

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0042.13
(250 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
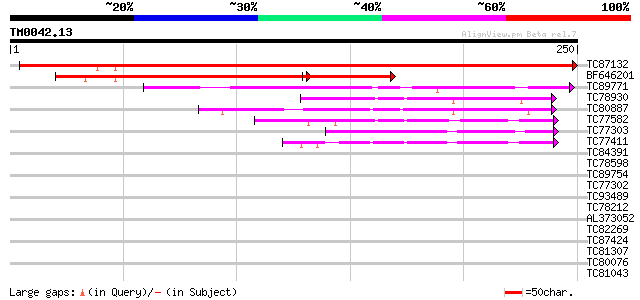
Score E
Sequences producing significant alignments: (bits) Value
TC87132 similar to GP|13702824|gb|AAK38500.1 putative FK506-bind... 400 e-112
BF646201 similar to GP|12247991|gb| unknown protein {Arabidopsis... 161 5e-54
TC89771 similar to GP|6143884|gb|AAF04431.1| unknown protein {Ar... 69 2e-12
TC78930 similar to GP|21535744|emb|CAD35362. FK506 binding prote... 56 1e-08
TC80887 similar to GP|6686798|emb|CAB64721.1 FKBP like protein {... 47 5e-06
TC77582 similar to SP|Q43207|FKB7_WHEAT 70 kDa peptidylprolyl is... 46 1e-05
TC77303 similar to SP|Q43207|FKB7_WHEAT 70 kDa peptidylprolyl is... 41 5e-04
TC77411 homologue to SP|Q41649|FKB2_VICFA FK506-binding protein ... 41 5e-04
TC84391 similar to PIR|E86340|E86340 protein F2D10.32 [imported]... 40 0.001
TC78598 similar to SP|Q43207|FKB7_WHEAT 70 kDa peptidylprolyl is... 39 0.001
TC89754 similar to GP|21618273|gb|AAM67323.1 unknown {Arabidopsi... 39 0.002
TC77302 similar to GP|1354207|gb|AAB82061.1| rof1 {Arabidopsis t... 38 0.003
TC93489 homologue to PIR|T47848|T47848 hypothetical protein T8B1... 35 0.036
TC78212 similar to GP|17381160|gb|AAL36392.1 unknown protein {Ar... 35 0.036
AL373052 similar to GP|21593014|gb| putative aldolase {Arabidops... 34 0.046
TC82269 weakly similar to GP|14581464|gb|AAK31804.1 protocadheri... 34 0.046
TC87424 similar to GP|19699309|gb|AAL91265.1 AT4g10840/F25I24_50... 34 0.061
TC81307 weakly similar to GP|4335772|gb|AAD17449.1| unknown prot... 33 0.079
TC80076 similar to GP|10177459|dbj|BAB10850. gene_id:MQB2.13~unk... 33 0.10
TC81043 weakly similar to GP|6523547|emb|CAB62280.1 hydroxyproli... 32 0.18
>TC87132 similar to GP|13702824|gb|AAK38500.1 putative FK506-binding protein
{Oryza sativa}, partial (68%)
Length = 857
Score = 400 bits (1028), Expect = e-112
Identities = 207/252 (82%), Positives = 224/252 (88%), Gaps = 6/252 (2%)
Frame = +2
Query: 5 FGSPPVFSLPLTRNSHHIASSSPTPPPIPPQQS--QSPTASSS----QSVQGTAKAQPQN 58
FGS +F LP TR SHHI SSS TPPP PQ S SP ++++ QSVQ +AKAQ Q
Sbjct: 17 FGSSSIFLLPNTRTSHHITSSSATPPPQQPQPSGPSSPQSTTNLNEDQSVQVSAKAQQQI 196
Query: 59 PIKPVTSSTKVDSTDWIATSLTRRFGLGAGLAWVGFLAFGVVSEQIKTRLEVSQQEANTR 118
PIKPV SSTKVDSTDWIATSLTRRFGLGAGLAWVGFLAFGVVSEQIKTRLEVSQQE+NTR
Sbjct: 197 PIKPVVSSTKVDSTDWIATSLTRRFGLGAGLAWVGFLAFGVVSEQIKTRLEVSQQESNTR 376
Query: 119 DVEKEEEVTLPNGIRYCELKVGGGASPRPGDLVVIDIMGKVESTGEVFVNTFEGDKKPLA 178
+VE+E+EV LPNGIRY ELK+GGG PR GDLVVIDIMGKVESTGEVFVNTFEGDKK LA
Sbjct: 377 NVEEEKEVILPNGIRYYELKIGGGDMPRRGDLVVIDIMGKVESTGEVFVNTFEGDKKALA 556
Query: 179 LVMGSRPYSKGVCEGIEYVLKTMKAGGKRKVIVPPQLGFRENGADLGSGVQIPPLATLEY 238
LVMGSRPYSKGVCEGIEYV+K+MKAGGKRKVIVPP+LGFRENGADLGSGV+IPPLATLEY
Sbjct: 557 LVMGSRPYSKGVCEGIEYVIKSMKAGGKRKVIVPPELGFRENGADLGSGVEIPPLATLEY 736
Query: 239 IVEVDKVSIAPA 250
+V+VDKVSIAPA
Sbjct: 737 VVQVDKVSIAPA 772
>BF646201 similar to GP|12247991|gb| unknown protein {Arabidopsis thaliana},
partial (40%)
Length = 609
Score = 161 bits (408), Expect(2) = 5e-54
Identities = 90/132 (68%), Positives = 97/132 (73%), Gaps = 19/132 (14%)
Frame = +1
Query: 21 HIASSSPTPPPI---------PPQQSQSPTASSS----------QSVQGTAKAQPQNPIK 61
H S SP P P+ PP Q P+ SS QSVQ +AKAQ Q PIK
Sbjct: 16 HQFSYSPIPEPVITSLLHQQRPPPQQPQPSGPSSPQSTTNLNEDQSVQVSAKAQQQIPIK 195
Query: 62 PVTSSTKVDSTDWIATSLTRRFGLGAGLAWVGFLAFGVVSEQIKTRLEVSQQEANTRDVE 121
PV SSTKVDSTDWIATSLTRRFGLGAGLAWVGFLAFGVVSEQIKTRLEVSQQE+NTR+VE
Sbjct: 196 PVVSSTKVDSTDWIATSLTRRFGLGAGLAWVGFLAFGVVSEQIKTRLEVSQQESNTRNVE 375
Query: 122 KEEEVTLPNGIR 133
+E+EV LPNGIR
Sbjct: 376 EEKEVILPNGIR 411
Score = 67.0 bits (162), Expect(2) = 5e-54
Identities = 33/41 (80%), Positives = 34/41 (82%)
Frame = +2
Query: 130 NGIRYCELKVGGGASPRPGDLVVIDIMGKVESTGEVFVNTF 170
N RY ELK+GGG PR GDLVVIDIMGKV STGEVFVNTF
Sbjct: 485 NMYRYYELKIGGGDMPRRGDLVVIDIMGKVXSTGEVFVNTF 607
Score = 35.8 bits (81), Expect = 0.016
Identities = 25/69 (36%), Positives = 35/69 (50%), Gaps = 2/69 (2%)
Frame = +3
Query: 5 FGSPPVFSLPLTRNSHHIASSSPTPPPIPPQQSQSPTASSSQSVQGTA--KAQPQNPIKP 62
FGS +F LP TR SHHI SSS T S + T ++ +S+ T K + ++
Sbjct: 6 FGSSSIFLLPNTRTSHHITSSSAT--------SSTTTTATLRSIISTVNNKFE*RSVSAS 161
Query: 63 VTSSTKVDS 71
+ ST DS
Sbjct: 162LCQSTATDS 188
>TC89771 similar to GP|6143884|gb|AAF04431.1| unknown protein {Arabidopsis
thaliana}, partial (65%)
Length = 1003
Score = 68.9 bits (167), Expect = 2e-12
Identities = 52/198 (26%), Positives = 91/198 (45%), Gaps = 8/198 (4%)
Frame = +3
Query: 60 IKPVTSSTKVDSTDWIATSLTRRFGLGAGLAWVGFLAFGVVSEQIKTRLEVSQQEANTRD 119
++P++ S +++ + TSL F GV + + + S++
Sbjct: 276 VEPISMSLQIEGRRALLTSLLTTFA-------------GVYACDVAEAVSTSRRALRGAK 416
Query: 120 VEKEEEVTLPNGIRYCELKVGGGASPRPGDLVVIDIMGKVESTGEVFVNTFEGDKKPLAL 179
+ + + TLPNG++Y +LKVG GA G V I + K G F+ + +G
Sbjct: 417 IPESDFKTLPNGLKYYDLKVGDGAEAVKGSRVAIHYVAKWR--GITFMTSRQG-----MG 575
Query: 180 VMGSRPYS--------KGVCEGIEYVLKTMKAGGKRKVIVPPQLGFRENGADLGSGVQIP 231
V G PY V +G++ ++ M+ GG+R +IVPP+L + G +IP
Sbjct: 576 VGGGTPYGFDVGESARGNVLKGLDVGVEGMRVGGQRLLIVPPELAYGSRGVQ-----EIP 740
Query: 232 PLATLEYIVEVDKVSIAP 249
P AT+E +E+ + +P
Sbjct: 741 PNATIEMDIELLAIKQSP 794
>TC78930 similar to GP|21535744|emb|CAD35362. FK506 binding protein 1
{Arabidopsis thaliana}, partial (58%)
Length = 842
Score = 55.8 bits (133), Expect = 1e-08
Identities = 40/116 (34%), Positives = 60/116 (51%), Gaps = 3/116 (2%)
Frame = +1
Query: 129 PNGIRYCELKVGGGASPRPGDLVVIDIMGKVESTGEVFVNTFEGDKKPLALVMGSRPYSK 188
P+G+ YC+ VG G G L+ +G++E+ G+VF +++ K PL +G K
Sbjct: 271 PSGLSYCDKVVGYGPQAVKGQLIKAHYVGRLEN-GKVFDSSYNRGK-PLTFRVGVGEVIK 444
Query: 189 GVCEGI--EYVLKTMKAGGKRKVIVPPQLGFRENGADL-GSGVQIPPLATLEYIVE 241
G GI + + M GGKR + +PP+ G+ GA G IPP A L + VE
Sbjct: 445 GWDVGILGDDGIPPMLTGGKRTLKLPPEFGYGSRGAGCKGGSCVIPPDAVLLFDVE 612
>TC80887 similar to GP|6686798|emb|CAB64721.1 FKBP like protein {Arabidopsis
thaliana}, partial (65%)
Length = 742
Score = 47.4 bits (111), Expect = 5e-06
Identities = 46/162 (28%), Positives = 75/162 (45%), Gaps = 4/162 (2%)
Frame = +2
Query: 84 GLGAGLAWV-GFLAFGVVSEQIKTRLEVSQQEANTRDVEKEEEVTLPNGIRYCELKVGGG 142
G+G AWV G A + +I+ V + V+ +G+ YC+ G G
Sbjct: 245 GIGIVAAWVMGLTALDADATRIEYYATVGEPMCELNYVK--------SGLGYCDYVEGFG 400
Query: 143 ASPRPGDLVVIDIMGKVESTGEVFVNTFEGDKKPLALVMGSRPYSKGVCEGI--EYVLKT 200
G+L+ I + + G VF ++++ KPL + +G +G+ +GI +
Sbjct: 401 DEAPLGELIDIHYTARF-ADGIVFDSSYKR-AKPLTMRIGVGKVIRGLDQGILGGEGVPP 574
Query: 201 MKAGGKRKVIVPPQLGFRENGADLGSG-VQIPPLATLEYIVE 241
M+ GGKRK+ +PP L + A SG IP ATL Y ++
Sbjct: 575 MRIGGKRKLTIPPLLAYGPEPAGCFSGDCNIPGNATLLYDIK 700
>TC77582 similar to SP|Q43207|FKB7_WHEAT 70 kDa peptidylprolyl isomerase (EC
5.2.1.8) (Peptidylprolyl cis-trans isomerase) (PPiase),
partial (92%)
Length = 2130
Score = 46.2 bits (108), Expect = 1e-05
Identities = 44/141 (31%), Positives = 65/141 (45%), Gaps = 7/141 (4%)
Frame = +1
Query: 109 EVSQQEANTRDVEKEEEVTLPN------GIRYCELKVGGG-ASPRPGDLVVIDIMGKVES 161
E+ Q E D + EE G++ LK+G G +P GD V + G +
Sbjct: 118 EIPQAEEMNEDFDLPEEKVGDEREVGDRGLKKKLLKLGEGWDTPESGDEVQVHYTGTLLD 297
Query: 162 TGEVFVNTFEGDKKPLALVMGSRPYSKGVCEGIEYVLKTMKAGGKRKVIVPPQLGFRENG 221
G F ++ + D P + +G +G EGI KTMK G +PP+L + E+
Sbjct: 298 -GTKFDSSRDRDS-PFSFTLGQGQVIQGWDEGI----KTMKKGENALFTIPPELAYGES- 456
Query: 222 ADLGSGVQIPPLATLEYIVEV 242
GS IPP ATL++ VE+
Sbjct: 457 ---GSPPTIPPNATLQFDVEM 510
>TC77303 similar to SP|Q43207|FKB7_WHEAT 70 kDa peptidylprolyl isomerase (EC
5.2.1.8) (Peptidylprolyl cis-trans isomerase) (PPiase),
partial (50%)
Length = 995
Score = 40.8 bits (94), Expect = 5e-04
Identities = 29/103 (28%), Positives = 47/103 (45%)
Frame = +2
Query: 140 GGGASPRPGDLVVIDIMGKVESTGEVFVNTFEGDKKPLALVMGSRPYSKGVCEGIEYVLK 199
G P D V + G + G+ F ++ + + +G KG EG ++
Sbjct: 56 GDNPIPNLDDEVHVHFTGTLLLDGKKFDSSSDRGTSTFSFTLGRGQVIKGWGEG----MR 223
Query: 200 TMKAGGKRKVIVPPQLGFRENGADLGSGVQIPPLATLEYIVEV 242
TM+ G K +PP+L + ++G IPP ATL+Y VE+
Sbjct: 224 TMRKGEKALFTIPPELAYGKSGLP----PNIPPNATLQYDVEL 340
>TC77411 homologue to SP|Q41649|FKB2_VICFA FK506-binding protein 2 precursor
(EC 5.2.1.8) (Peptidyl-prolyl cis- trans isomerase)
(PPiase), partial (98%)
Length = 682
Score = 40.8 bits (94), Expect = 5e-04
Identities = 42/127 (33%), Positives = 63/127 (49%), Gaps = 5/127 (3%)
Frame = +2
Query: 121 EKEEEVT-LPNGIRY----CELKVGGGASPRPGDLVVIDIMGKVESTGEVFVNTFEGDKK 175
+KE +VT L G++Y CE++ GD V + GK+ + G VF ++FE +
Sbjct: 89 KKEADVTELQIGVKYKPKSCEVQA------HKGDKVKVHYRGKL-TDGTVFDSSFERNN- 244
Query: 176 PLALVMGSRPYSKGVCEGIEYVLKTMKAGGKRKVIVPPQLGFRENGADLGSGVQIPPLAT 235
P+ +G KG +G L M G KRK+ +P +LG+ E GS IP AT
Sbjct: 245 PIDFELGGGQVIKGWDQG----LLGMCLGEKRKLKIPAKLGYGEQ----GSPPTIPGGAT 400
Query: 236 LEYIVEV 242
L + E+
Sbjct: 401 LIFDTEL 421
>TC84391 similar to PIR|E86340|E86340 protein F2D10.32 [imported] -
Arabidopsis thaliana, partial (66%)
Length = 766
Score = 39.7 bits (91), Expect = 0.001
Identities = 33/157 (21%), Positives = 66/157 (42%), Gaps = 25/157 (15%)
Frame = +3
Query: 111 SQQEANTRDVEKEEEVTLPNGIRYCELKVGGGASPRPGDLVVIDI------MGKVESTGE 164
+++ +++ ++ +T P+G++Y + G G G V + + V S
Sbjct: 219 ARERRKKKNIPIDDYITSPDGLKYYDFLEGKGPIAEKGSTVQVHFDCLYRGITAVSSRES 398
Query: 165 VFVNTFEGDKKPLALVMGSRPYSKGVCEGIEY-------------------VLKTMKAGG 205
+ +P +G+ P + E ++ +++ M+ GG
Sbjct: 399 KLLAGNRVIAQPYEFKVGAPPGKERKREFVDNPNGLFSAQAAPKPPQAMYTIVEGMRVGG 578
Query: 206 KRKVIVPPQLGFRENGADLGSGVQIPPLATLEYIVEV 242
KR VIVPP+ G+ + G + +IPP AT E +E+
Sbjct: 579 KRTVIVPPEKGYGKKGMN-----EIPPGATFELNIEL 674
>TC78598 similar to SP|Q43207|FKB7_WHEAT 70 kDa peptidylprolyl isomerase (EC
5.2.1.8) (Peptidylprolyl cis-trans isomerase) (PPiase),
partial (92%)
Length = 2028
Score = 39.3 bits (90), Expect = 0.001
Identities = 42/139 (30%), Positives = 68/139 (48%), Gaps = 4/139 (2%)
Frame = +1
Query: 108 LEVSQQE---ANTRDVEKEEEVTLPNGIRYCELKVGGG-ASPRPGDLVVIDIMGKVESTG 163
+E+ ++E + T+ V +E+E+ NG++ +K G G +P GD V + G + G
Sbjct: 226 MEMPEEEEVMSPTQKVGEEKEIG-KNGLKKKLVKEGEGWETPDNGDQVEVHYTG-ILLDG 399
Query: 164 EVFVNTFEGDKKPLALVMGSRPYSKGVCEGIEYVLKTMKAGGKRKVIVPPQLGFRENGAD 223
F ++ + +G KG EGI KTMK G +PP+L + E+
Sbjct: 400 TKFDSSRDRGTT-FKFKLGQGQVIKGWDEGI----KTMKKGENAIFTIPPELAYGES--- 555
Query: 224 LGSGVQIPPLATLEYIVEV 242
GS IPP A L++ VE+
Sbjct: 556 -GSPPTIPPNAALQFDVEL 609
>TC89754 similar to GP|21618273|gb|AAM67323.1 unknown {Arabidopsis
thaliana}, partial (88%)
Length = 759
Score = 38.5 bits (88), Expect = 0.002
Identities = 19/58 (32%), Positives = 30/58 (50%)
Frame = +2
Query: 12 SLPLTRNSHHIASSSPTPPPIPPQQSQSPTASSSQSVQGTAKAQPQNPIKPVTSSTKV 69
S P ++HH + S+ P P PP S SP++SS S A + + PV++ K+
Sbjct: 152 SSPSPSSAHHASVSNSHPSPSPPPTSSSPSSSSPTSWSSPASSTTSSSNHPVSAQCKI 325
>TC77302 similar to GP|1354207|gb|AAB82061.1| rof1 {Arabidopsis thaliana},
partial (56%)
Length = 1221
Score = 38.1 bits (87), Expect = 0.003
Identities = 38/121 (31%), Positives = 54/121 (44%), Gaps = 1/121 (0%)
Frame = +1
Query: 123 EEEVTLPNGIRYCELKVGGG-ASPRPGDLVVIDIMGKVESTGEVFVNTFEGDKKPLALVM 181
EE+ G++ LK G G +P GD V + G + + + G +
Sbjct: 277 EEKEIGKQGLKKKLLKEGEGWDTPDVGDEVHVHYTGTLLDGTKFDSSRDRGTT--FNFTL 450
Query: 182 GSRPYSKGVCEGIEYVLKTMKAGGKRKVIVPPQLGFRENGADLGSGVQIPPLATLEYIVE 241
G KG EGI KTMK G +PP+L + E+ GS IPP ATL++ VE
Sbjct: 451 GQGQVIKGWDEGI----KTMKKGENALFTIPPELAYGES----GSPPTIPPNATLQFDVE 606
Query: 242 V 242
+
Sbjct: 607 L 609
>TC93489 homologue to PIR|T47848|T47848 hypothetical protein T8B10.30 -
Arabidopsis thaliana, partial (34%)
Length = 563
Score = 34.7 bits (78), Expect = 0.036
Identities = 17/40 (42%), Positives = 25/40 (62%)
Frame = +3
Query: 176 PLALVMGSRPYSKGVCEGIEYVLKTMKAGGKRKVIVPPQL 215
P + MG++ G EGI + M+AGGKR++I+PP L
Sbjct: 114 PAKIRMGNKALVPGFEEGI----RDMRAGGKRRIIIPPDL 221
>TC78212 similar to GP|17381160|gb|AAL36392.1 unknown protein {Arabidopsis
thaliana}, partial (67%)
Length = 1739
Score = 34.7 bits (78), Expect = 0.036
Identities = 22/69 (31%), Positives = 32/69 (45%), Gaps = 4/69 (5%)
Frame = +1
Query: 9 PVFSLPLTRNSHHIASSSPTPPPIPPQQSQSPTASSSQSVQGT----AKAQPQNPIKPVT 64
PV++ + NS SSSP P PQ PT++ +++ T AKA Q + +
Sbjct: 214 PVYTRQWSGNSSSTGSSSPVMSPAHPQSRLGPTSTGLSTIKRTQNVAAKAAAQRLARVMA 393
Query: 65 SSTKVDSTD 73
S T D D
Sbjct: 394 SQTVDDDDD 420
>AL373052 similar to GP|21593014|gb| putative aldolase {Arabidopsis
thaliana}, partial (19%)
Length = 459
Score = 34.3 bits (77), Expect = 0.046
Identities = 23/77 (29%), Positives = 36/77 (45%), Gaps = 10/77 (12%)
Frame = +3
Query: 8 PPVFSLPLTRNSHHIASSSPTPPPI--PPQQSQSPTASSS--------QSVQGTAKAQPQ 57
P + PL R H ++ SPT PP PPQ PTA+S+ + ++ + P
Sbjct: 129 PLTWQCPLER-IRHFSNPSPTTPPSQSPPQPPPHPTATSNPVSVLMKPSTASSSSPSPPL 305
Query: 58 NPIKPVTSSTKVDSTDW 74
+P P + +T S+ W
Sbjct: 306 SPKSPPSPATTSLSSTW 356
Score = 26.9 bits (58), Expect = 7.4
Identities = 13/32 (40%), Positives = 16/32 (49%)
Frame = +3
Query: 23 ASSSPTPPPIPPQQSQSPTASSSQSVQGTAKA 54
ASSS PP+ P+ SP +S S T A
Sbjct: 276 ASSSSPSPPLSPKSPPSPATTSLSSTWNTVTA 371
>TC82269 weakly similar to GP|14581464|gb|AAK31804.1 protocadherin 15 {Homo
sapiens}, partial (1%)
Length = 728
Score = 34.3 bits (77), Expect = 0.046
Identities = 23/72 (31%), Positives = 28/72 (37%), Gaps = 7/72 (9%)
Frame = +2
Query: 8 PPVFSLPLTRNSHHIASSSPTPPPIPPQQS-------QSPTASSSQSVQGTAKAQPQNPI 60
PP + P+ + SH S PTPP PP SP S++ S T
Sbjct: 170 PPSSTSPIPQISHQPPSPPPTPPNPPPPSKMKTLNPVHSPKWSNNSSTNATISQTTDTHH 349
Query: 61 KPVTSSTKVDST 72
K SSTK T
Sbjct: 350 KSTKSSTKTTFT 385
>TC87424 similar to GP|19699309|gb|AAL91265.1 AT4g10840/F25I24_50
{Arabidopsis thaliana}, partial (32%)
Length = 1014
Score = 33.9 bits (76), Expect = 0.061
Identities = 21/68 (30%), Positives = 35/68 (50%)
Frame = +2
Query: 6 GSPPVFSLPLTRNSHHIASSSPTPPPIPPQQSQSPTASSSQSVQGTAKAQPQNPIKPVTS 65
G PP +LP T +S +PT P+P ++ SP + SS+S + T++ N + S
Sbjct: 197 GPPPRITLPETPTQRSESSKTPTQSPLPNKKPPSP-SPSSRSKKKTSETPTTNSL---LS 364
Query: 66 STKVDSTD 73
+D+ D
Sbjct: 365 EASLDNPD 388
>TC81307 weakly similar to GP|4335772|gb|AAD17449.1| unknown protein
{Arabidopsis thaliana}, partial (31%)
Length = 1171
Score = 33.5 bits (75), Expect = 0.079
Identities = 24/64 (37%), Positives = 33/64 (51%), Gaps = 1/64 (1%)
Frame = +2
Query: 7 SPPVFSLPLTRNSHHIASSSPTPPPIPPQQSQSPTASSSQSVQGTAKAQPQNPI-KPVTS 65
S P F L L + S +SSSP+P P P S SP S++ A + P + + P +S
Sbjct: 491 SSPPFFLSLPQPSSPTSSSSPSPSPSPSSASPSP------SLKSFALSPPSSFLHSPFSS 652
Query: 66 STKV 69
ST V
Sbjct: 653 STSV 664
>TC80076 similar to GP|10177459|dbj|BAB10850. gene_id:MQB2.13~unknown
protein {Arabidopsis thaliana}, partial (21%)
Length = 910
Score = 33.1 bits (74), Expect = 0.10
Identities = 22/67 (32%), Positives = 34/67 (49%), Gaps = 11/67 (16%)
Frame = +1
Query: 12 SLPLTRN--SHHIASSSPTPPPIPPQQSQSPTASSSQSVQGTAKAQPQN---------PI 60
S PL ++ + +A+ SP+P P PP SPTA+ QG QP N +
Sbjct: 106 SQPLRKD*VASFMATQSPSPSPSPPPPLPSPTATDEN--QGADIIQPSNMDQQNAGDESV 279
Query: 61 KPVTSST 67
KP++S++
Sbjct: 280 KPISSTS 300
>TC81043 weakly similar to GP|6523547|emb|CAB62280.1 hydroxyproline-rich
glycoprotein DZ-HRGP {Volvox carteri f. nagariensis},
partial (19%)
Length = 555
Score = 32.3 bits (72), Expect = 0.18
Identities = 21/68 (30%), Positives = 30/68 (43%), Gaps = 6/68 (8%)
Frame = +1
Query: 6 GSPPVFSLPLTRNSHHIASSSPTPPPIPPQQSQSPTASSSQSVQGTAKAQPQ------NP 59
GSPP P + S + +P PPPIPP P+ S S + A P+ +P
Sbjct: 232 GSPPKPPAPPSPGSP--PAPAPAPPPIPPSPPAPPSPGSPPSPPPASPAPPKPTPPKGSP 405
Query: 60 IKPVTSST 67
+ P+ T
Sbjct: 406 LLPMPEKT 429
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.313 0.133 0.382
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 7,103,317
Number of Sequences: 36976
Number of extensions: 112404
Number of successful extensions: 2348
Number of sequences better than 10.0: 247
Number of HSP's better than 10.0 without gapping: 1785
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 2153
length of query: 250
length of database: 9,014,727
effective HSP length: 94
effective length of query: 156
effective length of database: 5,538,983
effective search space: 864081348
effective search space used: 864081348
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.9 bits)
S2: 57 (26.6 bits)
Lotus: description of TM0042.13


