

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0041.8
(562 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
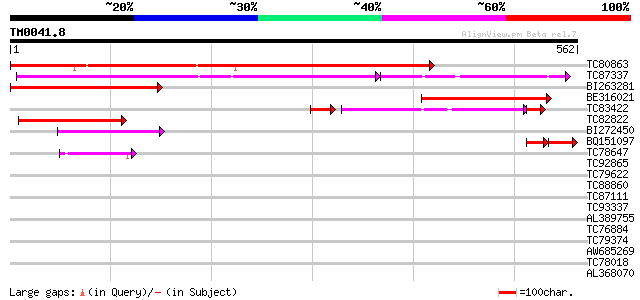
TC80863 similar to GP|498038|gb|AAA33934.1|| lipid transfer prot... 622 e-178
TC87337 similar to GP|10716610|gb|AAG21908.1 putative CER1 {Oryz... 290 e-106
BI263281 weakly similar to GP|498038|gb|A lipid transfer protein... 265 3e-71
BE316021 similar to GP|22900949|gb cuticle protein {Arabidopsis ... 255 4e-68
TC83422 weakly similar to GP|1209703|gb|AAB87721.1| maize gl1 ho... 107 2e-27
TC82822 similar to GP|2317909|gb|AAC24373.1| CER1-like protein {... 105 4e-23
BI272450 similar to GP|1199467|dbj possible aldehyde decarbonyla... 87 2e-17
BQ151097 46 2e-11
TC78647 similar to GP|20197199|gb|AAC95199.2 putative C-4 sterol... 50 2e-06
TC92865 homologue to GP|20197199|gb|AAC95199.2 putative C-4 ster... 40 0.003
TC79622 similar to PIR|B96718|B96718 probable sterol desaturase ... 35 0.10
TC88860 similar to GP|21593012|gb|AAM64961.1 putative C-4 sterol... 32 0.65
TC87111 similar to PIR|T02405|T02405 Nicotiana tabacum axi1 prot... 32 0.85
TC93337 weakly similar to GP|21240660|gb|AAM44371.1 hypothetical... 31 1.4
AL389755 similar to GP|21554158|gb amino acid permease-like prot... 31 1.4
TC76884 similar to SP|P26690|6DCS_SOYBN NAD(P)H dependent 6'-deo... 30 1.9
TC79374 similar to PIR|T45874|T45874 transcription factor NF-Y ... 30 3.2
AW685269 similar to GP|13899097|gb| Unknown protein {Arabidopsis... 29 5.5
TC78018 similar to PIR|T04499|T04499 auxin-induced protein homol... 28 7.2
AL368070 28 7.2
>TC80863 similar to GP|498038|gb|AAA33934.1|| lipid transfer protein
{Senecio odorus}, partial (75%)
Length = 1474
Score = 622 bits (1604), Expect = e-178
Identities = 305/425 (71%), Positives = 352/425 (82%), Gaps = 4/425 (0%)
Frame = +1
Query: 1 MLFLTRNRRILQQGVDFKQIDKEWDWDNFLILQTIIASMAYYMLPFLQNLPSWNVKGVIV 60
MLFLTRNRRILQQGVDFKQIDKEWDWDNFLILQT++A++ Y+ PFLQ+LP WN K
Sbjct: 205 MLFLTRNRRILQQGVDFKQIDKEWDWDNFLILQTLVATLVSYIFPFLQHLPLWNCKRYYC 384
Query: 61 AL--VLHVGISEPLYYWVHRKFHGDYLFTNYHSLHHSSPVPESLTVGHATIVEHAFISVV 118
L S + ++ R F GDYLF NYHSLHHSSPVP LT G+AT +EH + V
Sbjct: 385 CYDSTLWEFQSHFIIGYI-RNFTGDYLFKNYHSLHHSSPVPTPLTAGNATFLEHLILMAV 561
Query: 119 IGIPILGASVMGYGSASLVYGYVLGFDFLRCLGHCNVEVVPHGLFKTFPFMKYLLYTPTY 178
IGIPI GAS+MGYGS SLVYGYVL FDFLRCLGHCNVE++PH LFK FPF++Y+++TPTY
Sbjct: 562 IGIPIFGASMMGYGSGSLVYGYVLIFDFLRCLGHCNVEIIPHKLFKAFPFLRYVIHTPTY 741
Query: 179 HSLHHTDNKDTNFCLFMPLFDALGHTLNTKSWELHESFSSGKCS--RVPDFVFLAHIVDV 236
H+LHHT+ KDTNFCLFMPLFDALG+TLN SW LHE+ SSG + VP FVFLAH+VD+
Sbjct: 742 HNLHHTE-KDTNFCLFMPLFDALGNTLNKNSWTLHETLSSGSGNGATVPRFVFLAHMVDI 918
Query: 237 TSSMHAQFCLRSFASLPFRTRFFLIPFWPIAITTLLAMWVRSKAFLFSFYYLRDRLHQTW 296
+S MH F LRS +S + TR F IP P+ ++AMW+ SK FL SFYYLR RLHQTW
Sbjct: 919 SSCMHVPFVLRSASSFAYTTRLFFIPCLPVTFLVVMAMWLWSKTFLLSFYYLRGRLHQTW 1098
Query: 297 VVPRCGFQYFLPFAAEGINKKIEEAILTADKIGVKVISLAALNKNEALNGGGKLFVDKHP 356
VVPRCGFQYFLPFA +GINK+IE+AIL ADK+GVKVISLAALNKNE+LNGGGKLFVDKHP
Sbjct: 1099VVPRCGFQYFLPFATDGINKQIEQAILRADKMGVKVISLAALNKNESLNGGGKLFVDKHP 1278
Query: 357 NLRVRVVHGNTLTAAVIIDEIPKDVEEVFLTGATSKLGRAIALYLCQKKVKVLMLTLSAD 416
NL+VRVVHGNTLTAAVI++EIP+DVEEVFLTGATSKLGRAIALYLCQKKV+VLMLT+S D
Sbjct: 1279NLKVRVVHGNTLTAAVILNEIPQDVEEVFLTGATSKLGRAIALYLCQKKVRVLMLTISTD 1458
Query: 417 RFKRI 421
RF++I
Sbjct: 1459RFQKI 1473
>TC87337 similar to GP|10716610|gb|AAG21908.1 putative CER1 {Oryza sativa},
partial (70%)
Length = 2272
Score = 290 bits (743), Expect(2) = e-106
Identities = 150/362 (41%), Positives = 211/362 (57%), Gaps = 1/362 (0%)
Frame = +3
Query: 7 NRRILQQGVDFKQIDKEWDWDNFLILQTIIASMAYYMLPFLQNLPSWNVKGVIVALVLHV 66
N RIL +G++F Q+D+E DWD+ ++ ++ +A Y L LP W GVI+A++LH
Sbjct: 258 NNRILDKGIEFDQVDREKDWDDQILFNGLLYYLACYTLEGASRLPLWRTDGVIIAILLHA 437
Query: 67 GISEPLYYWVHRKFHGDYLFTNYHSLHHSSPVPESLTVGHATIVEHAFISVVIGIPILGA 126
G E LYYW+HR H +L++ YHS HHSS V E +T EH ++ IP L
Sbjct: 438 GAVEFLYYWLHRALHHHFLYSRYHSHHHSSIVTEPITSVIHPFAEHISYFLLFAIPKLTL 617
Query: 127 SVMGYGSASLVYGYVLGFDFLRCLGHCNVEVVPHGLFKTFPFMKYLLYTPTYHSLHHTDN 186
S + GYV DF+ LGHCN E+VP LF FP +KYL+YTP++HSLHHT
Sbjct: 618 VFTNRASIGAMVGYVTYIDFMNNLGHCNFEIVPKWLFDIFPPLKYLMYTPSFHSLHHTQF 797
Query: 187 KDTNFCLFMPLFDALGHTLNTKSWELHESFSSGKCSRVPDFVFLAHIVDVTSSMHAQFCL 246
+ TN+ LFMPL+D + T++ S ELHES K P+ V L H+ S H +F
Sbjct: 798 R-TNYSLFMPLYDYIYGTMDKASDELHESTLKRK-EETPNVVHLTHLTTPESIYHLRFGF 971
Query: 247 RSFASLPFRTRFFLIPFWPIAITTLLAMWVRSKAFLFSFYYLRDRLH-QTWVVPRCGFQY 305
+ AS P+ ++++L WP ++ WV + F+ D+L+ QTW +P+ QY
Sbjct: 972 AALASKPYTSKWYLWLMWPFTAWSMFLTWVYGRTFIVERNRFDDKLNLQTWAIPKYSVQY 1151
Query: 306 FLPFAAEGINKKIEEAILTADKIGVKVISLAALNKNEALNGGGKLFVDKHPNLRVRVVHG 365
+ INK IEEAIL ADK G+KV+SL +N+ E LN G L+V +HP L V+VV G
Sbjct: 1152SRQWQKVSINKMIEEAILDADKKGIKVVSLGLMNQEEELNIYGGLYVSRHPKLNVKVVDG 1331
Query: 366 NT 367
++
Sbjct: 1332SS 1337
Score = 114 bits (285), Expect(2) = e-106
Identities = 65/189 (34%), Positives = 99/189 (51%)
Frame = +1
Query: 368 LTAAVIIDEIPKDVEEVFLTGATSKLGRAIALYLCQKKVKVLMLTLSADRFKRIQKEAPQ 427
L A +I+ IPK +V L G +K+ A+A LCQ+ V++ T++ D + +++K
Sbjct: 1339 LAVAAVINSIPKGTTQVLLRGKLTKVAYALAYTLCQQGVQIA--TMNDDDYVKLKKALMH 1512
Query: 428 EYQSYLVQVTKYQAAQHCKTWIVGKWITPREQSWAPRGTHFHQFVVPPILPFRRDCTYGE 487
S +V + W+VG +T EQ AP+GT F + P R+DC Y
Sbjct: 1513 SSHSNIVNAKSFTQM----IWLVGDGLTEEEQLKAPKGTLFIPYSQFPPKKHRKDCLYHY 1680
Query: 488 LAAMRLPEDVEGLGSCEYTMERGVVHACHAGGVVHSLEGWTHHEVGAIDVDRIDVVWKAA 547
AM P +E + SCE + R V+ A G+VH LE W HE G ++ +D VW +A
Sbjct: 1681 TPAMLTPTSIENVHSCEDWLPRRVMSAWRVAGIVHCLEEWNEHECG-YNMINMDKVWPSA 1857
Query: 548 LKHGLRPVS 556
LKHG +P++
Sbjct: 1858 LKHGFKPLT 1884
>BI263281 weakly similar to GP|498038|gb|A lipid transfer protein {Senecio
odorus}, partial (28%)
Length = 681
Score = 265 bits (678), Expect = 3e-71
Identities = 119/151 (78%), Positives = 137/151 (89%)
Frame = +3
Query: 1 MLFLTRNRRILQQGVDFKQIDKEWDWDNFLILQTIIASMAYYMLPFLQNLPSWNVKGVIV 60
MLFLTRNR+IL+QGVDFKQ+DKEWDWDNFLILQTI+ASMAYYM PFLQNLP WN+KG+I
Sbjct: 216 MLFLTRNRQILKQGVDFKQLDKEWDWDNFLILQTILASMAYYMFPFLQNLPLWNIKGLIA 395
Query: 61 ALVLHVGISEPLYYWVHRKFHGDYLFTNYHSLHHSSPVPESLTVGHATIVEHAFISVVIG 120
AL+ HVGISEPLYYWVH+KFHG YLFTNYHSLHHS PVP+S+TVG T++EH F++VVIG
Sbjct: 396 ALMFHVGISEPLYYWVHKKFHGHYLFTNYHSLHHSIPVPQSVTVGTGTVLEHLFLTVVIG 575
Query: 121 IPILGASVMGYGSASLVYGYVLGFDFLRCLG 151
IPILGAS+MGYGS S++YGY+ FDFL LG
Sbjct: 576 IPILGASLMGYGSTSMIYGYIFFFDFLXSLG 668
>BE316021 similar to GP|22900949|gb cuticle protein {Arabidopsis thaliana},
partial (20%)
Length = 388
Score = 255 bits (651), Expect = 4e-68
Identities = 116/129 (89%), Positives = 123/129 (94%)
Frame = +1
Query: 409 LMLTLSADRFKRIQKEAPQEYQSYLVQVTKYQAAQHCKTWIVGKWITPREQSWAPRGTHF 468
LMLT+S DRF++IQKEAP EYQSYLVQVTKYQAAQ+CK WIVGKWITPREQSWAP GTHF
Sbjct: 1 LMLTISTDRFQKIQKEAPLEYQSYLVQVTKYQAAQNCKAWIVGKWITPREQSWAPSGTHF 180
Query: 469 HQFVVPPILPFRRDCTYGELAAMRLPEDVEGLGSCEYTMERGVVHACHAGGVVHSLEGWT 528
HQFVVPPIL FRRDCTYG+LAAMRLPEDVEGLG CEYTM+RGVVHACHAGGVVHSLEGWT
Sbjct: 181 HQFVVPPILSFRRDCTYGDLAAMRLPEDVEGLGCCEYTMDRGVVHACHAGGVVHSLEGWT 360
Query: 529 HHEVGAIDV 537
HHEVGA+DV
Sbjct: 361 HHEVGALDV 387
>TC83422 weakly similar to GP|1209703|gb|AAB87721.1| maize gl1 homolog
{Arabidopsis thaliana}, partial (13%)
Length = 683
Score = 107 bits (266), Expect(3) = 2e-27
Identities = 62/185 (33%), Positives = 100/185 (53%)
Frame = +3
Query: 330 VKVISLAALNKNEALNGGGKLFVDKHPNLRVRVVHGNTLTAAVIIDEIPKDVEEVFLTGA 389
VKV+SL N+ + LN G+L++ ++P L++++V G++L A++++ IPK+ +VFL G
Sbjct: 96 VKVLSLGLSNQGDLLNRYGELYIKRYPQLKMKIVDGSSLVVAIVLNSIPKEENQVFLCGR 275
Query: 390 TSKLGRAIALYLCQKKVKVLMLTLSADRFKRIQKEAPQEYQSYLVQVTKYQAAQHCKTWI 449
K+ AI LC++ KV T+ D + +Q + Q LV + + K W+
Sbjct: 276 LDKVSYAIVNALCERGTKV--TTMYRDDHENLQLRLSSKSQKNLV----FPGSNSAKIWL 437
Query: 450 VGKWITPREQSWAPRGTHFHQFVVPPILPFRRDCTYGELAAMRLPEDVEGLGSCEYTMER 509
VG EQ AP+G+ F F P FR+DC Y AM P ++ + SCE + R
Sbjct: 438 VGDQCEEVEQKKAPKGSLFVPFSQFPPKKFRKDCFYLXTPAMITPPNLANVHSCENWLPR 617
Query: 510 GVVHA 514
V+ A
Sbjct: 618 RVMSA 632
Score = 27.7 bits (60), Expect(3) = 2e-27
Identities = 11/20 (55%), Positives = 14/20 (70%), Gaps = 1/20 (5%)
Frame = +2
Query: 513 HACHA-GGVVHSLEGWTHHE 531
H C A G++H+LEGW HE
Sbjct: 623 HECMAIAGILHALEGWDVHE 682
Score = 26.6 bits (57), Expect(3) = 2e-27
Identities = 13/25 (52%), Positives = 15/25 (60%)
Frame = +2
Query: 299 PRCGFQYFLPFAAEGINKKIEEAIL 323
PR QY + E +NK IEEAIL
Sbjct: 2 PRFHVQYLFKWQRETLNKLIEEAIL 76
>TC82822 similar to GP|2317909|gb|AAC24373.1| CER1-like protein {Arabidopsis
thaliana}, partial (19%)
Length = 580
Score = 105 bits (263), Expect = 4e-23
Identities = 45/107 (42%), Positives = 67/107 (62%)
Frame = +1
Query: 9 RILQQGVDFKQIDKEWDWDNFLILQTIIASMAYYMLPFLQNLPSWNVKGVIVALVLHVGI 68
RI+ +G++F+Q+D+E WD+ ++ ++ +AY++ P NLP W + GVI+ +LH G
Sbjct: 229 RIVDKGLEFEQVDRETHWDDQMLFTVLVYCIAYFIFPMASNLPWWRIDGVILTAILHAGP 408
Query: 69 SEPLYYWVHRKFHGDYLFTNYHSLHHSSPVPESLTVGHATIVEHAFI 115
E LYYW+HR H YL++ YHS HHSS V E +T EH I
Sbjct: 409 VEFLYYWLHRALHHHYLYSRYHSHHHSSIVTEPITSVAHPFAEHLVI 549
>BI272450 similar to GP|1199467|dbj possible aldehyde decarbonylase
{Arabidopsis thaliana}, partial (18%)
Length = 320
Score = 86.7 bits (213), Expect = 2e-17
Identities = 40/106 (37%), Positives = 57/106 (53%)
Frame = +1
Query: 48 QNLPSWNVKGVIVALVLHVGISEPLYYWVHRKFHGDYLFTNYHSLHHSSPVPESLTVGHA 107
Q LP W GV++ ++LH G E LYYW+HR H +L++ YHS HHSS V +T
Sbjct: 1 QKLPIWRTSGVVMTILLHSGPVEFLYYWLHRALHHHFLYSRYHSHHHSSIVTXPITSVVH 180
Query: 108 TIVEHAFISVVIGIPILGASVMGYGSASLVYGYVLGFDFLRCLGHC 153
EH ++ IP+ ++ S + GY+ DF+ LGHC
Sbjct: 181 PFAEHIAYFLLFAIPLYTTAITNTASIASFAGYLAYIDFMNNLGHC 318
>BQ151097
Length = 355
Score = 46.2 bits (108), Expect(2) = 2e-11
Identities = 22/28 (78%), Positives = 24/28 (85%)
Frame = +3
Query: 535 IDVDRIDVVWKAALKHGLRPVSSGSTGK 562
I V RIDVVWKA LKHG+RP+SSGST K
Sbjct: 69 IYVYRIDVVWKATLKHGIRPLSSGSTVK 152
Score = 40.8 bits (94), Expect(2) = 2e-11
Identities = 17/22 (77%), Positives = 18/22 (81%)
Frame = +2
Query: 513 HACHAGGVVHSLEGWTHHEVGA 534
H HA GVVH+LEGWTHH VGA
Sbjct: 2 HEGHARGVVHNLEGWTHH*VGA 67
>TC78647 similar to GP|20197199|gb|AAC95199.2 putative C-4 sterol methyl
oxidase {Arabidopsis thaliana}, partial (94%)
Length = 1151
Score = 50.4 bits (119), Expect = 2e-06
Identities = 24/78 (30%), Positives = 43/78 (54%), Gaps = 2/78 (2%)
Frame = +3
Query: 50 LPSWNVKGVIVALVLHVGISEPLYYWVHRKFHGDYLFTNYHSLHHSSPVPESLTVGHATI 109
LPSWN+ ++ ++ + + + ++YW HR H +L+ + HS+HH P LT +A
Sbjct: 384 LPSWNI--ILTQIMFYFILEDFIFYWGHRILHTKWLYKHIHSVHHEYATPFGLTSEYAHP 557
Query: 110 VEHAFI--SVVIGIPILG 125
E F+ + ++G I G
Sbjct: 558 AEILFLGFATIVGPAITG 611
>TC92865 homologue to GP|20197199|gb|AAC95199.2 putative C-4 sterol methyl
oxidase {Arabidopsis thaliana}, partial (50%)
Length = 656
Score = 39.7 bits (91), Expect = 0.003
Identities = 19/55 (34%), Positives = 29/55 (52%), Gaps = 2/55 (3%)
Frame = +2
Query: 73 YYWVHRKFHGDYLFTNYHSLHHSSPVPESLTVGHATIVEHAFI--SVVIGIPILG 125
+YW HR H +L+ + HS+HH P LT +A E F+ + ++G I G
Sbjct: 2 FYWGHRILHTKWLYKHVHSVHHEYATPFGLTSEYAHPAEILFLGFATIVGPAITG 166
>TC79622 similar to PIR|B96718|B96718 probable sterol desaturase T6C23.16
[imported] - Arabidopsis thaliana, partial (58%)
Length = 937
Score = 34.7 bits (78), Expect = 0.10
Identities = 48/195 (24%), Positives = 77/195 (38%), Gaps = 6/195 (3%)
Frame = +3
Query: 74 YWVHRKFHGD-YLFTNYHSLHHSSPVPESLTVGHATIVEHAFISVVIGIPILGASVMGYG 132
Y++HR H + +L+ + HSLHH VP S + +E + + G S + G
Sbjct: 12 YFMHRYMHQNKFLYKHIHSLHHRLIVPYSFGALYNHPIEGLLLDTIGG----ALSFLLSG 179
Query: 133 SASLVYGYVLGFDFLRCL-GHCNVEVVPHGLFKTFPFMKYLLYTPTYHSLHHT--DNKDT 189
+ + F ++ + HC + +P LF F YH +HH NK
Sbjct: 180 MSPRASIFFFSFATIKTVDDHCGL-WLPGNLFHMF-----FNNNSAYHDVHHQLYGNKYN 341
Query: 190 NFCLFMPLFDALGHTLNTKSWELHES--FSSGKCSRVPDFVFLAHIVDVTSSMHAQFCLR 247
F ++D + T S E S F S C D V + + FCL
Sbjct: 342 FSQPFFVMWDKILGTHMPYSLEKRASGGFESRPCKAYKD-------E*VINMSN*SFCLM 500
Query: 248 SFASLPFRTRFFLIP 262
S ++ ++ FF +P
Sbjct: 501 SIFNV-YQYCFFFLP 542
>TC88860 similar to GP|21593012|gb|AAM64961.1 putative C-4 sterol methyl
oxidase {Arabidopsis thaliana}, partial (56%)
Length = 749
Score = 32.0 bits (71), Expect = 0.65
Identities = 15/73 (20%), Positives = 32/73 (43%)
Frame = +3
Query: 35 IIASMAYYMLPFLQNLPSWNVKGVIVALVLHVGISEPLYYWVHRKFHGDYLFTNYHSLHH 94
+I+ + M+ LP + ++ L+++ + + YW+HR H + + H +HH
Sbjct: 477 LISYPSIKMIGIRTGLPLPSGFEILSQLLVYFLVEDYTNYWIHRFLHNKWGYEKIHRVHH 656
Query: 95 SSPVPESLTVGHA 107
P +A
Sbjct: 657 EYQAPIGFAAPYA 695
>TC87111 similar to PIR|T02405|T02405 Nicotiana tabacum axi1 protein homolog
[imported] - Arabidopsis thaliana, partial (20%)
Length = 909
Score = 31.6 bits (70), Expect = 0.85
Identities = 32/143 (22%), Positives = 60/143 (41%), Gaps = 6/143 (4%)
Frame = +3
Query: 270 TLLAMWVRSKA--FLFSFYYLRDRLHQTWVVPRCGFQYFLPFAAEGINKKIEEAILTADK 327
T L W FLF+ ++L + + GF PF+ +E+++++
Sbjct: 186 TTLTKWKNPSRLFFLFTLFFLGG-----FALFMIGFNLKTPFSKSPCATNVEKSLISNGF 350
Query: 328 IGVKVISLAALNKNEALNGG----GKLFVDKHPNLRVRVVHGNTLTAAVIIDEIPKDVEE 383
+ + ++NKNE LNGG +L V ++P++ N + + + + +E
Sbjct: 351 GSKSELKVVSMNKNEILNGGVLVKSELGVVQNPSI------SNGFDSKSELGVVSVNKDE 512
Query: 384 VFLTGATSKLGRAIALYLCQKKV 406
V G+ S I + Q KV
Sbjct: 513 VLNDGSLSNSLPLITNVMLQAKV 581
>TC93337 weakly similar to GP|21240660|gb|AAM44371.1 hypothetical protein
{Dictyostelium discoideum}, partial (7%)
Length = 743
Score = 30.8 bits (68), Expect = 1.4
Identities = 22/90 (24%), Positives = 39/90 (42%), Gaps = 2/90 (2%)
Frame = -3
Query: 188 DTNFCLFMPLFDALGHTLNTKSWELHESFSSGKCSRVPDFVFLAHIVDVTSSMHA--QFC 245
D +FC+++ +F + + S G S +P +FL +++V + F
Sbjct: 375 DAHFCVYIRVFHTNCFCFSASCTSI---MSDGARSIIPLVLFLPSLMNVVAFNQTLISFI 205
Query: 246 LRSFASLPFRTRFFLIPFWPIAITTLLAMW 275
L F L FFL+PF +T + +W
Sbjct: 204 LAVFVLLLITLLFFLLPFIHKLLTAINHLW 115
>AL389755 similar to GP|21554158|gb amino acid permease-like protein
{Arabidopsis thaliana}, partial (36%)
Length = 509
Score = 30.8 bits (68), Expect = 1.4
Identities = 23/67 (34%), Positives = 28/67 (41%), Gaps = 11/67 (16%)
Frame = +2
Query: 149 CLGHC----NVEVVPHGLFKTFPF-------MKYLLYTPTYHSLHHTDNKDTNFCLFMPL 197
C G C + PHG K + F M L PT+HSL H N C L
Sbjct: 2 CCGQCLQIVYSNIAPHGALKLYDFIAMVTVVMIILSQLPTFHSLRH-----INLC---SL 157
Query: 198 FDALGHT 204
F +LG+T
Sbjct: 158 FLSLGYT 178
>TC76884 similar to SP|P26690|6DCS_SOYBN NAD(P)H dependent 6'-deoxychalcone
synthase (EC 1.1.-.-). [Soybean] {Glycine max}, partial
(95%)
Length = 1256
Score = 30.4 bits (67), Expect = 1.9
Identities = 21/60 (35%), Positives = 32/60 (53%), Gaps = 1/60 (1%)
Frame = +2
Query: 302 GFQYFLPFAAEGINKKIEEAILTADKIGVKVISLAALNKNEALNGGGKLFV-DKHPNLRV 360
G+++F AA G+ K + EAI A K+G+ ++ + L KL+V D HP L V
Sbjct: 215 GYRHFDAAAAYGVEKSVGEAIAEALKLGL-------ISSRDELFVTSKLWVTDNHPELIV 373
>TC79374 similar to PIR|T45874|T45874 transcription factor NF-Y
CCAAT-binding-like protein - Arabidopsis thaliana,
partial (56%)
Length = 997
Score = 29.6 bits (65), Expect = 3.2
Identities = 22/76 (28%), Positives = 42/76 (54%), Gaps = 9/76 (11%)
Frame = -1
Query: 355 HPNLRVRVVHGNTLTAAVIIDEIP-KDVEEVFLTGATSKLGRAIALYL-----CQ---KK 405
H L +R+ GNTL +++++ + + VFL S LGRA+++ L C+ K+
Sbjct: 625 HHMLTLRICVGNTL*ERTLMNKLRIRTLLNVFLARCVSTLGRALSITLHLSVSCKVNLKR 446
Query: 406 VKVLMLTLSADRFKRI 421
+ +++ S+ R K+I
Sbjct: 445 INIILKP*SSHRPKQI 398
>AW685269 similar to GP|13899097|gb| Unknown protein {Arabidopsis thaliana},
partial (36%)
Length = 634
Score = 28.9 bits (63), Expect = 5.5
Identities = 13/45 (28%), Positives = 22/45 (48%)
Frame = +1
Query: 168 FMKYLLYTPTYHSLHHTDNKDTNFCLFMPLFDALGHTLNTKSWEL 212
+MKY+ YT +H H + N C+F+ + + K+W L
Sbjct: 166 YMKYIPYTKIFHGFHLVNLHAHNNCIFIFIL*FIILKKKKKNWRL 300
>TC78018 similar to PIR|T04499|T04499 auxin-induced protein homolog
F8F16.140 - Arabidopsis thaliana, partial (58%)
Length = 803
Score = 28.5 bits (62), Expect = 7.2
Identities = 12/44 (27%), Positives = 25/44 (56%)
Frame = +3
Query: 427 QEYQSYLVQVTKYQAAQHCKTWIVGKWITPREQSWAPRGTHFHQ 470
Q+Y++Y+ Q++ +A H T I+ WI R + + + F++
Sbjct: 69 QKYKAYINQIS*TEAHHHIYTSIISTWIHQRNLTKSEKSLDFNR 200
>AL368070
Length = 234
Score = 28.5 bits (62), Expect = 7.2
Identities = 12/43 (27%), Positives = 22/43 (50%), Gaps = 5/43 (11%)
Frame = +2
Query: 272 LAMWVRSKAFLFSFYYLRDRLHQTWV-----VPRCGFQYFLPF 309
L W+ S F+++ + L++R H W+ + C QY L +
Sbjct: 77 LVYWIVS*CFIWTLFLLKNRNHVLWIFFNINIRSCNKQYLLTY 205
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.326 0.141 0.450
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 21,477,943
Number of Sequences: 36976
Number of extensions: 367451
Number of successful extensions: 2566
Number of sequences better than 10.0: 43
Number of HSP's better than 10.0 without gapping: 2524
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 2555
length of query: 562
length of database: 9,014,727
effective HSP length: 101
effective length of query: 461
effective length of database: 5,280,151
effective search space: 2434149611
effective search space used: 2434149611
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.7 bits)
S2: 61 (28.1 bits)
Lotus: description of TM0041.8


