

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
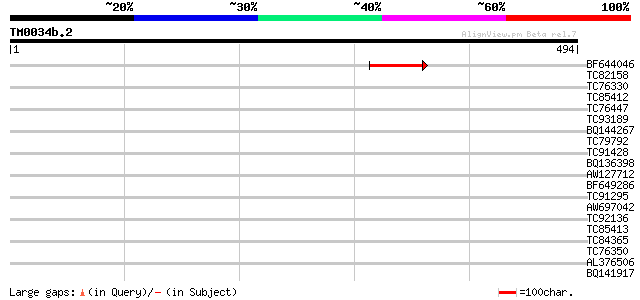
Query= TM0034b.2
(494 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
BF644046 45 5e-05
TC82158 weakly similar to GP|19920013|gb|AAM08453.1 hypothetical... 36 0.039
TC76330 similar to GP|8132441|gb|AAF73291.1| extensin {Pisum sat... 34 0.11
TC85412 homologue to GP|15450763|gb|AAK96653.1 elongation factor... 34 0.11
TC76447 homologue to GP|11121502|emb|CAC14888. putative extensin... 34 0.15
TC93189 similar to GP|18491187|gb|AAL69496.1 unknown protein {Ar... 34 0.15
BQ144267 similar to GP|13882294|gb PE_PGRS family protein {Mycob... 33 0.19
TC79792 similar to GP|14335158|gb|AAK59859.1 AT3g18420/MYF24_13 ... 33 0.33
TC91428 weakly similar to PIR|D71408|D71408 hypothetical protein... 32 0.43
BQ136398 weakly similar to GP|15145286|gb| Hypothetical protein ... 32 0.56
AW127712 similar to PIR|B71203|B712 hypothetical protein PH1895 ... 32 0.56
BF649286 32 0.73
TC91295 similar to PIR|T46192|T46192 acetylglutamate kinase-like... 32 0.73
AW697042 similar to GP|13122418|dbj putative glycin-rich protein... 32 0.73
TC92136 homologue to GP|22136812|gb|AAM91750.1 unknown protein {... 32 0.73
TC85413 similar to PIR|S23737|S23737 proline-rich protein precur... 32 0.73
TC84365 similar to GP|15811367|gb|AAL08940.1 zinc finger protein... 30 1.6
TC76350 similar to GP|8132441|gb|AAF73291.1| extensin {Pisum sat... 30 1.6
AL376506 homologue to GP|15021744|gb root nodule extensin {Pisum... 30 2.1
BQ141917 similar to GP|11993889|gb| virion-associated nuclear-sh... 30 2.1
>BF644046
Length = 597
Score = 45.4 bits (106), Expect = 5e-05
Identities = 20/51 (39%), Positives = 31/51 (60%)
Frame = +3
Query: 314 YEYMFSELGIRLPFSPFVQTVLRDINAAPCQLHPNAWAFIRCFEILSAAVG 364
Y ++F ++G + PF+ F L+ +N A QLHPN AF+ FEI ++G
Sbjct: 15 YSFVFEDIGFKFPFTNFECDFLKALNVASSQLHPNCCAFMCGFEISCESLG 167
>TC82158 weakly similar to GP|19920013|gb|AAM08453.1 hypothetical protein
{Dictyostelium discoideum}, partial (4%)
Length = 1230
Score = 35.8 bits (81), Expect = 0.039
Identities = 22/76 (28%), Positives = 35/76 (45%)
Frame = +2
Query: 55 PHVAAPRDASVTSISLFLRNRHFPFHNRPIVRSNSPLHATHLPKTIMTSNLPHASNTRHP 114
P +P + ++ S P H+RPI S P TH T T++ PHA+ T
Sbjct: 68 PTTTSPATPAHSTSSTASTPSETPTHSRPITAST*PATPTH--STSTTTSTPHATPTNST 241
Query: 115 SSFSPSPIKTLLPHSH 130
S+ + +P+ T +H
Sbjct: 242 STTTSNPLATPTHSTH 289
>TC76330 similar to GP|8132441|gb|AAF73291.1| extensin {Pisum sativum},
partial (72%)
Length = 1082
Score = 34.3 bits (77), Expect = 0.11
Identities = 38/130 (29%), Positives = 57/130 (43%), Gaps = 5/130 (3%)
Frame = +1
Query: 66 TSISLFLRNRHFPFHNRPIVRSNSPLHAT--HLPKTIMTSNLPHASNTRHPSSFSPSPIK 123
TSIS L H + N PLH + HL + S+LPH S T H F+
Sbjct: 223 TSISHLLHLSTHHLHQSTSI--NHPLHLSTHHLHQFTNISHLPHLS-THHHHQFTNI--- 384
Query: 124 TLLPHSHPFLRPLFT--SLIAYQTPSAHFRSGAVAYHP-FRYPLFTSYSGESHCPYLCIT 180
+ LPH LFT SL+ + + H + +++ P Y+ SH P+L
Sbjct: 385 SHLPHLSTHHLHLFTSISLLLHLSTHHHHQFTNISHLPHLSTHHLHQYTSISHLPHLSTH 564
Query: 181 HILIFSFLSL 190
H+ ++ +SL
Sbjct: 565 HLHQYTSISL 594
>TC85412 homologue to GP|15450763|gb|AAK96653.1 elongation factor EF-2
{Arabidopsis thaliana}, complete
Length = 3229
Score = 34.3 bits (77), Expect = 0.11
Identities = 19/63 (30%), Positives = 30/63 (47%)
Frame = +1
Query: 101 MTSNLPHASNTRHPSSFSPSPIKTLLPHSHPFLRPLFTSLIAYQTPSAHFRSGAVAYHPF 160
+ NLPH + HP +P+P+ T + +HP L P + + P H + A HP
Sbjct: 2896 LDGNLPHGHHHHHPHPPAPTPVHTPVAPAHPPLHPPVHTPVVPTHPPLHPPAPA---HPP 3066
Query: 161 RYP 163
+P
Sbjct: 3067 LHP 3075
>TC76447 homologue to GP|11121502|emb|CAC14888. putative extensin {Nicotiana
sylvestris}, partial (58%)
Length = 564
Score = 33.9 bits (76), Expect = 0.15
Identities = 46/167 (27%), Positives = 63/167 (37%), Gaps = 28/167 (16%)
Frame = +3
Query: 53 YPPHVAAPRDASVTSISLFLR-NRHFPFHNRPIVRSNSPLHATHLPKTIMTSNLPHA--- 108
+P H+ TSIS L + H H+ ++ PLH + TS PH
Sbjct: 39 HPLHLFTHHLHLFTSISHHLHLSTH---HHHQFTSTSHPLHLSTHHLHQFTSTSPHLHPS 209
Query: 109 ----------SNTRHPSSFSPSPIKTLLPHSHPFLRPL--FTSLIAYQTPSAH----FRS 152
S HPS+ + PH HP L FTS + PS H F S
Sbjct: 210 THHHHQFTSTSPLLHPSTHHLHQFTSTSPHLHPSTHHLHQFTSTSPHLHPSTHHHHQFTS 389
Query: 153 GAVAYHPFRYPL--FTSYS-----GESHCP-YLCITHILIFSFLSLH 191
+ HP + L FT+ S H P ++ +H L+ S LH
Sbjct: 390 TSHLLHPSTHHLHPFTNTSPHLHPSTHHLPRFISTSHHLLPSIYHLH 530
>TC93189 similar to GP|18491187|gb|AAL69496.1 unknown protein {Arabidopsis
thaliana}, partial (35%)
Length = 642
Score = 33.9 bits (76), Expect = 0.15
Identities = 32/106 (30%), Positives = 46/106 (43%), Gaps = 20/106 (18%)
Frame = -3
Query: 84 IVRSNSP-LHATHLPKTIMTSNLPHASNTR--------------HPSSFSPSPIKTLLPH 128
+V +++P L +TH+P+ PH ++R HP SFS I TL PH
Sbjct: 304 LVTTHNPWLSSTHMPQNCRMFLWPHVISSRRSLWSSNS*AFPFSHPFSFS---IPTLFPH 134
Query: 129 SHPFLRP---LFTSL--IAYQTPSAHFRSGAVAYHPFRYPLFTSYS 169
PF+R LF L I+ Q P + + +R P YS
Sbjct: 133 VDPFIRSRSNLFCKLNIISIQQPFLNL----ITTRLYRLPFIDRYS 8
>BQ144267 similar to GP|13882294|gb PE_PGRS family protein {Mycobacterium
tuberculosis CDC1551}, partial (1%)
Length = 1188
Score = 33.5 bits (75), Expect = 0.19
Identities = 20/46 (43%), Positives = 25/46 (53%), Gaps = 1/46 (2%)
Frame = -1
Query: 46 FKTLLNLYPPHVAAPRDASVT-SISLFLRNRHFPFHNRPIVRSNSP 90
+KT L L PP A P +S T S SL + +R FP +RP R P
Sbjct: 459 WKTSLRLPPPGFARPASSSFTISPSLMIIHRTFPMPHRPRTRRGGP 322
>TC79792 similar to GP|14335158|gb|AAK59859.1 AT3g18420/MYF24_13
{Arabidopsis thaliana}, partial (64%)
Length = 1289
Score = 32.7 bits (73), Expect = 0.33
Identities = 22/72 (30%), Positives = 35/72 (48%), Gaps = 5/72 (6%)
Frame = +2
Query: 64 SVTSISLFLRN----RHFPFHNRPIVRSNSPLHATHLPK-TIMTSNLPHASNTRHPSSFS 118
S+ I++F N H P H+ P + S H H P+ T+ + + + +HP S S
Sbjct: 5 SLHYIAIFFFNIVLSTHIPIHSNPKSKKKSFHHDFHYPQFTLFLHPIL*SHHHQHPFSIS 184
Query: 119 PSPIKTLLPHSH 130
PSP+ P +H
Sbjct: 185 PSPLSP--PSNH 214
>TC91428 weakly similar to PIR|D71408|D71408 hypothetical protein dl3335w -
Arabidopsis thaliana, partial (13%)
Length = 694
Score = 32.3 bits (72), Expect = 0.43
Identities = 21/76 (27%), Positives = 38/76 (49%), Gaps = 10/76 (13%)
Frame = +3
Query: 81 NRPIVRSNSPLHATHLPKTIMTSNLP----HASNTR------HPSSFSPSPIKTLLPHSH 130
NR + ++ P+ +T L T++ ++ H++ R HP+ F +P +TL H+H
Sbjct: 198 NRNLSQTIPPISSTFLKTTLLLNHSKSRRIHSTQPRYPSRHSHPTPFLQNPCQTLTRHNH 377
Query: 131 PFLRPLFTSLIAYQTP 146
LR L + Y+ P
Sbjct: 378 RRLRHLPQPPLQYRPP 425
>BQ136398 weakly similar to GP|15145286|gb| Hypothetical protein C27H5.3
{Caenorhabditis elegans}, partial (10%)
Length = 856
Score = 32.0 bits (71), Expect = 0.56
Identities = 25/88 (28%), Positives = 37/88 (41%), Gaps = 5/88 (5%)
Frame = +3
Query: 53 YPPHVAAPRDASVTSISLFLRNRHFPFHNRPIVRSNSPLHATHLPKTIMTSNL---PHAS 109
+PPH+ P S++ L ++ HFP P + N + I S L PH
Sbjct: 309 HPPHLLFPYPFSLSQFYLRNKSTHFP----PFLS*NKDKSSPSFFIPISLSPLGPNPHFL 476
Query: 110 NT--RHPSSFSPSPIKTLLPHSHPFLRP 135
+ P +F+P P+ L P PF P
Sbjct: 477 SLLFSRPLTFTPHPVPPLPPFPPPFTHP 560
>AW127712 similar to PIR|B71203|B712 hypothetical protein PH1895 - Pyrococcus
horikoshii, partial (10%)
Length = 309
Score = 32.0 bits (71), Expect = 0.56
Identities = 23/70 (32%), Positives = 30/70 (42%), Gaps = 2/70 (2%)
Frame = -1
Query: 54 PPHVAAPRDASVTSISLFLRNRHFPFHNRPIVRSNSPLHATHLPKTIMTSNLPH--ASNT 111
P H + PR S T SL P + PI SP+ +H P+T + + H S T
Sbjct: 291 PSHSSNPRQISQTPPSLTNPPPSIPPNTSPIYPPYSPISLSHSPRTRLFAQFSHPFLSLT 112
Query: 112 RHPSSFSPSP 121
P S S P
Sbjct: 111 PQPHSQSSYP 82
>BF649286
Length = 562
Score = 31.6 bits (70), Expect = 0.73
Identities = 21/63 (33%), Positives = 27/63 (42%)
Frame = -1
Query: 72 LRNRHFPFHNRPIVRSNSPLHATHLPKTIMTSNLPHASNTRHPSSFSPSPIKTLLPHSHP 131
L RH P HN P P + P+T + L HAS T H S K L HS+
Sbjct: 175 LHPRHLPHHNHP--TPTPPATPLNAPQT*LYRPLNHASGTHHQRHSS----KASLNHSNT 14
Query: 132 FLR 134
+ +
Sbjct: 13 YYK 5
>TC91295 similar to PIR|T46192|T46192 acetylglutamate kinase-like protein -
Arabidopsis thaliana, partial (47%)
Length = 720
Score = 31.6 bits (70), Expect = 0.73
Identities = 20/72 (27%), Positives = 32/72 (43%), Gaps = 4/72 (5%)
Frame = +2
Query: 92 HATHLPKTIMTSNLPHASNTRHPSSFSPSPIKTLLPHSHPFLRPLFTS----LIAYQTPS 147
H++H PK+I SN ++ HP + SP++ P H + TS ++Y+
Sbjct: 95 HSSHQPKSISKSNAHKPTSLFHPMAVVVSPLQPRHPSQHLRVNSESTSSLNRFLSYKNSE 274
Query: 148 AHFRSGAVAYHP 159
A S A P
Sbjct: 275 AKLSSSNTAAPP 310
>AW697042 similar to GP|13122418|dbj putative glycin-rich protein {Oryza
sativa (japonica cultivar-group)}, partial (11%)
Length = 730
Score = 31.6 bits (70), Expect = 0.73
Identities = 21/62 (33%), Positives = 28/62 (44%), Gaps = 5/62 (8%)
Frame = -3
Query: 90 PLHATHLPKTIMTSNLPHASNTRHPSSFSPSPIKTLLPHSHP-----FLRPLFTSLIAYQ 144
P HA P + + LP T H + P P+ +L PH HP L P T+L+A
Sbjct: 728 PPHAAPPPPPLHLAPLPLP--TLHSPTLPPPPLTSLSPHPHPHLPLCLLTPPPTTLLAPS 555
Query: 145 TP 146
P
Sbjct: 554 AP 549
>TC92136 homologue to GP|22136812|gb|AAM91750.1 unknown protein {Arabidopsis
thaliana}, partial (2%)
Length = 707
Score = 31.6 bits (70), Expect = 0.73
Identities = 18/67 (26%), Positives = 31/67 (45%)
Frame = -3
Query: 80 HNRPIVRSNSPLHATHLPKTIMTSNLPHASNTRHPSSFSPSPIKTLLPHSHPFLRPLFTS 139
H+ ++ S S + + T+ S++PH + + P + PSPI+ S P PL T
Sbjct: 354 HSSLLLSSLSIHNTSPTNTTLPNSHIPHHHSIQSPPQYPPSPIQLSFSDSPPVSSPL*TP 175
Query: 140 LIAYQTP 146
+ P
Sbjct: 174 THQHAPP 154
>TC85413 similar to PIR|S23737|S23737 proline-rich protein precursor -
kidney bean, partial (56%)
Length = 1139
Score = 31.6 bits (70), Expect = 0.73
Identities = 29/126 (23%), Positives = 48/126 (38%), Gaps = 9/126 (7%)
Frame = +1
Query: 47 KTLLNLYPPHVA-APRDASVTSISLFLRNRHFPFHNRPIVRS--------NSPLHATHLP 97
+TL YPPH AP + + H H+ + ++PLH +
Sbjct: 118 ETLTPTYPPHTTPAPLHPPANAPH---HHHHHHIHSPTPAPTPTPSPSPIHTPLHPPYHS 288
Query: 98 KTIMTSNLPHASNTRHPSSFSPSPIKTLLPHSHPFLRPLFTSLIAYQTPSAHFRSGAVAY 157
+ H + HP +P+P+ T + +HP L P + + P H + A
Sbjct: 289 APVPAKPPTHGHHHHHPHPPAPTPVHTPVAPAHPPLHPPVHTPVVPTHPPLHPPAPA--- 459
Query: 158 HPFRYP 163
HP +P
Sbjct: 460 HPPLHP 477
>TC84365 similar to GP|15811367|gb|AAL08940.1 zinc finger protein
{Arabidopsis thaliana}, partial (6%)
Length = 597
Score = 30.4 bits (67), Expect = 1.6
Identities = 19/61 (31%), Positives = 31/61 (50%)
Frame = -2
Query: 122 IKTLLPHSHPFLRPLFTSLIAYQTPSAHFRSGAVAYHPFRYPLFTSYSGESHCPYLCITH 181
++++LP + +P SLI S H + +YH +YP + ES+C YL I H
Sbjct: 293 VQSILPQLDYY*QPRHHSLI--WNHSHHHGNSQNSYHQ-KYPTHYCWLPESYCHYLAICH 123
Query: 182 I 182
+
Sbjct: 122 L 120
>TC76350 similar to GP|8132441|gb|AAF73291.1| extensin {Pisum sativum},
partial (63%)
Length = 1008
Score = 30.4 bits (67), Expect = 1.6
Identities = 30/112 (26%), Positives = 38/112 (33%), Gaps = 15/112 (13%)
Frame = +3
Query: 53 YPPHVAAPRDASVTSISLFLRNRHFPFHNRPIVRSNSPLHATHLPKTIMTSNLPH----- 107
+P H++ TS S L F H +N PLH + TS H
Sbjct: 279 HPLHLSTHHLHQFTSTSPHLHP--FTHHLHQFTSTNHPLHLFTHHLHLFTSISHHLHLST 452
Query: 108 --------ASNTRHPSSFSPSPIKTLLPHSHPFLR--PLFTSLIAYQTPSAH 149
S+ HPS+ P PH HP P F S + PS H
Sbjct: 453 HHHHQFTSTSHLLHPSTHHLHPFTNTSPHLHPSTHHLPRFISTSHHLLPSIH 608
>AL376506 homologue to GP|15021744|gb root nodule extensin {Pisum sativum},
partial (68%)
Length = 470
Score = 30.0 bits (66), Expect = 2.1
Identities = 31/104 (29%), Positives = 40/104 (37%), Gaps = 8/104 (7%)
Frame = +2
Query: 54 PPHVAAPRDASV-TSISLFLRNRHFPFHN--RPIVRSNSP-LHATHLPKTIMTSNLP--H 107
PPH ++P V T + P H +P+ S P +H PK + S P H
Sbjct: 116 PPHYSSPPPPPVHTYPKPVYHSPPPPVHTYPKPVYHSPPPPVHKYPHPKPVYHSPPPPVH 295
Query: 108 ASNTRHPSSFSPSPIKTLLPHSHPFLR--PLFTSLIAYQTPSAH 149
HP SP P PH HP P +Y+ PS H
Sbjct: 296 KYPHPHPVYHSPPPPVHKYPHPHPVYHSPPPPPPKKSYKYPSPH 427
>BQ141917 similar to GP|11993889|gb| virion-associated nuclear-shuttling
protein {Mus musculus}, partial (4%)
Length = 1117
Score = 30.0 bits (66), Expect = 2.1
Identities = 28/90 (31%), Positives = 31/90 (34%)
Frame = +2
Query: 95 HLPKTIMTSNLPHASNTRHPSSFSPSPIKTLLPHSHPFLRPLFTSLIAYQTPSAHFRSGA 154
HL T+ N P S HP FSP+ I LP + RP Y TP
Sbjct: 575 HLRFTLHPINYPTQSPHYHPPPFSPTHIPQHLPITPNS*RPNTNPAHIYTTP-------- 730
Query: 155 VAYHPFRYPLFTSYSGESHCPYLCITHILI 184
HP S SHC H LI
Sbjct: 731 ---HPAVVRFLVSPP*LSHCVIYLRYHYLI 811
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.322 0.138 0.434
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 18,770,515
Number of Sequences: 36976
Number of extensions: 332060
Number of successful extensions: 2379
Number of sequences better than 10.0: 70
Number of HSP's better than 10.0 without gapping: 2330
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 2367
length of query: 494
length of database: 9,014,727
effective HSP length: 100
effective length of query: 394
effective length of database: 5,317,127
effective search space: 2094948038
effective search space used: 2094948038
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 60 (27.7 bits)
Lotus: description of TM0034b.2


