

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
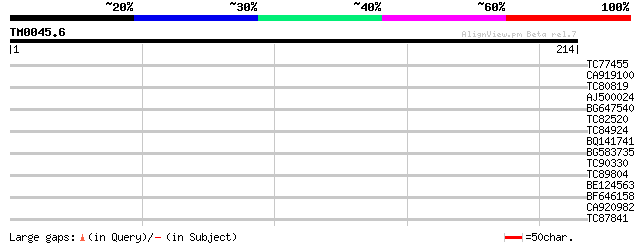
Query= TM0045.6
(214 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC77455 similar to GP|22335695|dbj|BAC10549. nine-cis-epoxycarot... 34 0.048
CA919100 homologue to PIR|G90291|G902 endoglucanase precursor [i... 33 0.063
TC80819 weakly similar to GP|10140689|gb|AAG13524.1 putative non... 33 0.082
AJ500024 33 0.11
BG647540 32 0.24
TC82520 29 1.2
TC84924 weakly similar to PIR|F96545|F96545 hypothetical protein... 28 2.6
BQ141741 GP|7243239|db KIAA1429 protein {Homo sapiens}, partial ... 27 4.5
BG583735 27 5.9
TC90330 weakly similar to PIR|D96593|D96593 myosin-like protein ... 27 5.9
TC89804 similar to GP|14719339|gb|AAK73157.1 putative protein ki... 27 5.9
BE124563 27 5.9
BF646158 similar to GP|4530126|gb| receptor-like protein kinase ... 27 5.9
CA920982 similar to PIR|H86215|H862 protein T6D22.22 [imported] ... 27 5.9
TC87841 similar to PIR|C86334|C86334 hypothetical protein AAF799... 27 5.9
>TC77455 similar to GP|22335695|dbj|BAC10549. nine-cis-epoxycarotenoid
dioxygenase1 {Pisum sativum}, partial (43%)
Length = 1865
Score = 33.9 bits (76), Expect = 0.048
Identities = 14/33 (42%), Positives = 20/33 (60%)
Frame = -2
Query: 86 FSHVWHFAAPSNVKAFAWRFLLVKIPSRLNLWK 118
F +W AP+ V AF+W L +IP+ +NL K
Sbjct: 331 FRELWKSKAPAKVLAFSWTLFLDRIPTMVNLGK 233
>CA919100 homologue to PIR|G90291|G902 endoglucanase precursor [imported] -
Sulfolobus solfataricus, partial (3%)
Length = 789
Score = 33.5 bits (75), Expect = 0.063
Identities = 14/39 (35%), Positives = 24/39 (60%), Gaps = 1/39 (2%)
Frame = -2
Query: 79 ALAQVLGFSH-VWHFAAPSNVKAFAWRFLLVKIPSRLNL 116
+L+ +L ++ +WH P V FAWR L ++P++ NL
Sbjct: 617 SLSLILPYAEMIWHRQVPLKVSVFAWRLLRDRLPTKSNL 501
>TC80819 weakly similar to GP|10140689|gb|AAG13524.1 putative non-LTR
retroelement reverse transcriptase {Oryza sativa
(japonica cultivar-group)}, partial (2%)
Length = 1262
Score = 33.1 bits (74), Expect = 0.082
Identities = 24/77 (31%), Positives = 34/77 (43%), Gaps = 5/77 (6%)
Frame = +2
Query: 89 VWHFAAPSNVKAFAWRFLLVKIPSRLNLWKLG---ATRGLISFGY--NRSKDEFLVVLIK 143
VWH PS V F WR L ++P++ NL G AT G + S +
Sbjct: 626 VWHKHIPSKVSLFVWRLLRNRLPTKDNLVHRGVLLATNAACVCGCVDSESTTHLFLHCNV 805
Query: 144 LCNLLHLAKNKITIISL 160
C+L L +N + I S+
Sbjct: 806 FCSLWSLVRNWLGIPSM 856
>AJ500024
Length = 538
Score = 32.7 bits (73), Expect = 0.11
Identities = 23/73 (31%), Positives = 38/73 (51%), Gaps = 3/73 (4%)
Frame = -1
Query: 127 SFGYNRSKDEFLVVLIKLCNLLHLAKNKITIISLRNYVRDYFDIADTEYYFYKFNLDYIG 186
SFGY++ KD++LVVL+ L + + SLR+ V + D Y Y+ N D +
Sbjct: 493 SFGYDQLKDDYLVVLMSYDPALDNIYSCLEFFSLRDNV---WKEMDCPYCPYRSNFDEMP 323
Query: 187 QFLC---DSLYWL 196
+ C +++WL
Sbjct: 322 KAGCLYNGAIHWL 284
>BG647540
Length = 625
Score = 31.6 bits (70), Expect = 0.24
Identities = 24/79 (30%), Positives = 35/79 (43%), Gaps = 8/79 (10%)
Frame = +3
Query: 32 FQNMWRNIGAYEDGSVRDMITLAKYNGGDLD--------TYFEHPDFEPLVEGPEALAQV 83
F + WR E+G VR Y GGDL+ + ++ + EG +V
Sbjct: 294 FGSCWRIWTDIEEGKVR------MYGGGDLEENGWFSVNSMYKKLEGMRFEEGSLTEMRV 455
Query: 84 LGFSHVWHFAAPSNVKAFA 102
F+H+W +APS V A A
Sbjct: 456 KVFTHIWKSSAPSKVVAEA 512
>TC82520
Length = 833
Score = 29.3 bits (64), Expect = 1.2
Identities = 11/28 (39%), Positives = 16/28 (56%)
Frame = +3
Query: 89 VWHFAAPSNVKAFAWRFLLVKIPSRLNL 116
VW PS V F WR ++P+++NL
Sbjct: 186 VWQKNIPSKVSMFVWRLFHNRLPTKVNL 269
>TC84924 weakly similar to PIR|F96545|F96545 hypothetical protein F8A12.9
[imported] - Arabidopsis thaliana, partial (9%)
Length = 575
Score = 28.1 bits (61), Expect = 2.6
Identities = 11/25 (44%), Positives = 19/25 (76%)
Frame = +1
Query: 125 LISFGYNRSKDEFLVVLIKLCNLLH 149
L FGY++S+D+++VV I +L+H
Sbjct: 466 LYGFGYDQSRDDYVVVFIVQ*SLIH 540
>BQ141741 GP|7243239|db KIAA1429 protein {Homo sapiens}, partial (0%)
Length = 1177
Score = 27.3 bits (59), Expect = 4.5
Identities = 22/83 (26%), Positives = 42/83 (50%), Gaps = 3/83 (3%)
Frame = +2
Query: 88 HVWHFAAPSNVKAFAWRFLLVKIPSRLNLWKLGATRGLISFGYNRSKDEFLVV-LIKLCN 146
H H N + A+R++ + +P L+ R + NR ++E V+ LI++C
Sbjct: 476 HHLHLLLLLNSGSVAYRYISLVLPRYRELF-----RKRV*LKTNRKRNEGAVLCLIEICK 640
Query: 147 LLHLAKN--KITIISLRNYVRDY 167
L HLA + + ++SL + +R +
Sbjct: 641 LCHLAHSGFVLPLLSLSSVLRRF 709
>BG583735
Length = 550
Score = 26.9 bits (58), Expect = 5.9
Identities = 12/29 (41%), Positives = 14/29 (47%)
Frame = +3
Query: 92 FAAPSNVKAFAWRFLLVKIPSRLNLWKLG 120
F PS K F WR L K+P + W G
Sbjct: 198 FDIPSVRKYFGWRLLQDKLPMCVEFWNRG 284
>TC90330 weakly similar to PIR|D96593|D96593 myosin-like protein
97843-94399 [imported] - Arabidopsis thaliana, partial
(36%)
Length = 1191
Score = 26.9 bits (58), Expect = 5.9
Identities = 11/37 (29%), Positives = 19/37 (50%)
Frame = -1
Query: 90 WHFAAPSNVKAFAWRFLLVKIPSRLNLWKLGATRGLI 126
WH+ P++++ F W IPS W+L G++
Sbjct: 147 WHWWWPTSIRTFRW------IPSSGTHWRLSMNTGVL 55
>TC89804 similar to GP|14719339|gb|AAK73157.1 putative protein kinase {Oryza
sativa}, partial (33%)
Length = 874
Score = 26.9 bits (58), Expect = 5.9
Identities = 7/17 (41%), Positives = 13/17 (76%)
Frame = +1
Query: 101 FAWRFLLVKIPSRLNLW 117
F+W+F L+++P +N W
Sbjct: 676 FSWKFCLMRVPKSMNSW 726
>BE124563
Length = 562
Score = 26.9 bits (58), Expect = 5.9
Identities = 12/29 (41%), Positives = 14/29 (47%)
Frame = +1
Query: 92 FAAPSNVKAFAWRFLLVKIPSRLNLWKLG 120
F PS K F WR L K+P + W G
Sbjct: 112 FDIPSVRKYFGWRLLQDKLPMCVEFWNRG 198
>BF646158 similar to GP|4530126|gb| receptor-like protein kinase homolog
RK20-1 {Phaseolus vulgaris}, partial (19%)
Length = 653
Score = 26.9 bits (58), Expect = 5.9
Identities = 21/64 (32%), Positives = 30/64 (46%), Gaps = 1/64 (1%)
Frame = +3
Query: 119 LGATRGLISFGYNRS-KDEFLVVLIKLCNLLHLAKNKITIISLRNYVRDYFDIADTEYYF 177
+G RG + RS ++ V+L KLC A + LR R F I DT + +
Sbjct: 207 IGLCRGDVKPDLCRSCLNDSRVLLTKLCPTQKEAISWYDNCMLRYSNRSIFGIMDTSFSY 386
Query: 178 YKFN 181
YK+N
Sbjct: 387 YKWN 398
>CA920982 similar to PIR|H86215|H862 protein T6D22.22 [imported] -
Arabidopsis thaliana, partial (17%)
Length = 773
Score = 26.9 bits (58), Expect = 5.9
Identities = 16/41 (39%), Positives = 22/41 (53%), Gaps = 7/41 (17%)
Frame = -1
Query: 49 DMITLAKYNGGDLDTYFEHPD-------FEPLVEGPEALAQ 82
DM T++K +D F+HP F+PL EGPE +Q
Sbjct: 638 DMKTVSK---SKMDETFQHPSIELFIMGFKPLAEGPENSSQ 525
>TC87841 similar to PIR|C86334|C86334 hypothetical protein AAF79906.1
[imported] - Arabidopsis thaliana, partial (18%)
Length = 1126
Score = 26.9 bits (58), Expect = 5.9
Identities = 10/21 (47%), Positives = 15/21 (70%)
Frame = +3
Query: 173 TEYYFYKFNLDYIGQFLCDSL 193
T++YF F L +G+F+CD L
Sbjct: 912 TDFYFAGFVLFVVGEFVCDRL 974
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.327 0.144 0.451
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 7,529,682
Number of Sequences: 36976
Number of extensions: 110766
Number of successful extensions: 684
Number of sequences better than 10.0: 30
Number of HSP's better than 10.0 without gapping: 683
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 684
length of query: 214
length of database: 9,014,727
effective HSP length: 92
effective length of query: 122
effective length of database: 5,612,935
effective search space: 684778070
effective search space used: 684778070
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.7 bits)
S2: 56 (26.2 bits)
Lotus: description of TM0045.6


