

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0046a.5
(327 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
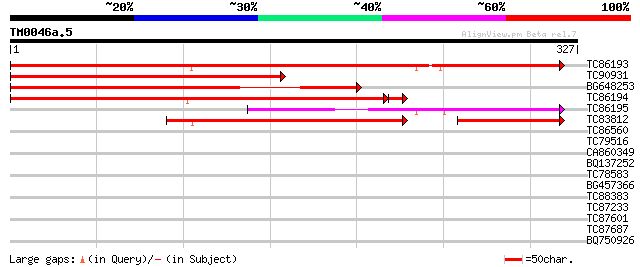
Score E
Sequences producing significant alignments: (bits) Value
TC86193 similar to GP|16974678|gb|AAL32439.1 SH3 domain-containi... 325 1e-89
TC90931 similar to GP|21928139|gb|AAM78097.1 AT4g18060/F15J5_30 ... 276 1e-74
BG648253 similar to GP|21928139|gb AT4g18060/F15J5_30 {Arabidops... 268 2e-72
TC86194 similar to GP|16974678|gb|AAL32439.1 SH3 domain-containi... 230 5e-61
TC86195 similar to GP|16974678|gb|AAL32439.1 SH3 domain-containi... 158 3e-39
TC83812 weakly similar to PIR|D86440|D86440 unknown protein [imp... 115 2e-26
TC86560 similar to GP|8843737|dbj|BAA97285.1 myosin heavy chain-... 37 0.008
TC79516 similar to GP|8777406|dbj|BAA96996.1 gb|AAF13095.1~gene_... 33 0.20
CA860349 weakly similar to GP|7294461|gb|A CG10083-PA {Drosophil... 30 0.98
BQ137252 similar to GP|6523547|emb| hydroxyproline-rich glycopro... 30 0.98
TC78583 similar to GP|4097585|gb|AAD09518.1| NTGP4 {Nicotiana ta... 30 1.3
BG457366 30 1.7
TC88383 similar to GP|21592515|gb|AAM64465.1 nuclear receptor bi... 29 2.8
TC87233 similar to PIR|T06048|T06048 kinesin-related protein kat... 28 3.7
TC87601 similar to GP|7671528|emb|CAB89490.1 CRK1 protein {Beta ... 28 6.3
TC87687 similar to GP|8894548|emb|CAB95829.1 hypothetical protei... 27 8.3
BQ750926 similar to PIR|T51049|T510 related to nucleolar phospho... 27 8.3
>TC86193 similar to GP|16974678|gb|AAL32439.1 SH3 domain-containing protein
2 {Arabidopsis thaliana}, partial (97%)
Length = 1311
Score = 325 bits (833), Expect = 1e-89
Identities = 183/367 (49%), Positives = 239/367 (64%), Gaps = 47/367 (12%)
Frame = +1
Query: 1 MDALRKQASKLREQVSKQQQAVIKQFSTSGYESSDVVVIDEGEMQIHQQLEKLYKATRTM 60
MDA+RKQASKLREQV++QQQAV+KQF GY SD +V DE E+ +HQ+LEKLY +TR
Sbjct: 115 MDAIRKQASKLREQVARQQQAVLKQFGGGGYGGSDNMVTDERELHLHQKLEKLYISTRAG 294
Query: 61 RDFQKEIVKAAETFTAIGYKHIETGTKLSEDCCKYGAENNIDN--ILAKAASVLGDARKH 118
+ +Q+++V+ E + G K +E GTKLSED KYGAEN + L +AA AR
Sbjct: 295 KHYQRDVVRGVEGYIVTGSKQVEIGTKLSEDSRKYGAENTCTSGGTLCRAALSYSRARAQ 474
Query: 119 VEKEHEELNRLLASQVLDPLRQMINGVPLEDARHLAQRYSRMRQEAETLREEISKRQVRV 178
+EKE L + L +QV +PLR M+ G PLEDARHLAQRY RMRQ+AE E+SKRQ +V
Sbjct: 475 MEKERGNLLKALGTQVAEPLRAMVVGAPLEDARHLAQRYDRMRQDAEAQAIEVSKRQAKV 654
Query: 179 RESPTSEQVA-KLHAAEARMKELKANMAVLGKEAAAALASVDAQQQRLTFQRLVAM---- 233
RE P + ++A KL AAEA++++LK+NM +LG+EAAAALA+V+AQQQRLT QRL+AM
Sbjct: 655 RELPGNSEIAMKLEAAEAKLQDLKSNMNILGREAAAALAAVEAQQQRLTLQRLIAMVEAE 834
Query: 234 -----------------MVSDRQKKESAPPV----------------------VVSENGS 254
M+S+RQ+ E APP + NGS
Sbjct: 835 RSYHQVVLQILDQLEGEMISERQRIE-APPTPSMDNSMPPPPPYEEVNGVYASQTTHNGS 1011
Query: 255 GKTM-YFLAEATHPFYGDSEKELSFSKGDFIVVRKVTPTGWSEGECNGKAGWFPSAYVEK 313
+M YFL E P+ SE EL+ S GD+IV+RKVT GW+EGEC G+AGWFP +Y+E+
Sbjct: 1012TDSMGYFLGEVLFPYSAVSEVELNLSVGDYIVIRKVTNNGWAEGECKGRAGWFPFSYIER 1191
Query: 314 RQRVPSS 320
R+RV +S
Sbjct: 1192RERVLAS 1212
>TC90931 similar to GP|21928139|gb|AAM78097.1 AT4g18060/F15J5_30
{Arabidopsis thaliana}, partial (44%)
Length = 595
Score = 276 bits (705), Expect = 1e-74
Identities = 141/159 (88%), Positives = 147/159 (91%)
Frame = +3
Query: 1 MDALRKQASKLREQVSKQQQAVIKQFSTSGYESSDVVVIDEGEMQIHQQLEKLYKATRTM 60
MDA RKQASKLREQV KQQQAVIKQFS SGYESSDVVVIDE EMQ HQ +EKLY+ATR
Sbjct: 102 MDAFRKQASKLREQVVKQQQAVIKQFSGSGYESSDVVVIDEVEMQRHQHMEKLYRATRAG 281
Query: 61 RDFQKEIVKAAETFTAIGYKHIETGTKLSEDCCKYGAENNIDNILAKAASVLGDARKHVE 120
RDFQKEIVKAAETFTAI YKHIETGTKLSE+CC+YGAENN DNILAKAASV GDARKHVE
Sbjct: 282 RDFQKEIVKAAETFTAISYKHIETGTKLSEECCRYGAENNSDNILAKAASVYGDARKHVE 461
Query: 121 KEHEELNRLLASQVLDPLRQMINGVPLEDARHLAQRYSR 159
KEHEELNRLL+SQVLDPLRQMING PLEDARHLAQRYS+
Sbjct: 462 KEHEELNRLLSSQVLDPLRQMINGPPLEDARHLAQRYSQ 578
>BG648253 similar to GP|21928139|gb AT4g18060/F15J5_30 {Arabidopsis
thaliana}, partial (46%)
Length = 563
Score = 268 bits (685), Expect = 2e-72
Identities = 146/203 (71%), Positives = 157/203 (76%)
Frame = +1
Query: 1 MDALRKQASKLREQVSKQQQAVIKQFSTSGYESSDVVVIDEGEMQIHQQLEKLYKATRTM 60
MDA RKQASKLREQV KQQQAVIKQFS SGYESSDVVVIDE EMQ HQ +EKLY+ATR
Sbjct: 55 MDAFRKQASKLREQVVKQQQAVIKQFSGSGYESSDVVVIDEVEMQRHQHMEKLYRATRAG 234
Query: 61 RDFQKEIVKAAETFTAIGYKHIETGTKLSEDCCKYGAENNIDNILAKAASVLGDARKHVE 120
RDFQKEIVKAAETFTAI YKHIETGTKLSE+CC+YGAENN DNILAKAASV GDARKHVE
Sbjct: 235 RDFQKEIVKAAETFTAISYKHIETGTKLSEECCRYGAENNSDNILAKAASVYGDARKHVE 414
Query: 121 KEHEELNRLLASQVLDPLRQMINGVPLEDARHLAQRYSRMRQEAETLREEISKRQVRVRE 180
KEHEELNRLL+SQ +EEIS+RQ RVRE
Sbjct: 415 KEHEELNRLLSSQ----------------------------------KEEISRRQARVRE 492
Query: 181 SPTSEQVAKLHAAEARMKELKAN 203
SPT+EQVAKLHAAEA+M+EL+AN
Sbjct: 493 SPTAEQVAKLHAAEAKMQELEAN 561
>TC86194 similar to GP|16974678|gb|AAL32439.1 SH3 domain-containing protein
2 {Arabidopsis thaliana}, partial (62%)
Length = 929
Score = 230 bits (587), Expect(2) = 5e-61
Identities = 122/221 (55%), Positives = 160/221 (72%), Gaps = 3/221 (1%)
Frame = +1
Query: 1 MDALRKQASKLREQVSKQQQAVIKQFSTSGYESSDVVVIDEGEMQIHQQLEKLYKATRTM 60
M+A+RKQASKLREQV++QQQAV+KQF GY SD +V DE E+Q HQ+LEKLY +TR
Sbjct: 148 MEAIRKQASKLREQVARQQQAVLKQFGAGGYGGSDNMVTDEVELQQHQKLEKLYISTRAG 327
Query: 61 RDFQKEIVKAAETFTAIGYKHIETGTKLSEDCCKYGAENNI--DNILAKAASVLGDARKH 118
+ +Q++IV+ E + G K +E GTKLSED KYG+EN + L++AA AR
Sbjct: 328 KHYQRDIVRGVEGYIVTGSKQVEIGTKLSEDSRKYGSENTCTSGSTLSRAALNYAHARAQ 507
Query: 119 VEKEHEELNRLLASQVLDPLRQMINGVPLEDARHLAQRYSRMRQEAETLREEISKRQVRV 178
+EKE L + L +QV +PLR M+ G PLEDARHLAQRY RMRQ+AE E+SKRQ +V
Sbjct: 508 MEKERGNLLKALGTQVAEPLRAMVMGAPLEDARHLAQRYDRMRQDAEAQAIEVSKRQAKV 687
Query: 179 RESP-TSEQVAKLHAAEARMKELKANMAVLGKEAAAALASV 218
RE+P +E KL AAE ++++LK+NMA+LGKEAAAA+ +V
Sbjct: 688 RETPGNAENTMKLEAAETKLQDLKSNMAILGKEAAAAMTAV 810
Score = 21.9 bits (45), Expect(2) = 5e-61
Identities = 9/11 (81%), Positives = 10/11 (90%)
Frame = +2
Query: 219 DAQQQRLTFQR 229
+AQQQRLT QR
Sbjct: 812 EAQQQRLTLQR 844
>TC86195 similar to GP|16974678|gb|AAL32439.1 SH3 domain-containing protein
2 {Arabidopsis thaliana}, partial (58%)
Length = 848
Score = 158 bits (399), Expect = 3e-39
Identities = 97/226 (42%), Positives = 126/226 (54%), Gaps = 43/226 (19%)
Frame = +2
Query: 138 LRQMINGVPLEDARHLAQRYSRMRQEAETLREEISKRQVRVRESPTSEQVAKLHAAEARM 197
L+ M G PLEDARHLAQRY RMRQ+AE E+SKRQ +VRE P + ++A
Sbjct: 38 LKGMWVGAPLEDARHLAQRYDRMRQDAEAQAIEVSKRQAKVRELPGNSEIA--------- 190
Query: 198 KELKANMAVLGKEAAAALASVDAQQQRLTFQRLVAM---------------------MVS 236
+ EAAAALA+V+AQQQRLT QRL+AM M+S
Sbjct: 191 ---------MKLEAAAALAAVEAQQQRLTLQRLIAMVEAERSYHQVVLQILDQLEGEMIS 343
Query: 237 DRQKKESAPPVVV---------------------SENGSGKTM-YFLAEATHPFYGDSEK 274
+RQ+ E+ P + + NGS +M YFL E P+ SE
Sbjct: 344 ERQRIEAPPTPSMDNSMPPPPPYEEVNGVYASQTTHNGSTDSMGYFLGEVLFPYSAVSEV 523
Query: 275 ELSFSKGDFIVVRKVTPTGWSEGECNGKAGWFPSAYVEKRQRVPSS 320
EL+ S GD+IV+RKVT GW+EGEC G+AGWFP +Y+E+R+RV +S
Sbjct: 524 ELNLSVGDYIVIRKVTNNGWAEGECKGRAGWFPFSYIERRERVLAS 661
>TC83812 weakly similar to PIR|D86440|D86440 unknown protein [imported] -
Arabidopsis thaliana, partial (33%)
Length = 921
Score = 115 bits (288), Expect = 2e-26
Identities = 62/143 (43%), Positives = 89/143 (61%), Gaps = 4/143 (2%)
Frame = +1
Query: 91 DCCKYGAENNIDNI---LAKAASVLGDARKHVEKEHEELNRLLASQVLDPLRQMINGVPL 147
DCCKYG EN LA+A+ G++ + +E E E L +L Q+ +PLR I G PL
Sbjct: 10 DCCKYGNENENQGSTYPLARASLQFGNSYEILENERETLLGILGDQISEPLRAQITGAPL 189
Query: 148 EDARHLAQRYSRMRQEAETLREEISKRQVRVRESPTS-EQVAKLHAAEARMKELKANMAV 206
EDARHL Y ++RQE E E+ +R+ ++R+S S E +L AE ++KE K+ +
Sbjct: 190 EDARHLTHNYDKLRQEVEGQAAEVLRRRSKLRDSSLSAESSMRLQNAEKKLKEHKSALVA 369
Query: 207 LGKEAAAALASVDAQQQRLTFQR 229
LG+EA AA++SV+ QQQ +T QR
Sbjct: 370 LGREATAAMSSVEEQQQHITLQR 438
Score = 78.6 bits (192), Expect = 3e-15
Identities = 35/62 (56%), Positives = 44/62 (70%)
Frame = +3
Query: 259 YFLAEATHPFYGDSEKELSFSKGDFIVVRKVTPTGWSEGECNGKAGWFPSAYVEKRQRVP 318
YF A+ HPF +E ELS S D++VVR+V GWSEGE NG AGWFPSAYV ++ VP
Sbjct: 651 YFFAKVIHPFDAQAEGELSLSVDDYVVVRQVAANGWSEGEFNGNAGWFPSAYVLRQDVVP 830
Query: 319 SS 320
++
Sbjct: 831 AN 836
>TC86560 similar to GP|8843737|dbj|BAA97285.1 myosin heavy chain-like
{Arabidopsis thaliana}, partial (28%)
Length = 2450
Score = 37.4 bits (85), Expect = 0.008
Identities = 48/205 (23%), Positives = 83/205 (40%), Gaps = 22/205 (10%)
Frame = +2
Query: 42 GEMQIHQQLEKLYKATRTM-RDFQ-----------KEIVKAAETFTAIGYKHIETGTKLS 89
GE IH++ K Y A R + ++ Q KE VK AET A +E +
Sbjct: 431 GERPIHKK-PKAYSAERVLAKETQLHVAEKELNKLKEQVKNAETTKAQALVELERAQRTV 607
Query: 90 EDCCKYGAENNIDNILAKAASVLGDARKHVEKEHEELNRLLASQVLDPLRQMINGVPLED 149
+D L + ++ ++R+ K E + + +NG E+
Sbjct: 608 DD-------------LTQKLKLITESRESAVKATEAAKSQAKQKYGES--DGVNGAWKEE 742
Query: 150 ARHLAQRYSRMRQEAETLREEISKRQVRVRESPTS--EQVAKLHAAEARMKELKANMAVL 207
+ QRY+ + E + ++++ K + S + V + AE MKE ++ L
Sbjct: 743 LENAVQRYASIMTELDAAKQDLRKTRQEYDSSSDARVSAVKRTEEAENAMKENTERVSEL 922
Query: 208 GKEAAAA--------LASVDAQQQR 224
KE +A LA V++QQQ+
Sbjct: 923 SKEISAVKESIEQTKLAYVESQQQQ 997
>TC79516 similar to GP|8777406|dbj|BAA96996.1
gb|AAF13095.1~gene_id:MIF21.5~strong similarity to
unknown protein {Arabidopsis thaliana}, partial (29%)
Length = 1077
Score = 32.7 bits (73), Expect = 0.20
Identities = 26/83 (31%), Positives = 41/83 (49%), Gaps = 2/83 (2%)
Frame = +3
Query: 162 QEAETLREEISKRQVRVRESPTSEQVAKLHAAEARMKELKANMAVLGKE--AAAALASVD 219
+EAE L+ E K++ ++ E E++ +L AEA M +LKAN A E ALA D
Sbjct: 336 REAEELKMERQKKKSQIEEL---ERIVRLKNAEADMFQLKANEAKREAERLQRIALAKSD 506
Query: 220 AQQQRLTFQRLVAMMVSDRQKKE 242
++ T L + +K+
Sbjct: 507 KSEEEYTSNYLKQKLSEAEAEKQ 575
>CA860349 weakly similar to GP|7294461|gb|A CG10083-PA {Drosophila
melanogaster}, partial (9%)
Length = 457
Score = 30.4 bits (67), Expect = 0.98
Identities = 18/55 (32%), Positives = 25/55 (44%), Gaps = 1/55 (1%)
Frame = +3
Query: 262 AEATHPFYGDSEKELSFSKGDFIV-VRKVTPTGWSEGECNGKAGWFPSAYVEKRQ 315
A A + + E+SF +G I + V+ W E G G FP+ YVE Q
Sbjct: 177 ARALYDYAAGEHNEISFEEGHIITDIDFVSEDWWQGTEEGGTKGLFPANYVELLQ 341
>BQ137252 similar to GP|6523547|emb| hydroxyproline-rich glycoprotein DZ-HRGP
{Volvox carteri f. nagariensis}, partial (10%)
Length = 1067
Score = 30.4 bits (67), Expect = 0.98
Identities = 22/73 (30%), Positives = 33/73 (45%)
Frame = +2
Query: 154 AQRYSRMRQEAETLREEISKRQVRVRESPTSEQVAKLHAAEARMKELKANMAVLGKEAAA 213
AQ+Y+R A R R+ R R PT+ + + H A R +E + A ++A
Sbjct: 374 AQQYARTTTRAPQCRASAQDRRRRARSIPTTTRTQQRHKACDR-EENDTSHARSARQARR 550
Query: 214 ALASVDAQQQRLT 226
+VD Q R T
Sbjct: 551 RTRAVDQQPHRET 589
>TC78583 similar to GP|4097585|gb|AAD09518.1| NTGP4 {Nicotiana tabacum},
partial (86%)
Length = 1269
Score = 30.0 bits (66), Expect = 1.3
Identities = 30/137 (21%), Positives = 52/137 (37%), Gaps = 14/137 (10%)
Frame = +3
Query: 6 KQASKLREQVSKQQQAVIKQFSTSGYESSDVVVIDEGEMQIHQQLEKLYKATRTMRDFQK 65
K+ S +Q+ ++ Q Y + +G M++H++ K+
Sbjct: 648 KKRSGQVQQLLSFVNLIVLQNGGQPYTDELFAELKKGAMKLHREQRKVDSLEGYSEGQIS 827
Query: 66 EIVKAAETFTAIGYKHI---------ETGTKLS-----EDCCKYGAENNIDNILAKAASV 111
E+ K + KHI E T+L E + AE++ K+
Sbjct: 828 ELKKHMQQTYEEQLKHITEMIESKLKEATTRLEKQLAEEQAARLRAEDSAKLAQKKSDDE 1007
Query: 112 LGDARKHVEKEHEELNR 128
+ RKH+EK HEEL +
Sbjct: 1008IRKLRKHLEKAHEELRK 1058
>BG457366
Length = 647
Score = 29.6 bits (65), Expect = 1.7
Identities = 23/108 (21%), Positives = 47/108 (43%), Gaps = 2/108 (1%)
Frame = +1
Query: 97 AENNIDNILAKAASVLGDARKHVEKEHEELNRLLASQVLDPLRQMINGVPLEDARH--LA 154
A+NNI ++ + A+S + + ++ +++ H E+ + L +A H L
Sbjct: 199 AQNNISDLKSLASSTIDELKRRIDRSHSEILKDL------------------EASHSRLH 324
Query: 155 QRYSRMRQEAETLREEISKRQVRVRESPTSEQVAKLHAAEARMKELKA 202
+R+ Q + +E K ++ E T + A + E M E +A
Sbjct: 325 KRFKMQSQGCQQAMDEAEKDYKKLSERITESREAMKASYEEFMAEAQA 468
>TC88383 similar to GP|21592515|gb|AAM64465.1 nuclear receptor binding
factor-like protein {Arabidopsis thaliana}, partial
(90%)
Length = 1313
Score = 28.9 bits (63), Expect = 2.8
Identities = 19/46 (41%), Positives = 25/46 (54%), Gaps = 7/46 (15%)
Frame = -3
Query: 115 ARKHVEKE---HEELNRLLASQVLDPLRQMING----VPLEDARHL 153
+RK VEKE H E N+L SQ +PL+ + N VP +HL
Sbjct: 405 SRKKVEKE*PNHRERNKLRRSQRSEPLQHLRNHQLQVVPASQDKHL 268
>TC87233 similar to PIR|T06048|T06048 kinesin-related protein katB -
Arabidopsis thaliana, partial (18%)
Length = 1488
Score = 28.5 bits (62), Expect = 3.7
Identities = 25/88 (28%), Positives = 40/88 (45%), Gaps = 5/88 (5%)
Frame = +3
Query: 210 EAAAALAS-VDAQQQRLTFQRLVAMMVSD----RQKKESAPPVVVSENGSGKTMYFLAEA 264
E AAA+ S + ++ RL F+R + D +++ E+A +VS N K + +
Sbjct: 708 EKAAAMESLIKEREARLDFERSQTTLSEDLGRAQRELETANQKIVSLNDMYKRLQEYITS 887
Query: 265 THPFYGDSEKELSFSKGDFIVVRKVTPT 292
+ G ELS +GD V K T
Sbjct: 888 LQQYNGKLHSELSTVEGDLKRVEKEKAT 971
>TC87601 similar to GP|7671528|emb|CAB89490.1 CRK1 protein {Beta vulgaris},
partial (50%)
Length = 1493
Score = 27.7 bits (60), Expect = 6.3
Identities = 18/65 (27%), Positives = 33/65 (50%), Gaps = 2/65 (3%)
Frame = -2
Query: 265 THPFYGDSEKELS--FSKGDFIVVRKVTPTGWSEGECNGKAGWFPSAYVEKRQRVPSSNF 322
TH +EK+L KG+ + ++ PTGW E C K F +++ Q++ S +
Sbjct: 307 THTIICGTEKQLRRPVPKGNHTICHRM-PTGWIEERCKPKVSNFEYSFI-VNQKI*SFDI 134
Query: 323 SSEVY 327
+ +V+
Sbjct: 133 AVKVH 119
>TC87687 similar to GP|8894548|emb|CAB95829.1 hypothetical protein {Cicer
arietinum}, partial (67%)
Length = 1969
Score = 27.3 bits (59), Expect = 8.3
Identities = 20/61 (32%), Positives = 29/61 (46%)
Frame = +3
Query: 163 EAETLREEISKRQVRVRESPTSEQVAKLHAAEARMKELKANMAVLGKEAAAALASVDAQQ 222
EAE +EE+ K++ V S E AK EA + + A + + +E AS QQ
Sbjct: 510 EAEQPKEEVEKKETEVAASSVDEDGAK--TVEAIEESVVAVASSVPEEPKVVEASSPEQQ 683
Query: 223 Q 223
Q
Sbjct: 684 Q 686
>BQ750926 similar to PIR|T51049|T510 related to nucleolar phosphoprotein
[imported] - Neurospora crassa, partial (9%)
Length = 779
Score = 27.3 bits (59), Expect = 8.3
Identities = 21/67 (31%), Positives = 34/67 (50%)
Frame = +2
Query: 157 YSRMRQEAETLREEISKRQVRVRESPTSEQVAKLHAAEARMKELKANMAVLGKEAAAALA 216
+S+ +++ E SK + +V +P E +KE KAN A KEAAAA +
Sbjct: 20 HSKETDPSDSKESEESKPEAKVTTTPLIEY----------LKEKKANKA---KEAAAAKS 160
Query: 217 SVDAQQQ 223
+ A+Q+
Sbjct: 161 AKQARQE 181
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.313 0.127 0.343
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 6,810,034
Number of Sequences: 36976
Number of extensions: 68768
Number of successful extensions: 317
Number of sequences better than 10.0: 34
Number of HSP's better than 10.0 without gapping: 307
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 307
length of query: 327
length of database: 9,014,727
effective HSP length: 96
effective length of query: 231
effective length of database: 5,465,031
effective search space: 1262422161
effective search space used: 1262422161
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.9 bits)
S2: 58 (26.9 bits)
Lotus: description of TM0046a.5


