

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
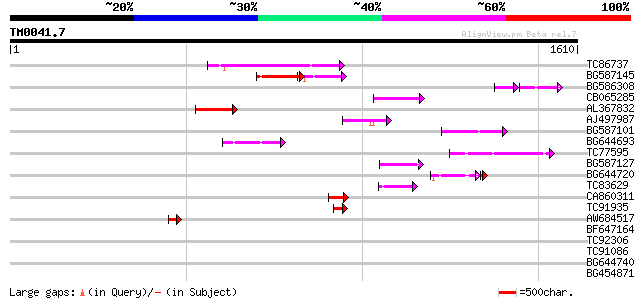
Query= TM0041.7
(1610 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC86737 weakly similar to GP|6683624|dbj|BAA89272.1 Pol {Alterna... 189 6e-48
BG587145 similar to PIR|H86337|H8 protein F5M15.26 [imported] - ... 132 1e-30
BG586308 weakly similar to PIR|F84528|F8 probable retroelement p... 67 1e-24
CB065285 weakly similar to PIR|A84500|A84 probable retroelement ... 93 1e-18
AL367832 weakly similar to GP|14091845|gb Putative retroelement ... 92 1e-18
AJ497987 weakly similar to GP|9927273|dbj Similar to Arabidopsis... 91 3e-18
BG587101 similar to GP|6691191|gb F7F22.15 {Arabidopsis thaliana... 91 5e-18
BG644693 weakly similar to GP|18767374|g Putative 22 kDa kafirin... 90 6e-18
TC77595 weakly similar to PIR|T18350|T18350 probable pol polypro... 75 3e-13
BG587127 weakly similar to PIR|H84506|H84 probable retroelement ... 65 2e-10
BG644720 47 5e-07
TC83629 weakly similar to GP|6692261|gb|AAF24611.1| putative RNa... 50 9e-06
CA860311 weakly similar to GP|7289872|gb|A CG17427 gene product ... 45 3e-04
TC91935 45 3e-04
AW684517 similar to GP|9927273|dbj Similar to Arabidopsis thalia... 44 5e-04
BF647164 weakly similar to GP|6691191|gb F7F22.15 {Arabidopsis t... 41 0.003
TC92306 40 0.010
TC91086 36 0.11
BG644740 similar to PIR|A84460|A84 probable retroelement pol pol... 36 0.11
BG454871 weakly similar to GP|10140673|g putative gag-pol polypr... 32 1.5
>TC86737 weakly similar to GP|6683624|dbj|BAA89272.1 Pol {Alternaria
alternata}, partial (21%)
Length = 1540
Score = 189 bits (481), Expect = 6e-48
Identities = 133/398 (33%), Positives = 206/398 (51%), Gaps = 9/398 (2%)
Frame = +1
Query: 562 LLKEFKDCFA*DNDEMPGLSRDLVELQLPIKEDKKLVK*LPRRFHP------DVLVKIKE 615
+L+EF D F + SR L++ +P+ DK P + P L+ +K+
Sbjct: 343 VLEEFPDLFNPEKAYQVPASRGLLDHAIPLIPDKD-GNDPPLPWGPLYGMSRQELLVLKK 519
Query: 616 EIERLLRCKFIRAAKYVEWLANVVPVIKKNGKMRVCIDFRDLNAATPKDEYHMPIAEMMV 675
+E LL FI+A+ A V+ V K G +R C+D+R LNA T KD Y +P+ +
Sbjct: 520 TLEDLLDKGFIKASGSAAG-APVLFVRKPGGGIRFCVDYRALNAITKKDRYPLPLISETL 696
Query: 676 DSAAGHEYLSLLDGYSGYNQIFIAKEDVSKTAFRCSGALGTYEWVVMPFGLKNAGATYQR 735
AG + + LD + ++++ I ED KTAFR G +EW+V PFGL A AT+QR
Sbjct: 697 RRVAGARWFTKLDVVAAFHKMRIKDEDQEKTAFRT--RYGLFEWIVCPFGLTGAPATFQR 870
Query: 736 VMNTIFHDFIETFMQVYIDD-LVVKSPSRDEHLSHLRKSFERMRIHGLKMNPLKCAFGVI 794
+N H+F++ F+ YIDD L+ + S+ +H + +R+ R+ GL ++P KC F V
Sbjct: 871 YINKTLHEFLDDFVTAYIDDVLIYTTGSKKDHEAQVRRVLRRLADAGLSLDPKKCEFSVT 1050
Query: 795 AGDFLGFVVHK-KGIEINKNKAKAILDTSPPTSKKQLQSLLGKVNFLRRFIDNLSDKTKP 853
++GF++ KG+ + K AI D PP S K +S LG N+ + FI S+ T+P
Sbjct: 1051TVKYVGFILTAGKGVSCDPLKLAAIRDWLPPGSVKGARSFLGFCNYYKDFIPGYSEITEP 1230
Query: 854 FSSLLRLKREDI-FRWEA*HQKAFDELKNYLAIPPVMIPPIKGKPMRLYISATDETIGSM 912
L RL R+D FRW A + AF +LK A PV+ + + +G +
Sbjct: 1231---LTRLTRKDFPFRWGAEQEAAFTKLKRLFAEEPVLRMFDPEAVTTVETDCSGFALGGV 1401
Query: 913 LAQEDEDGIERAVFYLSRVLNDAKTRYTMIEKLCLCLY 950
L QED G V + S+ L+ A+ Y + +K L ++
Sbjct: 1402LTQEDGTGAAHPVAFHSQRLSPAEYNYPIHDKELLAVW 1515
>BG587145 similar to PIR|H86337|H8 protein F5M15.26 [imported] - Arabidopsis
thaliana, partial (13%)
Length = 763
Score = 132 bits (332), Expect = 1e-30
Identities = 64/136 (47%), Positives = 97/136 (71%)
Frame = +2
Query: 701 EDVSKTAFRCSGALGTYEWVVMPFGLKNAGATYQRVMNTIFHDFIETFMQVYIDDLVVKS 760
+D+ KTAF GTY + VMPFGLKNAG+TYQR++N +F D + M+VYIDD++VKS
Sbjct: 11 DDLEKTAFITDR--GTYCYKVMPFGLKNAGSTYQRLVNRMFADKLGNTMEVYIDDMLVKS 184
Query: 761 PSRDEHLSHLRKSFERMRIHGLKMNPLKCAFGVIAGDFLGFVVHKKGIEINKNKAKAILD 820
+HL+HL++ F+ + + +K+NP KC FGV +G+FLG++V ++GIE+N + AILD
Sbjct: 185 LRATDHLNHLKE*FKTLDEYIMKLNPAKCTFGVTSGEFLGYIVTQQGIEVNPKQITAILD 364
Query: 821 TSPPTSKKQLQSLLGK 836
P + +++Q L G+
Sbjct: 365 LPSPKNSREVQRLTGE 412
Score = 81.6 bits (200), Expect = 2e-15
Identities = 49/145 (33%), Positives = 76/145 (51%), Gaps = 7/145 (4%)
Frame = +3
Query: 818 ILDTSPPTSKKQLQSLL-------GKVNFLRRFIDNLSDKTKPFSSLLRLKREDIFRWEA 870
IL SPP +Q + G++ L RFI +DK PF LL + F W+
Sbjct: 336 ILSRSPPY*TSLVQRIAERSSDSRGRIAALNRFISRSTDKCLPFYKLLCGNKR--FVWDE 509
Query: 871 *HQKAFDELKNYLAIPPVMIPPIKGKPMRLYISATDETIGSMLAQEDEDGIERAVFYLSR 930
++AF++LK YL PPV+ P G + LYI+ + + S+L +ED G ++ +FY S+
Sbjct: 510 KCEEAFEQLKQYLTTPPVLSKPEAGDTLSLYIAISSTAVSSVLIREDR-GEQKPIFYTSK 686
Query: 931 VLNDAKTRYTMIEKLCLCLYFSCTK 955
+ D +TRY +EK+ + S K
Sbjct: 687 RMTDPETRYPTLEKMAFAVITSARK 761
>BG586308 weakly similar to PIR|F84528|F8 probable retroelement pol polyprotein
[imported] - Arabidopsis thaliana, partial (7%)
Length = 686
Score = 66.6 bits (161), Expect(2) = 1e-24
Identities = 44/122 (36%), Positives = 68/122 (55%), Gaps = 1/122 (0%)
Frame = -3
Query: 1448 VLWAYRNSPREATGTTPFRLAYGQEAVLPAEVYLQSF-RIQRQEDIPSEVYWNMMLDELV 1506
VLW++R +PR AT +TPF +A+ EA+ PAEV + S R + ++I E+ + + + L
Sbjct: 465 VLWSHRTNPRGATKSTPFSMAHRVEAMAPAEVNVTSL*RSRMPQNI--ELNNDRLFNALE 292
Query: 1507 NLDEERVLALDVLTRQKDRIAKAYNKKVKNRSFVTGDYVWKVILPTDKKDRAYGKWAPNW 1566
++E R AL + + +I YNK VK++ GD V + K+ A GK NW
Sbjct: 291 TIEERRDQALLRIQNYQHQIESYYNKTVKSQPLKLGDIVLCKVFENTKELNA-GKLGTNW 115
Query: 1567 EG 1568
EG
Sbjct: 114 EG 109
Score = 66.6 bits (161), Expect(2) = 1e-24
Identities = 31/70 (44%), Positives = 42/70 (59%)
Frame = -2
Query: 1376 GLPESLTTDQGTMFVGRKVAAFAESWGIKLLTSIPYYAQANGQVEAANKILISLIKKHVG 1435
GLP + TD G+ F+ K F E W I+L T+ P Y Q+NGQ EA+NKI+I +KK +
Sbjct: 682 GLPYEIVTDNGSHFISNKFREFCERWRIRLNTASPRYPQSNGQAEASNKIIIDGLKKRLD 503
Query: 1436 RKPKSWHESL 1445
K W + L
Sbjct: 502 LKKGCWADEL 473
>CB065285 weakly similar to PIR|A84500|A84 probable retroelement gag/pol
polyprotein [imported] - Arabidopsis thaliana, partial
(2%)
Length = 592
Score = 92.8 bits (229), Expect = 1e-18
Identities = 52/146 (35%), Positives = 81/146 (54%)
Frame = -2
Query: 1033 WKLFFDDSSHKEGSGIGMFIVSPQGIPTKFMFRIRESCSNNESEYEALVSGLEILLALGA 1092
W L FD + + G GIG IVSPQG F RI C+NN +EYEA + G+E + +
Sbjct: 534 WGLVFDGAVNAYGKGIGAVIVSPQGHYIPFTARILFECTNNMAEYEACIFGIEEAIDMRI 355
Query: 1093 KNVEVKGDSELVVKQLTKEYKCISKNLAKYYVKAMSLLANFDQAGVSYIPRVSNQEANEL 1152
K++++ GDS LV+ Q+ E++ NL Y A LL F + + +IPR NQ A+ L
Sbjct: 354 KHLDIYGDSALVINQIKGEWETHHANLIPYRDYARRLLTYFTKVELHHIPRDENQMADAL 175
Query: 1153 AQIASGYMIDKQKLKELIRIKEKLSP 1178
A ++S + ++ +I+++ P
Sbjct: 174 ATLSSMFRVNHWNDVPIIKVQRLERP 97
>AL367832 weakly similar to GP|14091845|gb Putative retroelement {Oryza
sativa}, partial (2%)
Length = 384
Score = 92.4 bits (228), Expect = 1e-18
Identities = 43/119 (36%), Positives = 76/119 (63%)
Frame = -1
Query: 528 QDPLEKVDLGDGSKKRPTYISSLIDPELRGRMIELLKEFKDCFA*DNDEMPGLSRDLVEL 587
Q+ +E ++LG KR + + ++ ++ ++ +LL+E+ D FA ++MPGL +VE
Sbjct: 357 QEEIELINLGTEENKREIKVGAALEEGVKRKIFQLLREYLDIFACSYEDMPGLDPKIVEH 178
Query: 588 QLPIKEDKKLVK*LPRRFHPDVLVKIKEEIERLLRCKFIRAAKYVEWLANVVPVIKKNG 646
++P K + V+* RR HPD+ +KIK E+++ + F+ +Y EW+AN+VPV KK+G
Sbjct: 177 RIPTKPECPPVR*KLRRTHPDMALKIKSEVQKQIDAGFLMTVEYPEWVANIVPVPKKDG 1
>AJ497987 weakly similar to GP|9927273|dbj Similar to Arabidopsis thaliana
chromosome II BAC F26H6; putative retroelement pol
polyprotein, partial (1%)
Length = 636
Score = 91.3 bits (225), Expect = 3e-18
Identities = 53/167 (31%), Positives = 82/167 (48%), Gaps = 26/167 (15%)
Frame = -2
Query: 944 KLCLCLYFSCTKLKYYIKPIDVMVFSHFDIIKHMLSKPILHSRIGKWSLALTEYSLTYAP 1003
K C L ++ +L++Y+ + S D IK++ KP L RI +W + L+EY + Y
Sbjct: 635 KTCCALAWAAKRLRHYMINHTTWLVSKMDPIKYIFEKPALTGRIARWQMLLSEYDIEYRS 456
Query: 1004 LKTIKGQAVADFLADHTL-----------PEEIAYVGLQP---------------WKLFF 1037
K IKG +AD LA L EEI Y+ ++ W L F
Sbjct: 455 QKAIKGSILADHLAHQPLEDYRPIKFDFPDEEIMYLKMKDCDEPLFGEGPDPDSVWGLIF 276
Query: 1038 DDSSHKEGSGIGMFIVSPQGIPTKFMFRIRESCSNNESEYEALVSGL 1084
D + + G+GIG +++P+G F R+R C+NN +EYEA + G+
Sbjct: 275 DGAVNVYGNGIGAVLLTPKGAHIPFTARLRFDCTNNIAEYEACIMGI 135
>BG587101 similar to GP|6691191|gb F7F22.15 {Arabidopsis thaliana}, partial
(10%)
Length = 624
Score = 90.5 bits (223), Expect = 5e-18
Identities = 52/188 (27%), Positives = 90/188 (47%)
Frame = +2
Query: 1227 LFKKNVDGTLLKCLSEVDAFIAVSAAHGGLCGAHQAGAKMKWILFRQGMYWPTIMKDCIE 1286
L+K+ D ++C++E + + HG H A +K + + G +WPT+ KD
Sbjct: 41 LYKQCADNIYIRCVAEEEIPGILFHCHGSNYAGHFAVSKTVSKIQQAGFWWPTMFKDAHS 220
Query: 1287 YAKGCQDCQKHSGIQHVPASELHSIIKSWPFRGWAIDLIGEIHPALSRQHKYIIVAIDYF 1346
+ C CQ+ I + I++ F W ID +G S +KYI+VA+DY
Sbjct: 221 FISKCDPCQRQGNIS*RNEMPQNFILEVEVFDVWGIDFMGPFPS--SYNNKYILVAVDYV 394
Query: 1347 IKWVEAIPLQSVTQETVIEFIQNHIVYRFGLPESLTTDQGTMFVGRKVAAFAESWGIKLL 1406
KWVEAI + V++ ++ I RFG+P + +D G+ F+ + + G++
Sbjct: 395 SKWVEAIASPTNDATVVVKMFKSVIFPRFGVPRVVISDGGSHFINKVFEKLLKKNGVRHK 574
Query: 1407 TSIPYYAQ 1414
+ Y+ Q
Sbjct: 575 VATAYHPQ 598
>BG644693 weakly similar to GP|18767374|g Putative 22 kDa kafirin cluster;
Ty3-Gypsy type {Oryza sativa}, partial (15%)
Length = 716
Score = 90.1 bits (222), Expect = 6e-18
Identities = 56/179 (31%), Positives = 102/179 (56%)
Frame = +2
Query: 604 RFHPDVLVKIKEEIERLLRCKFIRAAKYVEWLANVVPVIKKNGKMRVCIDFRDLNAATPK 663
R +P L +K +++ LL FI+ + Y + V+ + KK+G +R+ ID+ LN K
Sbjct: 92 RINPLKLKVLKLQLKDLLEKGFIQPSIYP*GVV-VLFLKKKDGFLRMSIDYPQLNNVNIK 268
Query: 664 DEYHMPIAEMMVDSAAGHEYLSLLDGYSGYNQIFIAKEDVSKTAFRCSGALGTYEWVVMP 723
+Y +P+ + + D+ G ++ +D G +Q + EDV KTAFR G YE +VM
Sbjct: 269 IKYPLPLIDELFDNLQGSKWFFKIDLRLG*HQHRVIGEDVPKTAFRIR--YGHYEILVMS 442
Query: 724 FGLKNAGATYQRVMNTIFHDFIETFMQVYIDDLVVKSPSRDEHLSHLRKSFERMRIHGL 782
FG N + +MN +F D++++ + V+ +D+++ S + +EH +HLR + + ++ GL
Sbjct: 443 FG*TNPPMAFMELMNRVFQDYLDSLVIVFSNDILIYSKNENEHENHLRLALKVLKDIGL 619
>TC77595 weakly similar to PIR|T18350|T18350 probable pol polyprotein - rice
blast fungus gypsy retroelement (fragment), partial (14%)
Length = 1708
Score = 74.7 bits (182), Expect = 3e-13
Identities = 71/305 (23%), Positives = 125/305 (40%), Gaps = 7/305 (2%)
Frame = +2
Query: 1249 VSAAHGGLCGAHQAGAKMKWILFRQGMYWPTIMKDCIEYAKGCQDCQKHSGIQHVPASEL 1308
V +H H I+ R+ +WP + + + C C G H+
Sbjct: 182 VQESHDSTAAGHPGRNGTLEIVSRK-FFWPGQSQTVRRFVRNCDVC----GGIHIWRQAK 346
Query: 1309 HSIIKSWPFRG-----WAIDLIGEIHPALSRQHKYIIVAIDYFIKWVEAIPLQSVTQETV 1363
+K P ++D I + P R +Y+ V +D K V + ++ E
Sbjct: 347 RGFLKPLPVPNRLHSDLSMDFITSLPPTRGRGSQYLWVIVDRLSKSVTLEEMDTMEAEAC 526
Query: 1364 IE-FIQNHIVYRF-GLPESLTTDQGTMFVGRKVAAFAESWGIKLLTSIPYYAQANGQVEA 1421
+ F+ H YRF G+P+S+ +D+G+ +VGR F G+ L S Y+ Q +G E
Sbjct: 527 AQRFLSCH--YRFHGMPQSIVSDRGSNWVGRFWREFCRLTGVTQLLSTSYHPQTDGGTER 700
Query: 1422 ANKILISLIKKHVGRKPKSWHESLSQVLWAYRNSPREATGTTPFRLAYGQEAVLPAEVYL 1481
N+ + ++++ +V +W + L V A RN + G TPF + +G V P
Sbjct: 701 WNQEIQAVLRAYVCWSQDNWGDLLPTVQLALRNRHNSSIGATPFFVEHGYH-VDPIPTVE 877
Query: 1482 QSFRIQRQEDIPSEVYWNMMLDELVNLDEERVLALDVLTRQKDRIAKAYNKKVKNRSFVT 1541
+ + + + +++ M D + E V A Q+ A A ++ +
Sbjct: 878 DTGGVVSEGEAAAQLLVKRMKDVTGFIQAEIVAA------QQRSEASANKRRCPADRYQV 1039
Query: 1542 GDYVW 1546
GD VW
Sbjct: 1040 GDKVW 1054
>BG587127 weakly similar to PIR|H84506|H84 probable retroelement pol
polyprotein [imported] - Arabidopsis thaliana, partial
(13%)
Length = 415
Score = 65.5 bits (158), Expect = 2e-10
Identities = 39/125 (31%), Positives = 63/125 (50%)
Frame = +3
Query: 1049 GMFIVSPQGIPTKFMFRIRESCSNNESEYEALVSGLEILLALGAKNVEVKGDSELVVKQL 1108
G+ + SP K FR+ SNNE+ YEAL++G+ + L +N+ DS+LV Q
Sbjct: 3 GIRLTSPTNEILKQSFRLEFHASNNETNYEALIAGVRLAHGLKIRNIHAYCDSQLVASQF 182
Query: 1109 TKEYKCISKNLAKYYVKAMSLLANFDQAGVSYIPRVSNQEANELAQIASGYMIDKQKLKE 1168
+ EY+ + + Y L D ++ IPR N +A+ LA +AS +LK
Sbjct: 183 SGEYEARDELMDTYLKLVQKLAQKLDYFALTRIPRSENVQADALAALASS---SDPELKR 353
Query: 1169 LIRIK 1173
+I ++
Sbjct: 354 VIPVR 368
>BG644720
Length = 678
Score = 47.0 bits (110), Expect(2) = 5e-07
Identities = 46/150 (30%), Positives = 65/150 (42%), Gaps = 7/150 (4%)
Frame = -3
Query: 1194 WRKPIVEY------LQNPVGSTDIKVKYRALNYTIIGNELFKKNVDGTLLKCLSEVDAFI 1247
WR+PI+ Y L+NP ST I+ L+ L+ +G LL CL E +
Sbjct: 463 WRQPIIIYMCYGIFLENPRRSTYIRRHTPHLH*--YNETLYI*LFEGVLL*CLGEEETIQ 290
Query: 1248 AVSAAHGGLCGAHQAGAKMKWILFRQGMYWPT-IMKDCIEYAKGCQDCQKHSGIQHVPAS 1306
A AH +CG+H+ +K+ + + R G YW T IM A Q P
Sbjct: 289 AFQ*AHSRVCGSHK--SKLYFHIKRMGYYWNT*IMLKGALLANFIQQ----------PPE 146
Query: 1307 ELHSIIKSWPFRGWAIDLIGEIHPALSRQH 1336
LH I S F W +++ G + P S H
Sbjct: 145 VLHPTITS*LFESWELNVFG-LFPKSSGGH 59
Score = 26.6 bits (57), Expect(2) = 5e-07
Identities = 10/18 (55%), Positives = 13/18 (71%)
Frame = -1
Query: 1338 YIIVAIDYFIKWVEAIPL 1355
YI+ A+ YF KWVE + L
Sbjct: 54 YILAAVYYF*KWVEVVAL 1
>TC83629 weakly similar to GP|6692261|gb|AAF24611.1| putative RNase H
{Arabidopsis thaliana}, partial (21%)
Length = 1071
Score = 49.7 bits (117), Expect = 9e-06
Identities = 34/120 (28%), Positives = 56/120 (46%), Gaps = 7/120 (5%)
Frame = +2
Query: 1046 SGIGMFIVSPQGIPTKFMFRIRESCSNN---ESEYEALVSGLEILLALGAKNVEVKGDSE 1102
+G G +++ G ++ R+ + +EY AL+ GL+ G K V KGDSE
Sbjct: 599 AGAGALLLAEDG---SLLYGFRQGLGHQTKESAEYRALLLGLKHASMKGFKYVTAKGDSE 769
Query: 1103 LVVKQLTKEYKCISKNLAKYYVKAMSLLANFDQAGVSYIPRVSN----QEANELAQIASG 1158
LV+ Q+ +K ++L K +A+ L NF + +I R N + AN + G
Sbjct: 770 LVINQILDPWKIKDEHLKKLCAEALELSDNFHSFRIQHISRERNYGAVRNANRAINLTDG 949
>CA860311 weakly similar to GP|7289872|gb|A CG17427 gene product {Drosophila
melanogaster}, partial (20%)
Length = 192
Score = 44.7 bits (104), Expect = 3e-04
Identities = 20/57 (35%), Positives = 35/57 (61%)
Frame = +1
Query: 904 ATDETIGSMLAQEDEDGIERAVFYLSRVLNDAKTRYTMIEKLCLCLYFSCTKLKYYI 960
A D IG+ L+Q+DE+ E + Y SR+L A+ YT++E+ CL ++ ++Y+
Sbjct: 1 APDWGIGAGLSQKDEENHEHPIAYASRLLTAAERNYTVVERECLAAIWAIRNFRHYL 171
>TC91935
Length = 1144
Score = 44.7 bits (104), Expect = 3e-04
Identities = 18/40 (45%), Positives = 26/40 (65%)
Frame = +3
Query: 919 DGIERAVFYLSRVLNDAKTRYTMIEKLCLCLYFSCTKLKY 958
DG + A +YLS +L D T+YT I+KLCL ++C + Y
Sbjct: 984 DGTKIAAYYLSWILKDVGTKYTHIDKLCLSFLYACNNISY 1103
>AW684517 similar to GP|9927273|dbj Similar to Arabidopsis thaliana
chromosome II BAC F26H6; putative retroelement pol
polyprotein, partial (1%)
Length = 488
Score = 43.9 bits (102), Expect = 5e-04
Identities = 20/37 (54%), Positives = 25/37 (67%)
Frame = +1
Query: 451 FIVVPSKANYNLLLGREWIHGVGAVPSTLHQRISIWK 487
F V+ A+Y+ LLGR WIH GAV STLHQ++ K
Sbjct: 7 FQVMDINASYSCLLGRPWIHDAGAVTSTLHQKLKFVK 117
>BF647164 weakly similar to GP|6691191|gb F7F22.15 {Arabidopsis thaliana},
partial (6%)
Length = 469
Score = 41.2 bits (95), Expect = 0.003
Identities = 32/101 (31%), Positives = 51/101 (49%), Gaps = 1/101 (0%)
Frame = +1
Query: 1504 ELVNLDEERVLALDVLTRQKDRIAKAYNKKVKNRSFVTGDYVWKVILPTDKKDRAYGKWA 1563
EL L+E R+ A + K+R K +++++ R F G+ V +L + GK
Sbjct: 103 ELNELEELRLDAYENAKIYKERTKKWHDRRIIRREFREGELV---LLFNSRLKLFPGKLR 273
Query: 1564 PNWEGPFKVKKVLSNNAY-VIKELSGQRQFVTINDKYLKAY 1603
+W GPF+VK V+ + A V E +G T+N + LK Y
Sbjct: 274 SHWSGPFQVKNVMPSGAVEVWSESTGP---FTVNGQRLKHY 387
>TC92306
Length = 521
Score = 39.7 bits (91), Expect = 0.010
Identities = 23/73 (31%), Positives = 35/73 (47%)
Frame = +2
Query: 1081 VSGLEILLALGAKNVEVKGDSELVVKQLTKEYKCISKNLAKYYVKAMSLLANFDQAGVSY 1140
+ GL G ++V V+GD + V KQ +K + NL A+ L +NF V +
Sbjct: 2 ILGLNEARNQGYEHVHVRGDFQXVCKQFXGSWKVNNPNLRNLCNXAVELKSNFKSVSVEH 181
Query: 1141 IPRVSNQEANELA 1153
+PR N A+ A
Sbjct: 182 VPRGXNXAADAQA 220
>TC91086
Length = 697
Score = 36.2 bits (82), Expect = 0.11
Identities = 11/22 (50%), Positives = 21/22 (95%)
Frame = -1
Query: 625 FIRAAKYVEWLANVVPVIKKNG 646
FI+ ++Y++WL+N++P+IKK+G
Sbjct: 676 FIQTSRYIKWLSNILPIIKKHG 611
>BG644740 similar to PIR|A84460|A84 probable retroelement pol polyprotein
[imported] - Arabidopsis thaliana, partial (4%)
Length = 754
Score = 36.2 bits (82), Expect = 0.11
Identities = 15/36 (41%), Positives = 23/36 (63%)
Frame = -1
Query: 641 VIKKNGKMRVCIDFRDLNAATPKDEYHMPIAEMMVD 676
V KK+G R+CID+R N T K++Y +P + + D
Sbjct: 223 VRKKDGYFRMCIDYRQFNKVTTKNKYPLPRIDNLFD 116
>BG454871 weakly similar to GP|10140673|g putative gag-pol polyprotein {Oryza
sativa (japonica cultivar-group)}, partial (7%)
Length = 674
Score = 32.3 bits (72), Expect = 1.5
Identities = 19/71 (26%), Positives = 32/71 (44%)
Frame = +2
Query: 1402 GIKLLTSIPYYAQANGQVEAANKILISLIKKHVGRKPKSWHESLSQVLWAYRNSPREATG 1461
G L S Y+ ++GQ EA NK ++ + P W ++ + Y S +
Sbjct: 65 GTILTMSSAYHP*SDGQSEALNKGXEMYLRCLMFTDPLKWSKAFPWAEYWYNTSYNISAA 244
Query: 1462 TTPFRLAYGQE 1472
TPF+ YG++
Sbjct: 245 MTPFKALYGRD 277
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.325 0.139 0.420
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 48,482,152
Number of Sequences: 36976
Number of extensions: 679205
Number of successful extensions: 4086
Number of sequences better than 10.0: 44
Number of HSP's better than 10.0 without gapping: 3025
Number of HSP's successfully gapped in prelim test: 137
Number of HSP's that attempted gapping in prelim test: 993
Number of HSP's gapped (non-prelim): 3263
length of query: 1610
length of database: 9,014,727
effective HSP length: 109
effective length of query: 1501
effective length of database: 4,984,343
effective search space: 7481498843
effective search space used: 7481498843
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 65 (29.6 bits)
Lotus: description of TM0041.7


