

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0037a.10
(168 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
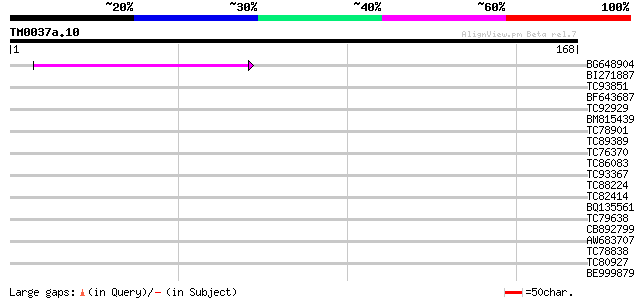
Score E
Sequences producing significant alignments: (bits) Value
BG648904 43 5e-05
BI271887 37 0.005
TC93851 weakly similar to GP|21734794|gb|AAM76972.1 reduced vern... 34 0.024
BF643687 31 0.27
TC92929 similar to GP|18252953|gb|AAL62403.1 transcription facto... 30 0.60
BM815439 29 0.78
TC78901 similar to GP|21734794|gb|AAM76972.1 reduced vernalizati... 29 0.78
TC89389 weakly similar to PIR|T45611|T45611 N-hydroxycinnamoyl/b... 28 1.7
TC76370 similar to GP|21734794|gb|AAM76972.1 reduced vernalizati... 28 1.7
TC86083 homologue to GP|13873334|dbj|BAB44155. hydroxypyruvate r... 28 2.3
TC93367 similar to GP|17381261|gb|AAL36049.1 At1g61590/T25B24_6 ... 27 3.9
TC88224 similar to GP|11994771|dbj|BAB03161. myosin-like protein... 27 3.9
TC82414 similar to PIR|T49915|T49915 pre-mRNA splicing factor AT... 24 4.5
BQ135561 27 5.1
TC79638 similar to PIR|T10242|T10242 (S)-2-hydroxy-acid oxidase ... 27 5.1
CB892799 weakly similar to PIR|T05768|T05 subtilisin-like protei... 27 5.1
AW683707 similar to GP|1814424|gb|A homeodomain protein AHDP {Ar... 26 6.6
TC78838 homologue to SP|O04136|HKL3_MALDO Homeobox protein knott... 26 6.6
TC80927 similar to PIR|B86411|B86411 protein F3M18.4 [imported] ... 26 6.6
BE999879 homologue to GP|17064906|gb Unknown protein {Arabidopsi... 26 8.6
>BG648904
Length = 770
Score = 43.1 bits (100), Expect = 5e-05
Identities = 20/65 (30%), Positives = 31/65 (46%)
Frame = +2
Query: 8 FAVEFAHDLGLQWKLVDPFGSVHLVSYESNSEEPYLFEGLLEIRKFYSLQGMNWFLLKYQ 67
F ++ ++L QW L+D G+ H V + + P L +G +R FY G +Y
Sbjct: 572 FVAQYFYELPEQWHLLDGDGNTHSVMFNKSVSRPLLTDGWSALRLFYEFSGSKIIAFQYL 751
Query: 68 GCSNF 72
G S F
Sbjct: 752 GGSRF 766
>BI271887
Length = 623
Score = 36.6 bits (83), Expect = 0.005
Identities = 21/69 (30%), Positives = 33/69 (47%)
Frame = +1
Query: 6 RKFAVEFAHDLGLQWKLVDPFGSVHLVSYESNSEEPYLFEGLLEIRKFYSLQGMNWFLLK 65
R F +F +G L DP G+ V +S + + +L +G IR FY + W L
Sbjct: 283 RMFGSDFGDQIGRFATLTDPKGNQFEVLVDSINGDFFLTKGWKAIRDFYGISLGAWITLI 462
Query: 66 YQGCSNFDL 74
+ G +FD+
Sbjct: 463 FVGVGHFDM 489
>TC93851 weakly similar to GP|21734794|gb|AAM76972.1 reduced vernalization
response 1 {Arabidopsis thaliana}, partial (9%)
Length = 710
Score = 34.3 bits (77), Expect = 0.024
Identities = 17/56 (30%), Positives = 31/56 (55%)
Frame = +1
Query: 49 EIRKFYSLQGMNWFLLKYQGCSNFDLFIYNDSIEEVCYLTRSLPPSITPTTHSHTS 104
+ ++YSL+ + +Y+G SNF + I++ + E+CY P TP+T T+
Sbjct: 415 QFAEYYSLRYGCFLSFRYEGNSNFSVIIFDATSVEICY------PLKTPSTSGETN 564
>BF643687
Length = 576
Score = 30.8 bits (68), Expect = 0.27
Identities = 20/50 (40%), Positives = 27/50 (54%), Gaps = 4/50 (8%)
Frame = +1
Query: 12 FAHDLGLQWK----LVDPFGSVHLVSYESNSEEPYLFEGLLEIRKFYSLQ 57
F+HD G Q + LVDP + V E N++ YL +G IR FY +Q
Sbjct: 379 FSHDFGDQIQQYATLVDPKRNQFEVLVERNNQGIYLTKGWHAIRDFYKVQ 528
>TC92929 similar to GP|18252953|gb|AAL62403.1 transcription factor putative
{Arabidopsis thaliana}, partial (58%)
Length = 598
Score = 29.6 bits (65), Expect = 0.60
Identities = 12/25 (48%), Positives = 16/25 (64%)
Frame = +2
Query: 92 PPSITPTTHSHTSFHGECTTGSKSV 116
PPS+ P T+SHTS H T S ++
Sbjct: 221 PPSLIPPTNSHTSKHSISTINSSNI 295
>BM815439
Length = 705
Score = 29.3 bits (64), Expect = 0.78
Identities = 19/70 (27%), Positives = 28/70 (39%)
Frame = -2
Query: 62 FLLKYQGCSNFDLFIYNDSIEEVCYLTRSLPPSITPTTHSHTSFHGECTTGSKSVTEEWI 121
FLL + C + I ++E +C+L S+ + SF +T S V
Sbjct: 611 FLLFFLACFDLQGLIAASTLESLCFLESLTSSSLESLCSTTPSFFSSKSTSSSGVCLPTR 432
Query: 122 FYPCFFTTRL 131
F FF RL
Sbjct: 431 FLSLFFLFRL 402
>TC78901 similar to GP|21734794|gb|AAM76972.1 reduced vernalization response
1 {Arabidopsis thaliana}, partial (37%)
Length = 1760
Score = 29.3 bits (64), Expect = 0.78
Identities = 20/79 (25%), Positives = 36/79 (45%)
Frame = +2
Query: 8 FAVEFAHDLGLQWKLVDPFGSVHLVSYESNSEEPYLFEGLLEIRKFYSLQGMNWFLLKYQ 67
F ++ D+ L P GSV V + + + +G E + YS+ + KY+
Sbjct: 281 FMRKYGGDISPTVTLTVPDGSVWRVIMKKVDNKFWFLDGWNEFVQNYSISTGYLLVFKYE 460
Query: 68 GCSNFDLFIYNDSIEEVCY 86
G S+F + I++ E+ Y
Sbjct: 461 GKSHFTVNIFSLPTSEINY 517
>TC89389 weakly similar to PIR|T45611|T45611
N-hydroxycinnamoyl/benzoyltransferase-like protein -
Arabidopsis thaliana, partial (20%)
Length = 914
Score = 28.1 bits (61), Expect = 1.7
Identities = 17/50 (34%), Positives = 23/50 (46%), Gaps = 2/50 (4%)
Frame = +2
Query: 93 PSITPTTHSHTSFHGECTTGSKSVTEEW--IFYPCFFTTRLSPNQVSSSQ 140
P T TH S ++ + T E IFYPC F T P++ + SQ
Sbjct: 98 PVPTNCTHQKGSSLSPSSSAKPNSTSETLSIFYPCLFPTTCRPSRNNRSQ 247
>TC76370 similar to GP|21734794|gb|AAM76972.1 reduced vernalization response
1 {Arabidopsis thaliana}, partial (36%)
Length = 1840
Score = 28.1 bits (61), Expect = 1.7
Identities = 19/66 (28%), Positives = 35/66 (52%), Gaps = 1/66 (1%)
Frame = +1
Query: 22 LVDPFGSVHLVSYESNSEEPYLFEGLLEIRKFYSLQGMNWFLL-KYQGCSNFDLFIYNDS 80
L P G+V + + + +G + + Y++ G+ +FL+ Y+G SNF + I+N S
Sbjct: 247 LTVPDGTVWRLGLKKVDNRIWFVDGWQDFVQRYAI-GIGYFLVFTYEGNSNFIVHIFNMS 423
Query: 81 IEEVCY 86
E+ Y
Sbjct: 424 TAELNY 441
>TC86083 homologue to GP|13873334|dbj|BAB44155. hydroxypyruvate reductase
{Bruguiera gymnorrhiza}, complete
Length = 1418
Score = 27.7 bits (60), Expect = 2.3
Identities = 14/33 (42%), Positives = 17/33 (51%)
Frame = -3
Query: 90 SLPPSITPTTHSHTSFHGECTTGSKSVTEEWIF 122
+LPPS+ H HTSF G C S V + F
Sbjct: 1293 TLPPSV----HMHTSFEGSCRIESTPVCLHYSF 1207
>TC93367 similar to GP|17381261|gb|AAL36049.1 At1g61590/T25B24_6
{Arabidopsis thaliana}, partial (41%)
Length = 791
Score = 26.9 bits (58), Expect = 3.9
Identities = 13/24 (54%), Positives = 15/24 (62%), Gaps = 2/24 (8%)
Frame = -3
Query: 55 SLQGMNW--FLLKYQGCSNFDLFI 76
SL +NW FLL Y CS+F L I
Sbjct: 618 SLYALNWSLFLLVYTNCSSFHLII 547
>TC88224 similar to GP|11994771|dbj|BAB03161. myosin-like protein
{Arabidopsis thaliana}, partial (19%)
Length = 1155
Score = 26.9 bits (58), Expect = 3.9
Identities = 17/51 (33%), Positives = 23/51 (44%), Gaps = 1/51 (1%)
Frame = -2
Query: 62 FLLKYQGCSNFDLFIYNDSIEEVC-YLTRSLPPSITPTTHSHTSFHGECTT 111
F + Q C N F N + + YLT L PT ++ TS H C+T
Sbjct: 716 FSVWIQTCFNLTRFYLNQELGIIAKYLTPLLKFFSQPTYNAQTSTHRSCST 564
>TC82414 similar to PIR|T49915|T49915 pre-mRNA splicing factor ATP-dependent
RNA helicase-like protein - Arabidopsis thaliana,
partial (13%)
Length = 740
Score = 24.3 bits (51), Expect(2) = 4.5
Identities = 10/29 (34%), Positives = 16/29 (54%)
Frame = -2
Query: 136 VSSSQLVMNQFCYVLMTNESVYGIFVSLN 164
+SSS V ++FC+ L ++ V F N
Sbjct: 358 ISSSMAVFSRFCFFLCSSSDVSDSFTEKN 272
Score = 20.8 bits (42), Expect(2) = 4.5
Identities = 9/26 (34%), Positives = 13/26 (49%)
Frame = -3
Query: 114 KSVTEEWIFYPCFFTTRLSPNQVSSS 139
KS + W F+P LSP ++ S
Sbjct: 447 KSAADSWPFFPSPSVYFLSPTPLAPS 370
>BQ135561
Length = 801
Score = 26.6 bits (57), Expect = 5.1
Identities = 16/49 (32%), Positives = 24/49 (48%)
Frame = -2
Query: 31 LVSYESNSEEPYLFEGLLEIRKFYSLQGMNWFLLKYQGCSNFDLFIYND 79
+ S +S+S Y+F+ + YS +N+F Q S F L IY D
Sbjct: 431 ITSIKSSSMNMYVFKKESPLENLYSNSIVNYFFTCLQVPSFFTLIIYID 285
>TC79638 similar to PIR|T10242|T10242 (S)-2-hydroxy-acid oxidase (EC 1.1.3.15)
- cucurbit, complete
Length = 1662
Score = 26.6 bits (57), Expect = 5.1
Identities = 13/32 (40%), Positives = 18/32 (55%)
Frame = -3
Query: 62 FLLKYQGCSNFDLFIYNDSIEEVCYLTRSLPP 93
FLLK+Q + F L Y + + C+L R L P
Sbjct: 1510 FLLKHQTRTKFILNSY*HPVHDACFLLRWLQP 1415
>CB892799 weakly similar to PIR|T05768|T05 subtilisin-like proteinase (EC
3.4.21.-) - Arabidopsis thaliana, partial (20%)
Length = 875
Score = 26.6 bits (57), Expect = 5.1
Identities = 15/44 (34%), Positives = 20/44 (45%), Gaps = 1/44 (2%)
Frame = +1
Query: 50 IRKFYSLQGMNWFLLKYQGCSNFDLFI-YNDSIEEVCYLTRSLP 92
+R F+ L NW Q CS+F + I Y + CY R P
Sbjct: 715 VRNFHGLSSCNWNCNLSQSCSSFMVSICYQVGHHDNCYNPR*AP 846
>AW683707 similar to GP|1814424|gb|A homeodomain protein AHDP {Arabidopsis
thaliana}, partial (3%)
Length = 620
Score = 26.2 bits (56), Expect = 6.6
Identities = 12/31 (38%), Positives = 22/31 (70%)
Frame = -1
Query: 137 SSSQLVMNQFCYVLMTNESVYGIFVSLNAEL 167
SSS L++ FC+++ + S++ IF+S+N L
Sbjct: 575 SSSLLLLKPFCFIIPMH*SIH-IFLSINISL 486
>TC78838 homologue to SP|O04136|HKL3_MALDO Homeobox protein knotted-1 like 3
(KNAP3). [Apple Malus sylvestris] {Malus domestica},
partial (65%)
Length = 2233
Score = 26.2 bits (56), Expect = 6.6
Identities = 16/43 (37%), Positives = 22/43 (50%)
Frame = -1
Query: 81 IEEVCYLTRSLPPSITPTTHSHTSFHGECTTGSKSVTEEWIFY 123
I+ +C+ R LPP P S S + T SKSVT +F+
Sbjct: 778 IQRMCHNLR-LPPPHLPVHQSSISNNRHLTLSSKSVTITVVFH 653
>TC80927 similar to PIR|B86411|B86411 protein F3M18.4 [imported] -
Arabidopsis thaliana, partial (41%)
Length = 1299
Score = 26.2 bits (56), Expect = 6.6
Identities = 13/45 (28%), Positives = 22/45 (48%)
Frame = -2
Query: 93 PSITPTTHSHTSFHGECTTGSKSVTEEWIFYPCFFTTRLSPNQVS 137
PS T + S +FHG+ + S + +PC F R +++S
Sbjct: 242 PSRTERSKSSGAFHGDFAVAAPSHSGMPTSFPCAFALRAEKSKLS 108
>BE999879 homologue to GP|17064906|gb Unknown protein {Arabidopsis thaliana},
partial (20%)
Length = 659
Score = 25.8 bits (55), Expect = 8.6
Identities = 12/26 (46%), Positives = 15/26 (57%)
Frame = +3
Query: 122 FYPCFFTTRLSPNQVSSSQLVMNQFC 147
F+P F TT +SP +S S N FC
Sbjct: 162 FFPLFLTTLISPKPLSLSS--SNHFC 233
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.322 0.136 0.423
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 6,793,319
Number of Sequences: 36976
Number of extensions: 113901
Number of successful extensions: 673
Number of sequences better than 10.0: 44
Number of HSP's better than 10.0 without gapping: 668
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 673
length of query: 168
length of database: 9,014,727
effective HSP length: 89
effective length of query: 79
effective length of database: 5,723,863
effective search space: 452185177
effective search space used: 452185177
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 55 (25.8 bits)
Lotus: description of TM0037a.10


