

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0036c.6
(70 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
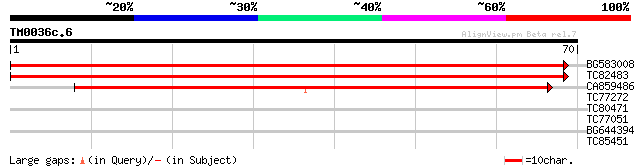
BG583008 similar to PIR|T52113|T521 probable transcription co-ac... 99 4e-22
TC82483 weakly similar to GP|13699910|dbj|BAB41214. putative tra... 77 2e-15
CA859486 weakly similar to PIR|T39903|T399 serine-rich protein -... 56 3e-09
TC77272 weakly similar to GP|19110917|gb|AAL85347.1 EDS1-like pr... 27 2.0
TC80471 weakly similar to GP|17529094|gb|AAL38757.1 unknown prot... 25 4.4
TC77051 similar to GP|16417950|gb|AAL18927.1 mevalonate disphosp... 25 4.4
BG644394 similar to PIR|S51324|S51 pullulanase - spinach, partia... 25 7.5
TC85451 similar to PIR|T06239|T06239 probable glutathione transf... 24 9.8
>BG583008 similar to PIR|T52113|T521 probable transcription co-activator KIWI
[imported] - Arabidopsis thaliana, partial (50%)
Length = 666
Score = 98.6 bits (244), Expect = 4e-22
Identities = 48/69 (69%), Positives = 53/69 (76%)
Frame = +3
Query: 1 DPDSIVVCEISKNRRVSVRNWQGRIVVDIREFYVKDGKQMPGKKGISLTMDQWNVLRNHI 60
D +V EI KNRRVSVR W + VDIREFY KDGKQ+PGKKGISL M+QW VLR+HI
Sbjct: 156 DDSGDIVFEIGKNRRVSVRTWNKQPWVDIREFYTKDGKQLPGKKGISLNMEQWIVLRDHI 335
Query: 61 EEIDKAVNE 69
EEID AV E
Sbjct: 336 EEIDNAVKE 362
>TC82483 weakly similar to GP|13699910|dbj|BAB41214. putative
transcriptional coactivator {Brassica rapa}, partial
(79%)
Length = 938
Score = 76.6 bits (187), Expect = 2e-15
Identities = 30/69 (43%), Positives = 50/69 (71%)
Frame = +2
Query: 1 DPDSIVVCEISKNRRVSVRNWQGRIVVDIREFYVKDGKQMPGKKGISLTMDQWNVLRNHI 60
D +V+CE+ K R+V++++++GR V IREFY KDGK++P KGISLT +QW + +
Sbjct: 320 DSGDLVICELGKKRKVTIQDFKGRTFVSIREFYTKDGKELPSSKGISLTQEQWLAFKKIV 499
Query: 61 EEIDKAVNE 69
+I++A+ +
Sbjct: 500 PDIEQAIQK 526
>CA859486 weakly similar to PIR|T39903|T399 serine-rich protein - fission
yeast (Schizosaccharomyces pombe), partial (3%)
Length = 606
Score = 55.8 bits (133), Expect = 3e-09
Identities = 25/60 (41%), Positives = 41/60 (67%), Gaps = 1/60 (1%)
Frame = +2
Query: 9 EISKNRRVSVRNWQGRIVVDIREFYVK-DGKQMPGKKGISLTMDQWNVLRNHIEEIDKAV 67
++S+ +RV+VR ++ I++D REF+ DG P KKGI+L +QWN L+ I +ID+ +
Sbjct: 263 KLSEKKRVTVRKFKNMILIDFREFFTNSDGDSNPTKKGIALQPEQWNKLKEFISDIDEEI 442
>TC77272 weakly similar to GP|19110917|gb|AAL85347.1 EDS1-like protein
{Nicotiana benthamiana}, partial (51%)
Length = 2168
Score = 26.6 bits (57), Expect = 2.0
Identities = 9/25 (36%), Positives = 17/25 (68%)
Frame = +1
Query: 39 QMPGKKGISLTMDQWNVLRNHIEEI 63
++ G+KGI + D WN+LR ++ +
Sbjct: 1669 KVEGQKGIGIHKDGWNILRRGMKAV 1743
>TC80471 weakly similar to GP|17529094|gb|AAL38757.1 unknown protein
{Arabidopsis thaliana}, partial (73%)
Length = 812
Score = 25.4 bits (54), Expect = 4.4
Identities = 10/26 (38%), Positives = 15/26 (57%)
Frame = +3
Query: 44 KGISLTMDQWNVLRNHIEEIDKAVNE 69
K I Q +LR H++ +D AVN+
Sbjct: 429 KSIDSLASQLKILRRHVDVLDSAVNK 506
>TC77051 similar to GP|16417950|gb|AAL18927.1 mevalonate disphosphate
decarboxylase {Hevea brasiliensis}, partial (98%)
Length = 1739
Score = 25.4 bits (54), Expect = 4.4
Identities = 13/38 (34%), Positives = 18/38 (47%)
Frame = +1
Query: 23 GRIVVDIREFYVKDGKQMPGKKGISLTMDQWNVLRNHI 60
GR +RE + KKGI +T + W+ L HI
Sbjct: 400 GRFQSCLREIRARACDVEDNKKGIKITKEDWSKLYVHI 513
>BG644394 similar to PIR|S51324|S51 pullulanase - spinach, partial (14%)
Length = 578
Score = 24.6 bits (52), Expect = 7.5
Identities = 13/29 (44%), Positives = 14/29 (47%), Gaps = 3/29 (10%)
Frame = +2
Query: 34 VKDGKQMPGKKGIS---LTMDQWNVLRNH 59
VKDG+ PGK IS MD W H
Sbjct: 230 VKDGQCYPGKSTIS*YWSIMDSWTYSNEH 316
>TC85451 similar to PIR|T06239|T06239 probable glutathione transferase (EC
2.5.1.18) 2 4-D inducible - soybean, complete
Length = 1155
Score = 24.3 bits (51), Expect = 9.8
Identities = 14/34 (41%), Positives = 20/34 (58%)
Frame = +3
Query: 26 VVDIREFYVKDGKQMPGKKGISLTMDQWNVLRNH 59
+V+IR+ K G + G +SL +D W V RNH
Sbjct: 774 IVEIRK---KFGLE*VGDVRLSLCLDVW*VWRNH 866
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.317 0.136 0.403
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 2,054,001
Number of Sequences: 36976
Number of extensions: 19859
Number of successful extensions: 86
Number of sequences better than 10.0: 16
Number of HSP's better than 10.0 without gapping: 86
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 86
length of query: 70
length of database: 9,014,727
effective HSP length: 46
effective length of query: 24
effective length of database: 7,313,831
effective search space: 175531944
effective search space used: 175531944
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 51 (24.3 bits)
Lotus: description of TM0036c.6


