

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
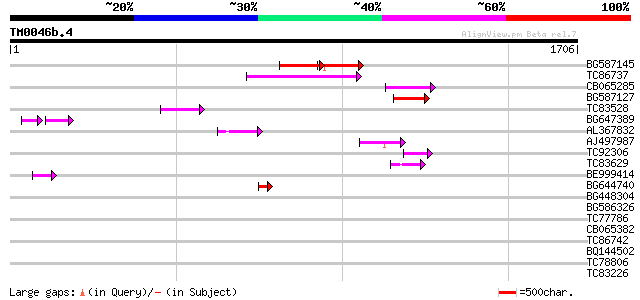
Query= TM0046b.4
(1706 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
BG587145 similar to PIR|H86337|H8 protein F5M15.26 [imported] - ... 173 4e-43
TC86737 weakly similar to GP|6683624|dbj|BAA89272.1 Pol {Alterna... 152 1e-36
CB065285 weakly similar to PIR|A84500|A84 probable retroelement ... 104 3e-22
BG587127 weakly similar to PIR|H84506|H84 probable retroelement ... 97 5e-20
TC83528 92 1e-18
BG647389 57 2e-15
AL367832 weakly similar to GP|14091845|gb Putative retroelement ... 79 2e-14
AJ497987 weakly similar to GP|9927273|dbj Similar to Arabidopsis... 66 1e-10
TC92306 53 9e-07
TC83629 weakly similar to GP|6692261|gb|AAF24611.1| putative RNa... 52 3e-06
BE999414 48 3e-05
BG644740 similar to PIR|A84460|A84 probable retroelement pol pol... 48 4e-05
BG448304 42 0.002
BG586326 similar to PIR|G84493|G8 probable retroelement pol poly... 39 0.017
TC77786 39 0.017
CB065382 similar to PIR|T09671|T096 RPE15 protein - alfalfa (fra... 38 0.038
TC86742 similar to PIR|T01541|T01541 hypothetical protein A_IG00... 36 0.15
BQ144502 weakly similar to GP|6523547|emb hydroxyproline-rich gl... 35 0.25
TC78806 similar to GP|8978273|dbj|BAA98164.1 receptor protein ki... 35 0.25
TC83226 weakly similar to PIR|G86419|G86419 probable reverse tra... 33 1.2
>BG587145 similar to PIR|H86337|H8 protein F5M15.26 [imported] - Arabidopsis
thaliana, partial (13%)
Length = 763
Score = 173 bits (439), Expect = 4e-43
Identities = 80/136 (58%), Positives = 106/136 (77%)
Frame = +2
Query: 811 MHPSDEESTTFMTNQANYCYKTMPFGLKNAGATYQRLMDKIFSKQVGRNMEVYVDDMIVK 870
MHP D E T F+T++ YCYK MPFGLKNAG+TYQRL++++F+ ++G MEVY+DDM+VK
Sbjct: 2 MHPDDLEKTAFITDRGTYCYKVMPFGLKNAGSTYQRLVNRMFADKLGNTMEVYIDDMLVK 181
Query: 871 SARASDHGGDLKEAFDQLRTYQMKLNPEKCSFGIQGGKFLGFMLTSRGIEVNPDKGRAIL 930
S RA+DH LKE F L Y MKLNP KC+FG+ G+FLG+++T +GIEVNP + AIL
Sbjct: 182 SLRATDHLNHLKE*FKTLDEYIMKLNPAKCTFGVTSGEFLGYIVTQQGIEVNPKQITAIL 361
Query: 931 EMKSPTSVKEVQRLTG 946
++ SP + +EVQRLTG
Sbjct: 362 DLPSPKNSREVQRLTG 409
Score = 110 bits (274), Expect = 6e-24
Identities = 60/146 (41%), Positives = 88/146 (60%), Gaps = 7/146 (4%)
Frame = +3
Query: 925 KGRAILEMKSPTSVKEVQRLT-------GRMAALSRFLPMAGDKAAPFFTCLKKNSKFQW 977
+ R IL P VQR+ GR+AAL+RF+ + DK PF+ L N +F W
Sbjct: 324 ESR*ILSRSPPY*TSLVQRIAERSSDSRGRIAALNRFISRSTDKCLPFYKLLCGNKRFVW 503
Query: 978 TEECEQAFTKLKETLATLPVLSKPTPSVPLVLYLAVTDKAVSTVLLQEEGKKQKVIYFVS 1037
E+CE+AF +LK+ L T PVLSKP L LY+A++ AVS+VL++E+ +QK I++ S
Sbjct: 504 DEKCEEAFEQLKQYLTTPPVLSKPEAGDTLSLYIAISSTAVSSVLIREDRGEQKPIFYTS 683
Query: 1038 HTLQGAELRYQKIEKAALAILKTARR 1063
+ E RY +EK A A++ +AR+
Sbjct: 684 KRMTDPETRYPTLEKMAFAVITSARK 761
>TC86737 weakly similar to GP|6683624|dbj|BAA89272.1 Pol {Alternaria
alternata}, partial (21%)
Length = 1540
Score = 152 bits (384), Expect = 1e-36
Identities = 101/347 (29%), Positives = 165/347 (47%), Gaps = 3/347 (0%)
Frame = +1
Query: 714 PRRRMSEEKNKAVQLETEKLIKARFIREVQYPTWLANVVMVKKANGKWRMCTDYTSLNKV 773
P MS ++ ++ E L+ FI+ A V+ V+K G R C DY +LN +
Sbjct: 475 PLYGMSRQELLVLKKTLEDLLDKGFIKASGSAAG-APVLFVRKPGGGIRFCVDYRALNAI 651
Query: 774 CPKDSYPLPNVDKLVDGASGNELLSLMDAYSGYNQIMMHPSDEESTTFMTNQANYCYKTM 833
KD YPLP + + + +G + +D + ++++ + D+E T F T + +
Sbjct: 652 TKKDRYPLPLISETLRRVAGARWFTKLDVVAAFHKMRIKDEDQEKTAFRTRYGLFEWIVC 831
Query: 834 PFGLKNAGATYQRLMDKIFSKQVGRNMEVYVDDMIV-KSARASDHGGDLKEAFDQLRTYQ 892
PFGL A AT+QR ++K + + + Y+DD+++ + DH ++ +L
Sbjct: 832 PFGLTGAPATFQRYINKTLHEFLDDFVTAYIDDVLIYTTGSKKDHEAQVRRVLRRLADAG 1011
Query: 893 MKLNPEKCSFGIQGGKFLGFMLTS-RGIEVNPDKGRAILEMKSPTSVKEVQRLTGRMAAL 951
+ L+P+KC F + K++GF+LT+ +G+ +P K AI + P SVK + G
Sbjct: 1012 LSLDPKKCEFSVTTVKYVGFILTAGKGVSCDPLKLAAIRDWLPPGSVKGARSFLGFCNYY 1191
Query: 952 SRFLPMAGDKAAPFFTCLKKNSKFQWTEECEQAFTKLKETLATLPVLSKPTPSVPLVLYL 1011
F+P + P +K+ F+W E E AFTKLK A PVL P +
Sbjct: 1192 KDFIPGYSEITEPLTRLTRKDFPFRWGAEQEAAFTKLKRLFAEEPVLRMFDPEAVTTVET 1371
Query: 1012 AVTDKAVSTVLLQEEGK-KQKVIYFVSHTLQGAELRYQKIEKAALAI 1057
+ A+ VL QE+G + F S L AE Y +K LA+
Sbjct: 1372 DCSGFALGGVLTQEDGTGAAHPVAFHSQRLSPAEYNYPIHDKELLAV 1512
>CB065285 weakly similar to PIR|A84500|A84 probable retroelement gag/pol
polyprotein [imported] - Arabidopsis thaliana, partial
(2%)
Length = 592
Score = 104 bits (260), Expect = 3e-22
Identities = 57/155 (36%), Positives = 87/155 (55%), Gaps = 3/155 (1%)
Frame = -2
Query: 1130 EGEKTQGEWVLSVDGSSNNTGSGAGITIESPDKMIIEQSLKFEFKASNNQSEYEALIAGL 1189
EG +W L DG+ N G G G I SP I + + F+ +NN +EYEA I G+
Sbjct: 558 EGPDPNSKWGLVFDGAVNAYGKGIGAVIVSPQGHYIPFTARILFECTNNMAEYEACIFGI 379
Query: 1190 RLAIELGVQKLFIKGDSQLVVKQVKGEYQVKDPQLSKYLEVVRRLMMEVKEIKIEHVPRG 1249
AI++ ++ L I GDS LV+ Q+KGE++ L Y + RRL+ ++++ H+PR
Sbjct: 378 EEAIDMRIKHLDIYGDSALVINQIKGEWETHHANLIPYRDYARRLLTYFTKVELHHIPRD 199
Query: 1250 QNERADVLAKLASTGRLGNYQTV---IQETLPRPS 1281
+N+ AD LA L+S R+ ++ V + L RPS
Sbjct: 198 ENQMADALATLSSMFRVNHWNDVPIIKVQRLERPS 94
>BG587127 weakly similar to PIR|H84506|H84 probable retroelement pol
polyprotein [imported] - Arabidopsis thaliana, partial
(13%)
Length = 415
Score = 97.1 bits (240), Expect = 5e-20
Identities = 48/110 (43%), Positives = 72/110 (64%)
Frame = +3
Query: 1154 GITIESPDKMIIEQSLKFEFKASNNQSEYEALIAGLRLAIELGVQKLFIKGDSQLVVKQV 1213
GI + SP I++QS + EF ASNN++ YEALIAG+RLA L ++ + DSQLV Q
Sbjct: 3 GIRLTSPTNEILKQSFRLEFHASNNETNYEALIAGVRLAHGLKIRNIHAYCDSQLVASQF 182
Query: 1214 KGEYQVKDPQLSKYLEVVRRLMMEVKEIKIEHVPRGQNERADVLAKLAST 1263
GEY+ +D + YL++V++L ++ + +PR +N +AD LA LAS+
Sbjct: 183 SGEYEARDELMDTYLKLVQKLAQKLDYFALTRIPRSENVQADALAALASS 332
>TC83528
Length = 555
Score = 92.4 bits (228), Expect = 1e-18
Identities = 51/138 (36%), Positives = 83/138 (59%), Gaps = 4/138 (2%)
Frame = +3
Query: 453 PQDFAHVIPHDNDPIVVTIRVNN-YVTKKVFLDQGSSADIIYGDAFDRLGLKESDLKPYK 511
P+ + P P+V+ +++NN + +VF++ S DI+Y AF ++ L+ES LKP +
Sbjct: 141 PKKYTFTGPDVALPMVIKLQINNNFSVLRVFVNPMSKVDILYWSAFLKMKLQESMLKPCQ 320
Query: 512 GTLVGFTGDRVSVRGYVEIPTAFGEGEFVKKFQVKYLVLACRAN---YNVLLGRDTLNKV 568
G L G G + V+GY+++ T FG+GE K +V+Y V+ + YNV+LG +L +
Sbjct: 321 GFLKGTFGKGLPVKGYIDLDTTFGKGENTKTIKVRYFVVESPPSVSIYNVVLGWPSLKDL 500
Query: 569 CAVISTAHLTVKYPACNG 586
AV+S A T+KYP +G
Sbjct: 501 KAVLSVAEFTIKYPVGDG 554
>BG647389
Length = 718
Score = 57.0 bits (136), Expect(2) = 2e-15
Identities = 25/84 (29%), Positives = 47/84 (55%)
Frame = +1
Query: 107 KGMAMQWFIRQPPFSIDNFTDLSTKFLTQFSANKTTKATMFDLISIHQQPGEKLKTYMAR 166
K QW++ P FSI + +++ K + QFSA+K K + +L ++ Q P E L Y+
Sbjct: 451 KEATQQWYMNLPRFSITGYQNMTHKLVHQFSASKHCKVSTTNLFNVRQDPNESLPEYLV* 630
Query: 167 FSKMAVQLEDENPDVCLASFKNGL 190
F+ +++ + ++ +F+NGL
Sbjct: 631 FNNATIKVVNPKQELFAGAFQNGL 702
Score = 45.4 bits (106), Expect(2) = 2e-15
Identities = 23/67 (34%), Positives = 40/67 (59%), Gaps = 2/67 (2%)
Frame = +2
Query: 35 QEQRLEAEDVVEF--QPFVPAITRVDIPKHLQTMALDAFSGESDPMEHLRYFNTKMVIGG 92
++ L EDV E QP I + ++L++++L +F +SDP+EH FNT+M + G
Sbjct: 233 EQDELHREDVDELDPQPLSMDIWNAPVLENLKSLSLLSFDDKSDPVEHATTFNTQMAVIG 412
Query: 93 ATDAVKC 99
A +++C
Sbjct: 413 ALKSLRC 433
>AL367832 weakly similar to GP|14091845|gb Putative retroelement {Oryza
sativa}, partial (2%)
Length = 384
Score = 79.0 bits (193), Expect = 2e-14
Identities = 42/134 (31%), Positives = 72/134 (53%)
Frame = -1
Query: 626 QEGFSLDPRDDADDFRPQPLEETKQVQVKDKFLKIGSGLTTEQEDRLITLLGDNLDLFAW 685
QE ++ P + + EE K+ +K+G+ L + ++ LL + LD+FA
Sbjct: 384 QERKAIQPHQEEIELINLGTEENKRE------IKVGAALEEGVKRKIFQLLREYLDIFAC 223
Query: 686 TINDVPGIDPKVITHKLAIRPGATPVIQPRRRMSEEKNKAVQLETEKLIKARFIREVQYP 745
+ D+PG+DPK++ H++ +P PV RR + ++ E +K I A F+ V+YP
Sbjct: 222 SYEDMPGLDPKIVEHRIPTKPECPPVR*KLRRTHPDMALKIKSEVQKQIDAGFLMTVEYP 43
Query: 746 TWLANVVMVKKANG 759
W+AN+V V K +G
Sbjct: 42 EWVANIVPVPKKDG 1
>AJ497987 weakly similar to GP|9927273|dbj Similar to Arabidopsis thaliana
chromosome II BAC F26H6; putative retroelement pol
polyprotein, partial (1%)
Length = 636
Score = 65.9 bits (159), Expect = 1e-10
Identities = 46/167 (27%), Positives = 72/167 (42%), Gaps = 29/167 (17%)
Frame = -2
Query: 1052 KAALAILKTARRLRPYFQSFQVKIKTDV-PLRQVLQKPDLSGRLVSWSVELSEYDIQYEP 1110
K A+ A+RLR Y + + + + P++ + +KP L+GR+ W + LSEYDI+Y
Sbjct: 635 KTCCALAWAAKRLRHYMINHTTWLVSKMDPIKYIFEKPALTGRIARWQMLLSEYDIEYRS 456
Query: 1111 RGQVTVQSLIDFVA----------------------------ELTPTEGEKTQGEWVLSV 1142
+ + L D +A E EG W L
Sbjct: 455 QKAIKGSILADHLAHQPLEDYRPIKFDFPDEEIMYLKMKDCDEPLFGEGPDPDSVWGLIF 276
Query: 1143 DGSSNNTGSGAGITIESPDKMIIEQSLKFEFKASNNQSEYEALIAGL 1189
DG+ N G+G G + +P I + + F +NN +EYEA I G+
Sbjct: 275 DGAVNVYGNGIGAVLLTPKGAHIPFTARLRFDCTNNIAEYEACIMGI 135
>TC92306
Length = 521
Score = 53.1 bits (126), Expect = 9e-07
Identities = 28/85 (32%), Positives = 38/85 (43%)
Frame = +2
Query: 1186 IAGLRLAIELGVQKLFIKGDSQLVVKQVKGEYQVKDPQLSKYLEVVRRLMMEVKEIKIEH 1245
I GL A G + + ++GD Q V KQ G ++V +P L L K + +EH
Sbjct: 2 ILGLNEARNQGYEHVHVRGDFQXVCKQFXGSWKVNNPNLRNLCNXAVELKSNFKSVSVEH 181
Query: 1246 VPRGQNERADVLAKLASTGRLGNYQ 1270
VPRG N AD A G +
Sbjct: 182 VPRGXNXAADAQANRGKNXGAGEVE 256
>TC83629 weakly similar to GP|6692261|gb|AAF24611.1| putative RNase H
{Arabidopsis thaliana}, partial (21%)
Length = 1071
Score = 51.6 bits (122), Expect = 3e-06
Identities = 33/108 (30%), Positives = 54/108 (49%), Gaps = 3/108 (2%)
Frame = +2
Query: 1147 NNTGSGAGITIESPDKMII---EQSLKFEFKASNNQSEYEALIAGLRLAIELGVQKLFIK 1203
N +GAG + + D ++ Q L + K S +EY AL+ GL+ A G + + K
Sbjct: 587 NRGPAGAGALLLAEDGSLLYGFRQGLGHQTKES---AEYRALLLGLKHASMKGFKYVTAK 757
Query: 1204 GDSQLVVKQVKGEYQVKDPQLSKYLEVVRRLMMEVKEIKIEHVPRGQN 1251
GDS+LV+ Q+ +++KD L K L +I+H+ R +N
Sbjct: 758 GDSELVINQILDPWKIKDEHLKKLCAEALELSDNFHSFRIQHISRERN 901
>BE999414
Length = 613
Score = 48.1 bits (113), Expect = 3e-05
Identities = 24/74 (32%), Positives = 38/74 (50%)
Frame = +3
Query: 68 LDAFSGESDPMEHLRYFNTKMVIGGATDAVKCRLLPSTFKGMAMQWFIRQPPFSIDNFTD 127
+D++ G DP EH+ + AVKC+L +T + AM WF SI ++ D
Sbjct: 9 MDSYDGT*DPDEHMENIEVVLTYRSVRGAVKCKLFVTTLRRGAMTWFKNLRRNSIGSWGD 188
Query: 128 LSTKFLTQFSANKT 141
L +F T F+ ++T
Sbjct: 189 LCHEFTTHFTVSRT 230
>BG644740 similar to PIR|A84460|A84 probable retroelement pol polyprotein
[imported] - Arabidopsis thaliana, partial (4%)
Length = 754
Score = 47.8 bits (112), Expect = 4e-05
Identities = 20/41 (48%), Positives = 27/41 (65%)
Frame = -1
Query: 749 ANVVMVKKANGKWRMCTDYTSLNKVCPKDSYPLPNVDKLVD 789
A ++ V+K +G +RMC DY NKV K+ YPLP +D L D
Sbjct: 238 AALLFVRKKDGYFRMCIDYRQFNKVTTKNKYPLPRIDNLFD 116
>BG448304
Length = 637
Score = 42.0 bits (97), Expect = 0.002
Identities = 23/73 (31%), Positives = 35/73 (47%)
Frame = +2
Query: 325 PPPPRRPPANVDTNRWCEYHKALGHTTDNCWNLRREIDRLIKAGHLANFVKDTAAPEVAK 384
P P + D ++C +HK H TD+C +L+ I+ LI+ G L K+ PE
Sbjct: 65 PKTPTQENKGTDKTKYCRFHKCHRHLTDDCIHLKDTIEILIQRGRLNQLTKN-PEPEKQT 241
Query: 385 ITQGDKGKGKEIV 397
+ K K+IV
Sbjct: 242 VKLITDKKNKDIV 280
>BG586326 similar to PIR|G84493|G8 probable retroelement pol polyprotein
[imported] - Arabidopsis thaliana, partial (13%)
Length = 736
Score = 38.9 bits (89), Expect = 0.017
Identities = 32/115 (27%), Positives = 55/115 (47%), Gaps = 4/115 (3%)
Frame = +2
Query: 996 PVLSKPTPSVPLVLYLAVTDKAVS---TVLLQEEGKKQKVIYFVSHTLQGAELRYQKIEK 1052
P+L P L+ Y+ TD +++ VL Q E KVI + S L+ E Y +
Sbjct: 17 PILVLPE----LITYVVYTDASITGLGCVLTQHE----KVIAYASRQLRKHEGNYPTHDL 172
Query: 1053 AALAILKTARRLRPYFQSFQVKIKTD-VPLRQVLQKPDLSGRLVSWSVELSEYDI 1106
A++ + R Y +V+I TD L+ + +P+L+ R W +++YD+
Sbjct: 173 EMAAVVFALKIWRSYLYGAKVQIHTDHKSLKYIFTQPELNLRQRRWMEFVADYDL 337
>TC77786
Length = 1094
Score = 38.9 bits (89), Expect = 0.017
Identities = 17/42 (40%), Positives = 24/42 (56%)
Frame = +3
Query: 333 ANVDTNRWCEYHKALGHTTDNCWNLRREIDRLIKAGHLANFV 374
A D + C YH++ GH T+ C+ LR I+ LIK L F+
Sbjct: 801 ARTDKAK*CCYHRSRGHDTEECFQLREAIEDLIKKVKLGRFI 926
>CB065382 similar to PIR|T09671|T096 RPE15 protein - alfalfa (fragment),
partial (9%)
Length = 624
Score = 37.7 bits (86), Expect = 0.038
Identities = 18/55 (32%), Positives = 29/55 (52%), Gaps = 1/55 (1%)
Frame = -2
Query: 317 GQTNVVQYPPPPRRPPANVDTNRWCEYHK-ALGHTTDNCWNLRREIDRLIKAGHL 370
G + Q PPP P ++ C++H+ ALGH + C+ L+ + +LI G L
Sbjct: 422 GHCTIRQGKPPPDPLPPRFRSDLKCDFHQGALGHDVEGCYALKHIVKKLINQGKL 258
>TC86742 similar to PIR|T01541|T01541 hypothetical protein A_IG005I10.16 -
Arabidopsis thaliana, partial (19%)
Length = 2073
Score = 35.8 bits (81), Expect = 0.15
Identities = 23/83 (27%), Positives = 35/83 (41%)
Frame = +3
Query: 256 AQTPRVPRPGPYQNSKPGFQNTWHRNPQGPPAATAGQTGASPAPVPLTKLNAPLSTILRA 315
+ +P P P P +S P+ T T A P+P P T + +PL+T+L
Sbjct: 702 SSSPSPPSPSPPTSSH--------------PSTTTSSTPAPPSPPPSTTVFSPLNTLL-- 833
Query: 316 VGQTNVVQYPPPPRRPPANVDTN 338
+PPPP P+ T+
Sbjct: 834 --------FPPPPIPSPSQTKTH 878
>BQ144502 weakly similar to GP|6523547|emb hydroxyproline-rich glycoprotein
DZ-HRGP {Volvox carteri f. nagariensis}, partial (39%)
Length = 1358
Score = 35.0 bits (79), Expect = 0.25
Identities = 29/108 (26%), Positives = 41/108 (37%), Gaps = 9/108 (8%)
Frame = -3
Query: 235 KQSQSGGVSQDKSKAGKETRIAQTPRVPRPGPYQNSKPGFQNTWHRNPQGP--------- 285
+++ +Q S+A + P P P ++P Q W P G
Sbjct: 753 RRTLRASAAQHPSRAPRRHPPPPPPPPPSSPPGMGTRPSCQENWWAAPPGARPSRPSRYI 574
Query: 286 PAATAGQTGASPAPVPLTKLNAPLSTILRAVGQTNVVQYPPPPRRPPA 333
P T+ + PAPVP +P + RA PPPRRPPA
Sbjct: 573 PPPTSPPPASPPAPVPP---QSPSTIRGRAPPPA------PPPRRPPA 457
Score = 30.4 bits (67), Expect = 6.1
Identities = 28/97 (28%), Positives = 34/97 (34%), Gaps = 22/97 (22%)
Frame = -1
Query: 259 PRVPR------PGPYQNSKPGFQNTWHRNPQG-----------PPAATAGQTGASPAPVP 301
PR PR PGP + P HR P+ PPAA +P P P
Sbjct: 413 PRAPRQGVAVVPGPRSGNSP---RAAHRGPEPRPLPRPPPPTRPPAAPGPGPPPAPGPTP 243
Query: 302 LTKLNA-----PLSTILRAVGQTNVVQYPPPPRRPPA 333
L+K A P + + P PP PPA
Sbjct: 242 LSKPPAAEGIPPATAPPPRARRLRPPSAPAPPPTPPA 132
>TC78806 similar to GP|8978273|dbj|BAA98164.1 receptor protein kinase-like
{Arabidopsis thaliana}, partial (28%)
Length = 1962
Score = 35.0 bits (79), Expect = 0.25
Identities = 62/279 (22%), Positives = 115/279 (40%), Gaps = 23/279 (8%)
Frame = +2
Query: 1073 VKIKTDVPLRQVLQKPDLSGRLVSWSVELSEYDIQYE--PRGQVTVQSLIDFVAELTPTE 1130
V+ KT++ L + +L LV + E E + YE P G +L+D ++
Sbjct: 692 VEFKTEIELLSRVHHKNLVS-LVGFCYEKGEQMLVYEYVPNG-----TLLDSLS------ 835
Query: 1131 GEKTQGEWVLSVDGSSNNTGSGAGITI--ESPDKMIIEQSLKFEFKASNNQSEYEALIAG 1188
+ G W+ + G+ G+T E D II + +K S+N LIA
Sbjct: 836 --RKSGIWMDWIRRLKVTLGAARGLTYLHELADPPIIHRDIK-----SSNILLDNHLIAK 994
Query: 1189 LRLAIELGVQKLFIKGDSQLVVKQVKGEYQVKDPQLSKYLEVVRR--------LMMEVKE 1240
+ + G+ KL + + V QVKG DP+ ++ + LM+E+
Sbjct: 995 VA---DFGLSKLLVDSERGHVTTQVKGTMGYLDPEYYMTQQLTEKSDVYSFGVLMLELAT 1165
Query: 1241 IKIEHVPRGQNERADVLAKLASTGRLGNYQTVIQETL-----PRPSIDLVEIKLKAVKSV 1295
+ + + +G+ +V+ + ++ L N +++ ++L P+ VE+ L+ VK
Sbjct: 1166 SR-KPIEQGKYIVREVMRVMDTSKELYNLHSILDQSLLKGTRPKGLERYVELALRCVKEY 1342
Query: 1296 SEGEISW------MESIKTFLESPPKEDDLNTRTKRREA 1328
+ S +ESI + P + +T EA
Sbjct: 1343 AAERPSMAEVAKEIESIIELVGVNPNSESASTTENYEEA 1459
>TC83226 weakly similar to PIR|G86419|G86419 probable reverse transcriptase
100033-105622 [imported] - Arabidopsis thaliana, partial
(2%)
Length = 885
Score = 32.7 bits (73), Expect = 1.2
Identities = 23/78 (29%), Positives = 41/78 (52%), Gaps = 3/78 (3%)
Frame = +3
Query: 1140 LSVDGSSNNTGS-GAGITIESPDKMIIEQSLKFEFKASNNQSEYEA--LIAGLRLAIELG 1196
++ D + N G G GI I +++ S +E ++ E EA L+ G+R A + G
Sbjct: 654 VNTDANLQNHGKWGLGIIIRDEVGLVMAAST-WETDGNDRALEAEAYALLTGMRFAKDCG 830
Query: 1197 VQKLFIKGDSQLVVKQVK 1214
K+ +GD++ ++K VK
Sbjct: 831 FXKVXFEGDNEKLMKMVK 884
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.324 0.139 0.418
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 44,169,416
Number of Sequences: 36976
Number of extensions: 575029
Number of successful extensions: 4122
Number of sequences better than 10.0: 60
Number of HSP's better than 10.0 without gapping: 3009
Number of HSP's successfully gapped in prelim test: 158
Number of HSP's that attempted gapping in prelim test: 898
Number of HSP's gapped (non-prelim): 3399
length of query: 1706
length of database: 9,014,727
effective HSP length: 110
effective length of query: 1596
effective length of database: 4,947,367
effective search space: 7895997732
effective search space used: 7895997732
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 65 (29.6 bits)
Lotus: description of TM0046b.4


