + + +N + +M+ P
Sbjct: 870 HVNDLKDSPGPGKWELRQATKQAL----QLFQDNYPEFVAKQVFINVPWWYLAVNRMISP 1037
Query: 270 AAQKFCDPKTIAKIQIL-DSKSLYKLLEVIDSSQLPDFLGG 309
F +T +K SKS LL I QLP GG
Sbjct: 1038----FLTQRTKSKFVFAGPSKSTETLLSYIAPEQLPVKYGG 1148
>TC85154 similar to GP|8894548|emb|CAB95829.1 hypothetical protein {Cicer
arietinum}, partial (83%)
Length = 1811
Score = 66.6 bits (161), Expect = 3e-11
Identities = 75/318 (23%), Positives = 139/318 (43%), Gaps = 25/318 (7%)
Frame = +3
Query: 17 VENSEDERRSSKIGTLRKKAMNASSKFTHSLKKRGKRKIDYRVPAVSIEDVRDEGEETVV 76
VE +E E+ K+ + + K S ++ G + ++ ++ V EGE+ V
Sbjct: 225 VETAEPEKEEKKV---EETVVEVVEKIAASTEEDGAKTVEAIQESIVSVPVT-EGEQPVA 392
Query: 77 HELRQRLIERGLLPPRHDDYY------------TLLRFLKARDFNIEKTIQMWEEMLLWR 124
+ + +E + P + + LL+FL+ARDF +++ M ++ +LWR
Sbjct: 393 EPVAE--VEVTPIVPEEVEIWGIPLLADERSDVILLKFLRARDFKVKEAFTMIKQTVLWR 566
Query: 125 KEYGTDTILEEFEFEELEEVLQYYPQGYHGVDKEGRPVYIERLGKAHPSRLMRITTID-- 182
KE+G + +L+E + ++V+ G DKEG PVY G+ L + T D
Sbjct: 567 KEFGVEALLQEDLGTDWDKVV-----FTDGTDKEGHPVYYNVFGEFEDKDLYQKTFSDEE 731
Query: 183 ---RYLKYHVQEFERALQEKFPACSIAAKRRIFSTTTILDVQG----LGMKNFTRTAANL 235
+++++ +Q E+++++ A S I + I D++ LG K ++
Sbjct: 732 KRTKFVRWWIQSLEKSVRKLDFAPS-----GISTLVQINDLKNSPGLLGKKELRQSIKQT 896
Query: 236 LAAMTKIDTNYYPETLHQMYIVNAG---SGFKKMLWPAAQKFCDPKTIAKIQIL-DSKSL 291
L + + YPE + + +N F +M+ P F +T +K SKS
Sbjct: 897 LQLL----QDNYPEFVAKQIFINVPWWYLAFSRMISP----FLTQRTKSKFVFAGSSKSA 1052
Query: 292 YKLLEVIDSSQLPDFLGG 309
L + I Q+P GG
Sbjct: 1053ETLFKYIAPEQVPVKYGG 1106
>TC79314 similar to GP|16930483|gb|AAL31927.1 AT3g51670/T18N14_50
{Arabidopsis thaliana}, partial (33%)
Length = 868
Score = 65.9 bits (159), Expect = 5e-11
Identities = 42/106 (39%), Positives = 61/106 (56%), Gaps = 7/106 (6%)
Frame = +1
Query: 99 LLRFLKARDFNIEKTIQMWEEMLLWRKEYGTDTILEE--FEFEELEEVLQYYPQGYHGVD 156
LL+FL+ARDF + + + M + L WRKE+ DTI+EE +F+ELE V+ Y QGY D
Sbjct: 496 LLKFLRARDFRVNEALNMITKCLAWRKEFNADTIVEEDLSQFKELEGVIAYM-QGY---D 663
Query: 157 KEGRPVYIERLGKAHPSRLM-RI----TTIDRYLKYHVQEFERALQ 197
KEG PV G + RI + ++L++ VQ ER ++
Sbjct: 664 KEGHPVCYNAYGLFRDKDMYERIFGDEEKLKKFLRWRVQVLERGIR 801
>TC85190 similar to GP|8894548|emb|CAB95829.1 hypothetical protein {Cicer
arietinum}, partial (50%)
Length = 1496
Score = 61.6 bits (148), Expect = 9e-10
Identities = 47/199 (23%), Positives = 95/199 (47%), Gaps = 17/199 (8%)
Frame = +3
Query: 17 VENSEDERRSSKIGTLRKKAMNASSKFTHSLKKRGKRKIDYRVPAVSIEDVRDEGEETVV 76
VE +E E+ K+ + + K S ++ G + ++ ++ V EGE+ V
Sbjct: 681 VETAEPEKEEKKV---EETVVEVVEKIAASTEEDGAKTVEAIQESIVSVPVT-EGEQPVA 848
Query: 77 HELRQRLIERGLLPPRHDDYY------------TLLRFLKARDFNIEKTIQMWEEMLLWR 124
+ + +E + P + + LL+FL+ARDF +++ M ++ +LWR
Sbjct: 849 EPVAE--VEVTPIVPEEVEIWGIPLLADERSDVILLKFLRARDFKVKEAFTMIKQTVLWR 1022
Query: 125 KEYGTDTILEEFEFEELEEVLQYYPQGYHGVDKEGRPVYIERLGKAHPSRLMRITTID-- 182
KE+G + +L+E + ++V+ G DKEG PVY G+ L + T D
Sbjct: 1023KEFGVEALLQEDLGTDWDKVV-----FTDGTDKEGHPVYYNVFGEFEDKDLYQKTFSDEE 1187
Query: 183 ---RYLKYHVQEFERALQE 198
+++++ +Q E+++++
Sbjct: 1188KRTKFVRWWIQSLEKSVRK 1244
>AW689381 weakly similar to GP|11994666|dbj
phosphatidylinositol/phosphatidylcholine transfer
protein-like {Arabidopsis thaliana}, partial (7%)
Length = 654
Score = 52.8 bits (125), Expect(2) = 1e-09
Identities = 26/57 (45%), Positives = 38/57 (66%)
Frame = +1
Query: 2 EGQCSTDEIRERRSDVENSEDERRSSKIGTLRKKAMNASSKFTHSLKKRGKRKIDYR 58
EG ++ R R + E SEDE R S+ +LR++AM AS++ T+SL+KR KR DY+
Sbjct: 316 EGLLVQEDERSRSLEPETSEDEWRKSRARSLRRRAMTASTRLTYSLRKRNKRVADYQ 486
Score = 28.1 bits (61), Expect(2) = 1e-09
Identities = 15/41 (36%), Positives = 20/41 (48%)
Frame = +3
Query: 73 ETVVHELRQRLIERGLLPPRHDDYYTLLRFLKARDFNIEKT 113
E V+ R L+ R LLP HD+ FLK + + KT
Sbjct: 531 EEAVNSFRLALLTRDLLPDSHDNITQC*GFLKGXNLTLIKT 653
>TC88628 similar to PIR|B86282|B86282 protein F10B6.22 [imported] -
Arabidopsis thaliana, partial (27%)
Length = 1087
Score = 56.6 bits (135), Expect = 3e-08
Identities = 55/214 (25%), Positives = 96/214 (44%), Gaps = 2/214 (0%)
Frame = +1
Query: 98 TLLRFLKARDFNIEKTIQMWEEMLLWRKE-YGTDTILEEFEFEELEEVLQYYPQGYHGVD 156
TL+RFL AR + +K +M+ + WR D + + E + E + + Q G+
Sbjct: 190 TLMRFLIARSMDSDKAAKMFVQWQKWRATMVPNDGFISDSEVPDELETRKIFLQ---GLS 360
Query: 157 KEGRPVYIERLGKAHPSRLMRITTIDRYLKYHVQEFERALQEKFPACSIAAKRRIFSTTT 216
K+ PV I + + PS+ ++ K+ V ++ + F + ++ I
Sbjct: 361 KDKYPVMIVQASRHFPSK-----DQIQFKKFIVHLLDKTIASAFKGREVGNEKLI----G 513
Query: 217 ILDVQGLGMKNFTRTAANLLAAMTKIDTNYYPETLHQMYIVNAGSGFKKMLWPAAQKFCD 276
+LD+QG+ KN A L+ + +YYPE L + YI++ F +W F D
Sbjct: 514 VLDLQGISYKNV--DARGLITGFQFLQ-SYYPECLAKCYILHM-PWFFVSVWRFVSGFLD 681
Query: 277 PKTIAKIQILDSKSLYKL-LEVIDSSQLPDFLGG 309
T KI I+ ++ KL + + LP+ GG
Sbjct: 682 KATQEKIVIISNEEEKKLFVSEVGEDILPEEYGG 783
>TC87543 similar to PIR|A96745|A96745 probable cytosolic factor T9N14.8
[imported] - Arabidopsis thaliana, partial (74%)
Length = 1718
Score = 55.8 bits (133), Expect = 5e-08
Identities = 56/217 (25%), Positives = 96/217 (43%), Gaps = 6/217 (2%)
Frame = +1
Query: 99 LLRFLKARDFNIEKTIQMWEEMLLWRKEYGTDTILEEFEFEELEEVLQYYPQGYHGVDKE 158
LL+FL+ARDF +++ M + L WRK + D +L+E ++L++V+ HG +E
Sbjct: 514 LLKFLRARDFKPKESHTMLKNTLQWRKSFNIDALLDEDLGDDLDKVV-----FMHGFSRE 678
Query: 159 GRPVYIERLGKAHPSRLMRIT-----TIDRYLKYHVQEFERALQEKFPACSIAAKRRIFS 213
G PV G+ L T +R+L++ VQ E+++++ S +F
Sbjct: 679 GHPVCYNVYGEFQNKELYEKTFGSEEKRERFLRWRVQFLEKSIRKL--DFSPGGVNTLFQ 852
Query: 214 TTTILDVQGLGMKNFTRTAANLLAAMTKIDTNYYPETLHQMYIVNAGSGFKKMLWPAAQK 273
+ + G K R A + A+ + N YPE + + +N + +
Sbjct: 853 VNDLKNSPGPAKKEL-RVATKM--ALELLQDN-YPEFVAKQVFINV-PWWYLAFYTILNP 1017
Query: 274 FCDPKTIAKIQIL-DSKSLYKLLEVIDSSQLPDFLGG 309
F +T +K SKS L + I Q+P GG
Sbjct: 1018FLTQRTKSKFVFAGTSKSPDTLFKYITPEQVPVQYGG 1128
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.317 0.133 0.382
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 17,581,926
Number of Sequences: 36976
Number of extensions: 222333
Number of successful extensions: 1316
Number of sequences better than 10.0: 63
Number of HSP's better than 10.0 without gapping: 1282
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1294
length of query: 619
length of database: 9,014,727
effective HSP length: 102
effective length of query: 517
effective length of database: 5,243,175
effective search space: 2710721475
effective search space used: 2710721475
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 61 (28.1 bits)
Lotus: description of TM0047c.1


