

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0035.12
(125 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
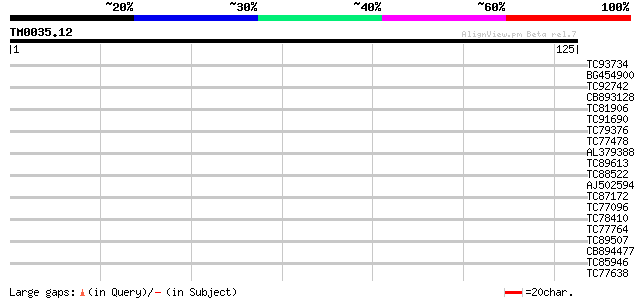
Score E
Sequences producing significant alignments: (bits) Value
TC93734 similar to PIR|T46027|T46027 hypothetical protein T10K17... 29 0.42
BG454900 weakly similar to PIR|T39903|T399 serine-rich protein -... 28 0.93
TC92742 similar to PIR|T00804|T00804 hypothetical protein At2g32... 27 2.1
CB893128 weakly similar to PIR|F96671|F966 hypothetical protein ... 27 2.1
TC81906 similar to GP|20259338|gb|AAM13994.1 unknown protein {Ar... 27 2.7
TC91690 similar to GP|15081684|gb|AAK82497.1 At2g24020/T29E15.22... 26 3.5
TC79376 homologue to GP|13877645|gb|AAK43900.1 Unknown protein {... 26 4.6
TC77478 similar to GP|9884651|dbj|BAB11979.1 MGDG synthase type ... 26 4.6
AL379388 26 4.6
TC89613 similar to GP|14334448|gb|AAK59422.1 unknown protein {Ar... 26 4.6
TC88522 similar to PIR|T02461|T02461 hypothetical protein At2g45... 26 4.6
AJ502594 weakly similar to PIR|T05755|T057 hypothetical protein ... 26 4.6
TC87172 similar to PIR|T51357|T51357 calcineurin B-like protein ... 25 6.0
TC77096 similar to PIR|T05572|T05572 glucose-6-phosphate isomera... 25 6.0
TC78410 similar to PIR|A84906|A84906 probable auxin-regulated pr... 25 6.0
TC77764 similar to GP|12324063|gb|AAG51991.1 unknown protein; 39... 25 6.0
TC89507 similar to GP|20197269|gb|AAC31240.2 expressed protein {... 25 7.9
CB894477 weakly similar to GP|10178181|db gene_id:MQN23.13~pir||... 25 7.9
TC85946 homologue to PIR|S52261|S52261 NADH dehydrogenase (ubiqu... 25 7.9
TC77638 similar to GP|20161455|dbj|BAB90379. ankyrin-like protei... 25 7.9
>TC93734 similar to PIR|T46027|T46027 hypothetical protein T10K17.260 -
Arabidopsis thaliana, partial (10%)
Length = 686
Score = 29.3 bits (64), Expect = 0.42
Identities = 23/75 (30%), Positives = 30/75 (39%)
Frame = +1
Query: 4 NPGQHHHSHRRSFFTHINWAPDQHPHEVLAFMPSLFKSNDLARRKRASDGDKSLNKTQEE 63
N +H SH H+N PH PSLF S +L+ R+ S
Sbjct: 1 NTHKHAVSHDTKSHLHLN-----RPHH-----PSLFLSLNLSPNSRSVIHSSSSIPIHIN 150
Query: 64 KASIWNKTQIKTPLN 78
SIWN+T + LN
Sbjct: 151SNSIWNETIFSSKLN 195
>BG454900 weakly similar to PIR|T39903|T399 serine-rich protein - fission
yeast (Schizosaccharomyces pombe), partial (12%)
Length = 258
Score = 28.1 bits (61), Expect = 0.93
Identities = 14/42 (33%), Positives = 17/42 (40%), Gaps = 8/42 (19%)
Frame = -1
Query: 3 HNPGQHHHSHRRSFFTHIN--------WAPDQHPHEVLAFMP 36
H+ QHHH H R HI + P HPH + P
Sbjct: 186 HHHQQHHHHHPRLLHKHIQLLQLPHHLYHPLLHPHSLQVLPP 61
>TC92742 similar to PIR|T00804|T00804 hypothetical protein At2g32640
[imported] - Arabidopsis thaliana, partial (28%)
Length = 757
Score = 26.9 bits (58), Expect = 2.1
Identities = 11/26 (42%), Positives = 18/26 (68%)
Frame = +2
Query: 26 QHPHEVLAFMPSLFKSNDLARRKRAS 51
Q P ++++ +PSLF S+DLA + S
Sbjct: 332 QEPQQIVSTIPSLFTSSDLADKIEGS 409
>CB893128 weakly similar to PIR|F96671|F966 hypothetical protein F13O11.14
[imported] - Arabidopsis thaliana, partial (8%)
Length = 768
Score = 26.9 bits (58), Expect = 2.1
Identities = 13/30 (43%), Positives = 19/30 (63%), Gaps = 1/30 (3%)
Frame = +3
Query: 4 NPGQHHHSHRRSFF-THINWAPDQHPHEVL 32
+P QHHHS SF T ++W+ + P E+L
Sbjct: 12 SPSQHHHSASTSFMATEVDWS--ELPKELL 95
>TC81906 similar to GP|20259338|gb|AAM13994.1 unknown protein {Arabidopsis
thaliana}, partial (26%)
Length = 653
Score = 26.6 bits (57), Expect = 2.7
Identities = 8/22 (36%), Positives = 11/22 (49%)
Frame = -3
Query: 8 HHHSHRRSFFTHINWAPDQHPH 29
HHH +S F H+ + H H
Sbjct: 219 HHHEQHKSLFLHVRFLDKYHHH 154
>TC91690 similar to GP|15081684|gb|AAK82497.1 At2g24020/T29E15.22
{Arabidopsis thaliana}, partial (19%)
Length = 500
Score = 26.2 bits (56), Expect = 3.5
Identities = 10/17 (58%), Positives = 11/17 (63%)
Frame = -2
Query: 16 FFTHINWAPDQHPHEVL 32
FF HINW P+ EVL
Sbjct: 442 FFYHINWIPNPSSREVL 392
>TC79376 homologue to GP|13877645|gb|AAK43900.1 Unknown protein {Arabidopsis
thaliana}, partial (29%)
Length = 952
Score = 25.8 bits (55), Expect = 4.6
Identities = 17/44 (38%), Positives = 20/44 (44%), Gaps = 3/44 (6%)
Frame = +3
Query: 45 ARRKRASDGDKSLNKTQEEKA---SIWNKTQIKTPLNKASTFEA 85
A RK + D+ L K QE SIWNK N+ FEA
Sbjct: 210 ASRKLQGEIDRVLKKVQEGVEVFDSIWNKVYDTDNANQKEKFEA 341
>TC77478 similar to GP|9884651|dbj|BAB11979.1 MGDG synthase type A {Glycine
max}, partial (85%)
Length = 2229
Score = 25.8 bits (55), Expect = 4.6
Identities = 8/12 (66%), Positives = 9/12 (74%)
Frame = -3
Query: 3 HNPGQHHHSHRR 14
H+P QHHH H R
Sbjct: 796 HSPHQHHHHHHR 761
>AL379388
Length = 214
Score = 25.8 bits (55), Expect = 4.6
Identities = 8/21 (38%), Positives = 11/21 (52%)
Frame = -3
Query: 7 QHHHSHRRSFFTHINWAPDQH 27
Q+HHS R + + W P H
Sbjct: 77 QYHHSQHRCYTEEVFWRPPSH 15
>TC89613 similar to GP|14334448|gb|AAK59422.1 unknown protein {Arabidopsis
thaliana}, partial (54%)
Length = 1706
Score = 25.8 bits (55), Expect = 4.6
Identities = 9/22 (40%), Positives = 11/22 (49%)
Frame = -3
Query: 8 HHHSHRRSFFTHINWAPDQHPH 29
HHH +R+ H P QH H
Sbjct: 606 HHHQQQRASTVHSTTQPPQHGH 541
>TC88522 similar to PIR|T02461|T02461 hypothetical protein At2g45860
[imported] - Arabidopsis thaliana, partial (98%)
Length = 600
Score = 25.8 bits (55), Expect = 4.6
Identities = 15/52 (28%), Positives = 26/52 (49%)
Frame = +1
Query: 40 KSNDLARRKRASDGDKSLNKTQEEKASIWNKTQIKTPLNKASTFEATKLRSG 91
K NDLA+RKR + + K ++EK + +++ NK + K + G
Sbjct: 19 KKNDLAKRKRQHEFELRREKLEKEK----KEKKLQAKKNKMKVDGSDKKKKG 162
>AJ502594 weakly similar to PIR|T05755|T057 hypothetical protein M4I22.120 -
Arabidopsis thaliana, partial (41%)
Length = 353
Score = 25.8 bits (55), Expect = 4.6
Identities = 8/20 (40%), Positives = 10/20 (50%)
Frame = -3
Query: 3 HNPGQHHHSHRRSFFTHINW 22
H+P HHH H + H W
Sbjct: 324 HHPQYHHHHHLDHYLHHCCW 265
>TC87172 similar to PIR|T51357|T51357 calcineurin B-like protein 2
[imported] - Arabidopsis thaliana, complete
Length = 1162
Score = 25.4 bits (54), Expect = 6.0
Identities = 9/12 (75%), Positives = 9/12 (75%)
Frame = +1
Query: 8 HHHSHRRSFFTH 19
HHHS SFFTH
Sbjct: 31 HHHSSSSSFFTH 66
>TC77096 similar to PIR|T05572|T05572 glucose-6-phosphate isomerase (EC
5.3.1.9) - Arabidopsis thaliana, partial (89%)
Length = 2494
Score = 25.4 bits (54), Expect = 6.0
Identities = 9/16 (56%), Positives = 11/16 (68%)
Frame = +1
Query: 9 HHSHRRSFFTHINWAP 24
HHSHR T I+W+P
Sbjct: 1648 HHSHRARSNTSISWSP 1695
>TC78410 similar to PIR|A84906|A84906 probable auxin-regulated protein
[imported] - Arabidopsis thaliana, partial (61%)
Length = 1073
Score = 25.4 bits (54), Expect = 6.0
Identities = 11/30 (36%), Positives = 14/30 (46%)
Frame = +1
Query: 1 MIHNPGQHHHSHRRSFFTHINWAPDQHPHE 30
+I +HH+H SF HI H HE
Sbjct: 526 LIMGISSNHHNHHLSFHLHIPHLNFHHHHE 615
>TC77764 similar to GP|12324063|gb|AAG51991.1 unknown protein; 3976-4980
{Arabidopsis thaliana}, partial (54%)
Length = 1777
Score = 25.4 bits (54), Expect = 6.0
Identities = 11/28 (39%), Positives = 12/28 (42%)
Frame = -2
Query: 2 IHNPGQHHHSHRRSFFTHINWAPDQHPH 29
+H HHH H H P QHPH
Sbjct: 1272 LHCNFHHHHHHHCRHPRHPL*HPSQHPH 1189
>TC89507 similar to GP|20197269|gb|AAC31240.2 expressed protein {Arabidopsis
thaliana}, partial (95%)
Length = 843
Score = 25.0 bits (53), Expect = 7.9
Identities = 12/38 (31%), Positives = 19/38 (49%)
Frame = -1
Query: 2 IHNPGQHHHSHRRSFFTHINWAPDQHPHEVLAFMPSLF 39
IH P Q SHR S+ T+I+ + + P++F
Sbjct: 765 IHRPDQQVSSHRISYVTYIDSGTKLFHGMIFSSYPTVF 652
>CB894477 weakly similar to GP|10178181|db
gene_id:MQN23.13~pir||T07721~strong similarity to
unknown protein {Arabidopsis thaliana}, partial (34%)
Length = 812
Score = 25.0 bits (53), Expect = 7.9
Identities = 10/21 (47%), Positives = 12/21 (56%), Gaps = 2/21 (9%)
Frame = -2
Query: 3 HNPGQHHHSHRRSF--FTHIN 21
H PG HH RR+ F H+N
Sbjct: 769 HCPGHHHRKQRRNHPKFNHLN 707
>TC85946 homologue to PIR|S52261|S52261 NADH dehydrogenase (ubiquinone) (EC
1.6.5.3) flavoprotein 1 precursor - potato, partial (96%)
Length = 1611
Score = 25.0 bits (53), Expect = 7.9
Identities = 7/10 (70%), Positives = 8/10 (80%)
Frame = -1
Query: 3 HNPGQHHHSH 12
HNP +HHH H
Sbjct: 1269 HNPSEHHHIH 1240
>TC77638 similar to GP|20161455|dbj|BAB90379. ankyrin-like protein {Oryza
sativa (japonica cultivar-group)}, partial (62%)
Length = 2345
Score = 25.0 bits (53), Expect = 7.9
Identities = 15/60 (25%), Positives = 29/60 (48%)
Frame = +1
Query: 40 KSNDLARRKRASDGDKSLNKTQEEKASIWNKTQIKTPLNKASTFEATKLRSGTCIFRSGA 99
KS+ K++SD +++ + EEK + ++ +T + K +S +F SGA
Sbjct: 781 KSDSDESEKKSSDSNETTDSNVEEKVEQSQNKESDENASEKNTDDNAKDQSSNEVFPSGA 960
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.317 0.129 0.382
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 4,120,109
Number of Sequences: 36976
Number of extensions: 54922
Number of successful extensions: 657
Number of sequences better than 10.0: 43
Number of HSP's better than 10.0 without gapping: 632
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 651
length of query: 125
length of database: 9,014,727
effective HSP length: 84
effective length of query: 41
effective length of database: 5,908,743
effective search space: 242258463
effective search space used: 242258463
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 52 (24.6 bits)
Lotus: description of TM0035.12


