

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
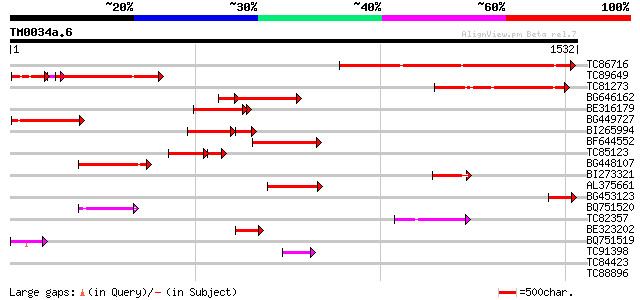
Query= TM0034a.6
(1532 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC86716 similar to PIR|T14279|T14279 myosin-like protein my5 - c... 506 e-143
TC89649 similar to PIR|T14276|T14276 myosin-like protein my2 - c... 320 e-113
TC81273 similar to PIR|T14278|T14278 myosin-like protein my4 - c... 342 9e-94
BG646162 similar to GP|15451591|g Putative myosin heavy chain {O... 286 6e-77
BE316179 similar to GP|15451591|g Putative myosin heavy chain {O... 273 3e-75
BG449727 similar to GP|7243765|gb unconventional myosin XI {Vall... 240 3e-63
BI265994 similar to PIR|F96587|F9 hypothetical protein T22H22.1 ... 175 3e-62
BF644552 similar to PIR|T14279|T1 myosin-like protein my5 - comm... 193 5e-49
TC85123 homologue to PIR|G96539|G96539 hypothetical protein F14I... 141 4e-42
BG448107 similar to GP|11994771|d myosin-like protein {Arabidops... 166 6e-41
BI273321 similar to GP|20503048|gb putative myosin heavy chain {... 148 1e-35
AL375661 weakly similar to PIR|S51824|S5 myosin heavy chain MYA2... 123 6e-28
BG453123 similar to PIR|D85390|D85 myosin-like protein [imported... 122 8e-28
BQ751520 weakly similar to SP|P70569|MY5 Myosin Vb (Myosin 5B) (... 119 7e-27
TC82357 similar to GP|7243765|gb|AAF43440.1| unconventional myos... 116 8e-26
BE323202 similar to PIR|S51823|S51 myosin heavy chain ATM2 - Ara... 85 2e-16
BQ751519 similar to GP|940860|emb|C MYO2 {Saccharomyces cerevisi... 66 9e-11
TC91398 similar to GP|9828627|gb|AAG00250.1| F1N21.13 {Arabidops... 46 1e-04
TC84423 similar to PIR|F84730|F84730 probable myosin heavy chain... 41 0.004
TC88896 weakly similar to GP|6728968|gb|AAF26966.1| unknown prot... 40 0.005
>TC86716 similar to PIR|T14279|T14279 myosin-like protein my5 - common
sunflower, partial (38%)
Length = 2675
Score = 506 bits (1302), Expect = e-143
Identities = 281/643 (43%), Positives = 405/643 (62%), Gaps = 7/643 (1%)
Frame = +2
Query: 892 KLEKQLDELTWRLHLEKKIRVSNEDAKQIEISKLQKMIEALNLELDAAKLATINECNKNA 951
KLEK+++ELTWRL +EK++R E+ K E++KL+ + A+ ++++ A I E +
Sbjct: 2 KLEKRVEELTWRLQIEKRLRTDLEEEKAQEVAKLRDALHAMQIQVEEANAKVIKEREASQ 181
Query: 952 VLQNQFELSIKEKSALKRELVAVDEIRKENAMLKVSLDAFEKKCTALEVELINAQKGRDE 1011
IKE + + ++ + E LK SL + + + E
Sbjct: 182 KAIQDAPPVIKETPVIIEDTEKINSLLAEVNCLKESLLLEREAKEEAKRAQAETEARSKE 361
Query: 1012 TIEKLREFEQKCSQLEQNVKSLEEKMLSLEDENHVLRQKALSAPLKSNRQGFAKSLSERY 1071
+K+ + ++K QL++ V+ LEEK+ + E EN VLRQ+AL+ AKSL+ R
Sbjct: 362 LFKKVEDSDRKADQLQELVQRLEEKISNSESENQVLRQQALAV------SPTAKSLAARP 523
Query: 1072 SNAVASRTERKPIF---ESPTPTKL---IPTFTPGMSDSRRSKLTAERHQDNCEFLSRCI 1125
+ + RT E+ TP+ + + S+ + K ++ Q+N + L +CI
Sbjct: 524 RSVIIQRTPENGNALNGEAKTPSDMTLALSNVREPESEGKPQKSLNDKQQENQDVLIKCI 703
Query: 1126 KENLGFKNGKPLAAPIIYKCLLHWHAFESERTAIFDYIIEGINEVLKARDDDDVLPYWLS 1185
++LGF GKP+AA +IYKCLLHW +FE ERT +FD II+ I ++A+D+ DVL YWLS
Sbjct: 704 SQDLGFSEGKPIAACVIYKCLLHWRSFEVERTTVFDRIIQTIASAVEAQDNTDVLAYWLS 883
Query: 1186 NTSALLCLLQRNLRSNGFLTTTGQRY-TGSAGLASRTVHVSYLVSTSINHSCGPKSPLKF 1244
NTS LL LLQR L+++G + T QR T S+ L R +S + S + P +
Sbjct: 884 NTSTLLMLLQRTLKASGAASLTPQRRRTASSSLFGR---MSQGLRGSPQSAGLPFINGRG 1054
Query: 1245 IGYDDGVSHVEARYPAILFKQQLTACVEKIFGLLRDDLKKELSPLLGLCIQAPKTGRQHG 1304
+ DG+ VEA+YPA+LFKQQLTA +EK++G++RD+LKKE+SPLLGLCIQAP+T RQ
Sbjct: 1055 LSRLDGLRQVEAKYPALLFKQQLTAFLEKLYGMIRDNLKKEISPLLGLCIQAPRTSRQGL 1234
Query: 1305 GKLSRSPSGLPQQPSGGQWANIVNFLDSLMSKLHGNHVPSFFIRKLVTQVFSFINITLFN 1364
K + + QQ W +IV L++ + + N+ P F +RK+ TQ+FSFI++ LFN
Sbjct: 1235 VKGRSHANAVAQQALIAHWQSIVKSLNNSLKIMKANYAPPFLVRKVFTQIFSFIDVQLFN 1414
Query: 1365 SLLLRRECCTFSNGEYMKSGLAELEKWIVNAKEEYAGTSWHELNYIRQAVGFLVIHQKRK 1424
SLLLRRECC+FSNGEY+K+GLAELE+W + A EEY G++W EL +IRQAVGFLVIHQK K
Sbjct: 1415 SLLLRRECCSFSNGEYVKTGLAELEQWCIEATEEYTGSAWEELKHIRQAVGFLVIHQKPK 1594
Query: 1425 KSLDEIRQDLCPALTVRQIYRISTMYWDDKYGTQSVSNEVVSEMREIVSSKDNQNITSNS 1484
KSL+EI ++LCP L+++Q+YRISTMYWDDKYGT SVS E + MR +V ++D+ N S S
Sbjct: 1595 KSLNEITKELCPGLSIQQLYRISTMYWDDKYGTHSVSTEGTTTMRAMV-AEDSTNAVSTS 1771
Query: 1485 FLLDDDMSIPFSAEDIDMAIPAIDPDEIDLPAFVPEYSCAQFL 1527
FLLDDD SIPFS +DI ++ ++ ++D P + E S FL
Sbjct: 1772 FLLDDDSSIPFSVDDISKSMQEVEVADVDPPPLIRENSGFGFL 1900
>TC89649 similar to PIR|T14276|T14276 myosin-like protein my2 - common
sunflower (fragment), partial (33%)
Length = 1356
Score = 320 bits (819), Expect(2) = e-113
Identities = 173/296 (58%), Positives = 210/296 (70%), Gaps = 4/296 (1%)
Frame = +3
Query: 124 LGELSPHVFAVADASYRAMMNEGKSQSILVSGESGAGKTETTKLIMQYLTFVGGRAAGDD 183
L ELSPHVFAVAD +YRAM+NEGKS SILVSGESGAGKTETTK++M+YL ++GGR+ +
Sbjct: 471 LEELSPHVFAVADVAYRAMINEGKSNSILVSGESGAGKTETTKMLMRYLAYLGGRSGVEG 650
Query: 184 RSVEQQVLESNPLLEAFGNARTVRNDNSSRFGKFVEIQFDSNGRISGAAIRTYLLERSRV 243
R+VEQQVLESNP+LEAFGNA+TVRN+NSSRFGKFVEIQFD+ GRISGAAIRTYLLERSRV
Sbjct: 651 RTVEQQVLESNPVLEAFGNAKTVRNNNSSRFGKFVEIQFDNKGRISGAAIRTYLLERSRV 830
Query: 244 VQITDPERNYHCFYQLCAFET-DAEKYELGHPSHFHYLNQSKIYELDGVSNVEEYVRTRR 302
Q++DPERNYHCFY LCA + EK++LG+PS FHYLNQSK Y LDGV +
Sbjct: 831 CQVSDPERNYHCFYLLCAAPAEEREKFKLGNPSSFHYLNQSKCYLLDGVR*C*RISCNSK 1010
Query: 303 AMNIVGISHEDQEAIFR---TLAAILHLGNIEFSPGKEYDSSVIKDEKSRFHMQMAANLF 359
+ F + L + N++ G+E DSSV+KDE SRFH+ A L
Sbjct: 1011SNGCCWNQ*GGARMQFLGS*QQSCTLEILNLQX--GEEIDSSVVKDENSRFHLNTTAELL 1184
Query: 360 MCDVDLLLSTLCTRSIQTREGSIVKALDCNAAVAGRDTLAKTVYARLFDWLVDKIN 415
CD + L L R + T + LD AA++ D AKT+Y+RLFDWLV+K N
Sbjct: 1185KCDANSLEDALIQRVMVTPXEVXTRTLDPVAALSSXDAXAKTMYSRLFDWLVEKXN 1352
Score = 109 bits (272), Expect(2) = e-113
Identities = 56/103 (54%), Positives = 74/103 (71%)
Frame = +1
Query: 5 MGSKVWVQDRDQAWVAAEVLASDGGGNRVQLVTDSGKKVLASPEKLCPRDADEEEHGGVE 64
+GS VWV+D AW+ EV +GG V + T GK V+ S K+ P+D +E GGV+
Sbjct: 121 VGSHVWVEDPALAWIGGEVTKINGG--EVHVRTGDGKTVVKSISKVFPKD-NEAPPGGVD 291
Query: 65 DMTRLAYLNEPGVLYNLKRRYTLNDIYTYTGSILIAVNPFTKL 107
DMT+L+YL+EPGVL NL RY LN+IYTYTG+ILIA+NPF+K+
Sbjct: 292 DMTKLSYLHEPGVLNNLATRYELNEIYTYTGNILIAINPFSKV 420
Score = 47.4 bits (111), Expect = 4e-05
Identities = 21/47 (44%), Positives = 25/47 (52%)
Frame = +2
Query: 103 PFTKLPHLYDNHMMEQYKGAPLGELSPHVFAVADASYRAMMNEGKSQ 149
PF +LPHLYD HMM+QYKGA G + F + GK Q
Sbjct: 407 PFQRLPHLYDTHMMQQYKGAAFGRVKSPCFCSCRCCIQGNDQRGKKQ 547
>TC81273 similar to PIR|T14278|T14278 myosin-like protein my4 - common
sunflower, partial (22%)
Length = 1078
Score = 342 bits (876), Expect = 9e-94
Identities = 178/364 (48%), Positives = 249/364 (67%)
Frame = +1
Query: 1149 WHAFESERTAIFDYIIEGINEVLKARDDDDVLPYWLSNTSALLCLLQRNLRSNGFLTTTG 1208
W +FE+ERT++FD +I+ I ++ +DD+D++ YWLSNTSALL LLQ++L+S G +T
Sbjct: 1 WKSFEAERTSVFDRLIQMIGSAIENQDDNDLMAYWLSNTSALLFLLQQSLKSGGSTDSTP 180
Query: 1209 QRYTGSAGLASRTVHVSYLVSTSINHSCGPKSPLKFIGYDDGVSHVEARYPAILFKQQLT 1268
R + + + + S S S +PL+ V V+A+YPA+LFKQQLT
Sbjct: 181 VRKPPNPTSLFGRMTMGFRSSPS---SANLPAPLEV------VRKVDAKYPALLFKQQLT 333
Query: 1269 ACVEKIFGLLRDDLKKELSPLLGLCIQAPKTGRQHGGKLSRSPSGLPQQPSGGQWANIVN 1328
A VEKI+G+LRD+LKKEL + LCIQAP+T + + RS + G W +I+
Sbjct: 334 AYVEKIYGILRDNLKKELQSWISLCIQAPRTSKG----VLRSGRSFSKDSPMGHWQSIIE 501
Query: 1329 FLDSLMSKLHGNHVPSFFIRKLVTQVFSFINITLFNSLLLRRECCTFSNGEYMKSGLAEL 1388
L++L+ + N VP I+K+ +Q FS+IN+ LFNSLLLRR+CCTFSNGEY+K+GLAEL
Sbjct: 502 SLNTLLCTMKENFVPPVLIQKIFSQTFSYINVQLFNSLLLRRDCCTFSNGEYVKAGLAEL 681
Query: 1389 EKWIVNAKEEYAGTSWHELNYIRQAVGFLVIHQKRKKSLDEIRQDLCPALTVRQIYRIST 1448
E W AKEEYAGTSW EL +IRQAVGFLVIHQK + S DEI DLCP ++V+Q+YR+ T
Sbjct: 682 ELWCCQAKEEYAGTSWDELKHIRQAVGFLVIHQKYRISYDEIINDLCPIMSVQQLYRVCT 861
Query: 1449 MYWDDKYGTQSVSNEVVSEMREIVSSKDNQNITSNSFLLDDDMSIPFSAEDIDMAIPAID 1508
+YWD Y T++VS +V+S M+ ++ ++D+ N S+SFLLDD SIPFS +D+ ++ D
Sbjct: 862 LYWDANYNTRTVSQDVLSSMK-VLMAEDSNNAQSDSFLLDDTSSIPFSVDDLSTSLQERD 1038
Query: 1509 PDEI 1512
E+
Sbjct: 1039 CSEM 1050
>BG646162 similar to GP|15451591|g Putative myosin heavy chain {Oryza sativa}
[Oryza sativa (japonica cultivar-group)], partial (12%)
Length = 716
Score = 286 bits (731), Expect = 6e-77
Identities = 146/181 (80%), Positives = 158/181 (86%)
Frame = +3
Query: 608 FKQQLQALMETLNSTEPHYVRCVKPNSLNRPQMFENGSVIHQLRCGGVLEAVRISLAGYP 667
FKQQLQALMETL +TEPHY+RCVKPNS N PQ FEN SV+HQLRCGGVLEAVRISLAGYP
Sbjct: 174 FKQQLQALMETLKTTEPHYIRCVKPNSSNLPQKFENTSVLHQLRCGGVLEAVRISLAGYP 353
Query: 668 TRRTYSEFVDRFGLIALEFMDGSYDDKAVAEKILQKLKLENFQLGRTKVFLRAGQIGILD 727
TRRTYSEFVDRFGLIA EFMDGSYDD+A +KILQKLKLENFQLGRTKVFLRAGQIGILD
Sbjct: 354 TRRTYSEFVDRFGLIAPEFMDGSYDDRATTQKILQKLKLENFQLGRTKVFLRAGQIGILD 533
Query: 728 SRRAEVLDNAARCIQRQLRTFIARRVFIAVRAAAVCLQACCRGYIGQKMYASKRETAAAI 787
SRR+EVLDNAA+ IQR+LRTFIA R F + ACCRG + +K+YASKRETAAAI
Sbjct: 534 SRRSEVLDNAAKFIQRRLRTFIAHRGFHLNPSCCCFS*ACCRGCLARKIYASKRETAAAI 713
Query: 788 S 788
S
Sbjct: 714 S 716
Score = 85.1 bits (209), Expect = 2e-16
Identities = 41/56 (73%), Positives = 45/56 (80%)
Frame = +1
Query: 564 RDYVVVEHCNLLSSSKCPFVSGLFPLPPEESSRSSYRFSSVASRFKQQLQALMETL 619
RDYVV+EHCN+LSSSKCPFVS LFP PEESSRSSY+FSSVASRFK + L
Sbjct: 1 RDYVVLEHCNVLSSSKCPFVSSLFPSLPEESSRSSYKFSSVASRFKVYISITFRVL 168
>BE316179 similar to GP|15451591|g Putative myosin heavy chain {Oryza sativa}
[Oryza sativa (japonica cultivar-group)], partial (9%)
Length = 478
Score = 273 bits (697), Expect(2) = 3e-75
Identities = 129/146 (88%), Positives = 137/146 (93%)
Frame = +2
Query: 496 KKPIGIVALLDEACMFPKSTHETFSTKLFQHFPSHPRLGKEKFSQTDFAISHYAGKVTYH 555
+ PIG++ALLDEACMFPKSTHETFSTKLFQ+F SHPRL E+FSQTDF ISHYAGKVTYH
Sbjct: 2 ENPIGVIALLDEACMFPKSTHETFSTKLFQNFRSHPRLASERFSQTDFIISHYAGKVTYH 181
Query: 556 TDTFLDKNRDYVVVEHCNLLSSSKCPFVSGLFPLPPEESSRSSYRFSSVASRFKQQLQAL 615
TD FLDKNRDYVVVEHCNLLSSS CPFVSGLFPL PEESSRSSY+FSSVA+RFKQQLQAL
Sbjct: 182 TDAFLDKNRDYVVVEHCNLLSSSNCPFVSGLFPLLPEESSRSSYKFSSVATRFKQQLQAL 361
Query: 616 METLNSTEPHYVRCVKPNSLNRPQMF 641
METL STEPHY+RCVKPNSLNRPQMF
Sbjct: 362 METLKSTEPHYIRCVKPNSLNRPQMF 439
Score = 29.6 bits (65), Expect(2) = 3e-75
Identities = 11/13 (84%), Positives = 12/13 (91%)
Frame = +3
Query: 641 FENGSVIHQLRCG 653
FEN S+IHQLRCG
Sbjct: 438 FENASIIHQLRCG 476
>BG449727 similar to GP|7243765|gb unconventional myosin XI {Vallisneria
gigantea}, partial (12%)
Length = 613
Score = 240 bits (613), Expect = 3e-63
Identities = 123/198 (62%), Positives = 155/198 (78%), Gaps = 2/198 (1%)
Frame = +3
Query: 5 MGSKVWVQDRDQAWVAAEVLASDGGGNRVQLVTDSGKKVLASPEKLCPRDADEEEHGGVE 64
+G++VWV+D D AW+ EVL + G ++++ SGK V + +D E GV+
Sbjct: 27 VGAQVWVEDSDVAWIDGEVL--EVNGEDIKVLCTSGKTVTVKASSVYHKDT-EAPPCGVD 197
Query: 65 DMTRLAYLNEPGVLYNLKRRYTLNDIYTYTGSILIAVNPFTKLPHLYDNHMMEQYKGAPL 124
DMT+LAYL+EPGVL NL+ RY +N+IYTYTG+ILIAVNPF KLPHLYD+HMM QYKGA
Sbjct: 198 DMTKLAYLHEPGVLDNLRSRYDINEIYTYTGNILIAVNPFIKLPHLYDSHMMAQYKGAGF 377
Query: 125 GELSPHVFAVADASYRAMMNEGKSQSILVSGESGAGKTETTKLIMQYLTFVGGRA--AGD 182
GELSPH FA+ADA+YR M+NEG SQSILVSGESGAGKTE+TKL+M+YL ++GGRA A +
Sbjct: 378 GELSPHPFAIADAAYRLMINEGISQSILVSGESGAGKTESTKLLMRYLAYMGGRAANAAE 557
Query: 183 DRSVEQQVLESNPLLEAF 200
R+VEQ+VLESNP+L AF
Sbjct: 558 XRTVEQKVLESNPVLXAF 611
>BI265994 similar to PIR|F96587|F9 hypothetical protein T22H22.1 [imported] -
Arabidopsis thaliana, partial (12%)
Length = 643
Score = 175 bits (444), Expect(2) = 3e-62
Identities = 86/130 (66%), Positives = 106/130 (81%)
Frame = +2
Query: 480 SYIEFVDNQDVLELIEKKPIGIVALLDEACMFPKSTHETFSTKLFQHFPSHPRLGKEKFS 539
SY+EFVDNQDVL+LIEKKP GI+ALLDEACMFPKSTHETFSTKL+Q F H R K K +
Sbjct: 2 SYLEFVDNQDVLDLIEKKPGGIIALLDEACMFPKSTHETFSTKLYQTFKDHKRFIKPKLA 181
Query: 540 QTDFAISHYAGKVTYHTDTFLDKNRDYVVVEHCNLLSSSKCPFVSGLFPLPPEESSRSSY 599
++DF + HYAG V Y ++ F+DKN+DYVV EH +LS+SKC FVSGLFP EE+++S+
Sbjct: 182 RSDFNVVHYAGVVQYQSELFIDKNKDYVVPEHQVMLSTSKCSFVSGLFPPLSEETAKSA- 358
Query: 600 RFSSVASRFK 609
+FSS+ SRFK
Sbjct: 359 KFSSIGSRFK 388
Score = 83.6 bits (205), Expect(2) = 3e-62
Identities = 39/57 (68%), Positives = 47/57 (82%)
Frame = +1
Query: 611 QLQALMETLNSTEPHYVRCVKPNSLNRPQMFENGSVIHQLRCGGVLEAVRISLAGYP 667
QL+ LME LN TEPHY+RCVKPN+L +P +FEN +V+ QLR GGVLEAVRI AG+P
Sbjct: 418 QLKHLMEALNLTEPHYIRCVKPNNLLKPGIFENMNVMQQLRSGGVLEAVRIKCAGFP 588
>BF644552 similar to PIR|T14279|T1 myosin-like protein my5 - common
sunflower, partial (13%)
Length = 601
Score = 193 bits (490), Expect = 5e-49
Identities = 98/188 (52%), Positives = 137/188 (72%)
Frame = +2
Query: 655 VLEAVRISLAGYPTRRTYSEFVDRFGLIALEFMDGSYDDKAVAEKILQKLKLENFQLGRT 714
VLEA+RIS AGYPTRRT+ EF++RFG++A E +DG+YDDK + IL K+ ++ +Q+G+T
Sbjct: 2 VLEAIRISCAGYPTRRTFYEFLNRFGVLAPEVLDGNYDDKVACQMILGKMGMKGYQIGKT 181
Query: 715 KVFLRAGQIGILDSRRAEVLDNAARCIQRQLRTFIARRVFIAVRAAAVCLQACCRGYIGQ 774
KVFLRAGQ+ LD+RR+EVL NAAR IQRQ RT IAR+ FI +R AA+ LQ+ RG + +
Sbjct: 182 KVFLRAGQMAELDARRSEVLGNAARIIQRQTRTHIARKEFIELRRAAISLQSNLRGILAR 361
Query: 775 KMYASKRETAAAISVQKYIRMWLRRKTYMKLCSSATIIQSCVRGFMTRQRFLHIKEHRAA 834
K+Y R AAA+ ++K + ++ RK+Y+K SSATI+Q+ +R R F K+ +AA
Sbjct: 362 KLYEKLRREAAALKIEKNFKGYIARKSYLKARSSATILQTGLRAMKARDEFXFRKQTKAA 541
Query: 835 TSIQACWR 842
IQA +R
Sbjct: 542 IRIQAHFR 565
>TC85123 homologue to PIR|G96539|G96539 hypothetical protein F14I3.6
[imported] - Arabidopsis thaliana, partial (14%)
Length = 544
Score = 141 bits (356), Expect(2) = 4e-42
Identities = 65/110 (59%), Positives = 83/110 (75%)
Frame = +3
Query: 428 IGVLDIYGFECFKDNSFEQFCINFANEKLQQHFNEHVFKMEQEEYGREEINWSYIEFVDN 487
I +LDIYGFE F NSFEQFCIN+ANE+LQQHFN H+FK+EQEEY ++ I+W+ +EF DN
Sbjct: 60 ISILDIYGFESFNRNSFEQFCINYANERLQQHFNRHLFKLEQEEYIQDGIDWAKVEFEDN 239
Query: 488 QDVLELIEKKPIGIVALLDEACMFPKSTHETFSTKLFQHFPSHPRLGKEK 537
QD L L EKKP+G+++LLDE FP T TF+ KL QH S+ +E+
Sbjct: 240 QDCLNLFEKKPLGLLSLLDEESTFPNGTDLTFANKLKQHLNSNSCFKEER 389
Score = 50.1 bits (118), Expect(2) = 4e-42
Identities = 24/52 (46%), Positives = 33/52 (63%)
Frame = +2
Query: 535 KEKFSQTDFAISHYAGKVTYHTDTFLDKNRDYVVVEHCNLLSSSKCPFVSGL 586
+ + S+ + HYAG+VTY T FL+KNRD + V+ LLSSSKC S +
Sbjct: 377 QRRTSKKPLLMRHYAGEVTYDTTAFLEKNRDLMHVDSIQLLSSSKCHLPSDI 532
>BG448107 similar to GP|11994771|d myosin-like protein {Arabidopsis
thaliana}, partial (17%)
Length = 599
Score = 166 bits (420), Expect = 6e-41
Identities = 91/200 (45%), Positives = 127/200 (63%), Gaps = 3/200 (1%)
Frame = +1
Query: 186 VEQQVLESNPLLEAFGNARTVRNDNSSRFGKFVEIQFDSNGRISGAAIRTYLLERSRVVQ 245
+E ++L++NP+LEAFGN +T+RNDNSSRFGK +EI F G+ISGA I+T+LLE+SRVVQ
Sbjct: 10 IEHEILKTNPILEAFGNGKTLRNDNSSRFGKLIEIHFSETGKISGANIQTFLLEKSRVVQ 189
Query: 246 ITDPERNYHCFYQLCAFETDA--EKYELGHPSHFHYLNQSKIYELDGVSNVEEYVRTRRA 303
+ ER+YH FYQLCA + EK L + YL QS Y ++ V + EE+ A
Sbjct: 190 CNEGERSYHIFYQLCAGAPSSLREKLNLRSVEDYKYLRQSNCYSINDVDDAEEFRIVTDA 369
Query: 304 MNIVGISHEDQEAIFRTLAAILHLGNIEFSP-GKEYDSSVIKDEKSRFHMQMAANLFMCD 362
+++V IS EDQE +F LAA+L LGNI F+ E ++DE + A L CD
Sbjct: 370 LDVVHISKEDQENVFAMLAAVLWLGNISFTVIDNENHVQAVEDE----GLFSTAKLIGCD 537
Query: 363 VDLLLSTLCTRSIQTREGSI 382
++ L TL TR ++ + +I
Sbjct: 538 IEDLKLTLSTRKMKVGKDTI 597
>BI273321 similar to GP|20503048|gb putative myosin heavy chain {Oryza sativa
(japonica cultivar-group)}, partial (3%)
Length = 294
Score = 148 bits (374), Expect = 1e-35
Identities = 76/106 (71%), Positives = 81/106 (75%)
Frame = +1
Query: 1143 YKCLLHWHAFESERTAIFDYIIEGINEVLKARDDDDVLPYWLSNTSALLCLLQRNLRSNG 1202
YKCLLHWHAFESERTAIFDYII+GINEV+K RDDD VLP LSNTSAL CLLQRN+RSNG
Sbjct: 16 YKCLLHWHAFESERTAIFDYIIDGINEVIKVRDDDIVLPXXLSNTSALXCLLQRNVRSNG 195
Query: 1203 FLTTTGQRYTGSAGLASRTVHVSYLVSTSINHSCGPKSPLKFIGYD 1248
FLTTT QRY GS+GL SR H G KSPLK IGY+
Sbjct: 196 FLTTTAQRYAGSSGLTSRIGH-------------GLKSPLKLIGYN 294
>AL375661 weakly similar to PIR|S51824|S5 myosin heavy chain MYA2 -
Arabidopsis thaliana, partial (10%)
Length = 486
Score = 123 bits (308), Expect = 6e-28
Identities = 62/147 (42%), Positives = 101/147 (68%)
Frame = +1
Query: 698 EKILQKLKLENFQLGRTKVFLRAGQIGILDSRRAEVLDNAARCIQRQLRTFIARRVFIAV 757
++IL+K+ L+ +Q+G+TKVFLRAGQ+ LD+ R+E+L +A IQR++R+++ARR F+++
Sbjct: 16 KRILEKVGLKGYQIGKTKVFLRAGQMAELDTCRSEILGKSASIIQRKVRSYLARRSFVSI 195
Query: 758 RAAAVCLQACCRGYIGQKMYASKRETAAAISVQKYIRMWLRRKTYMKLCSSATIIQSCVR 817
R +A+ LQA CRG + +++Y R+ A+++ +Q+ RM + RK Y +L SSA IQ+ +R
Sbjct: 196 RLSAIQLQAACRGQLARQVYEGLRQEASSLIIQRCFRMHIARKAYTELYSSAISIQTGMR 375
Query: 818 GFMTRQRFLHIKEHRAATSIQACWRMY 844
G R K+ AA IQ+ R Y
Sbjct: 376 GMAARCELRFRKQTSAAIIIQSHCRRY 456
>BG453123 similar to PIR|D85390|D85 myosin-like protein [imported] -
Arabidopsis thaliana, partial (5%)
Length = 642
Score = 122 bits (307), Expect = 8e-28
Identities = 63/77 (81%), Positives = 68/77 (87%)
Frame = +2
Query: 1455 YGTQSVSNEVVSEMREIVSSKDNQNITSNSFLLDDDMSIPFSAEDIDMAIPAIDPDEIDL 1514
YGTQSVSNEVVS MRE+V+ KDNQN+ SNSFLLDDD+SIPFSAED+DMAIP ID DEIDL
Sbjct: 2 YGTQSVSNEVVSVMREMVN-KDNQNMPSNSFLLDDDLSIPFSAEDVDMAIPPIDLDEIDL 178
Query: 1515 PAFVPEYSCAQFLNPIQ 1531
P FV EYSCAQFLN Q
Sbjct: 179 PLFVSEYSCAQFLNSHQ 229
>BQ751520 weakly similar to SP|P70569|MY5 Myosin Vb (Myosin 5B) (Myosin heavy
chain myr 6). [Rat] {Rattus norvegicus}, partial (7%)
Length = 549
Score = 119 bits (299), Expect = 7e-27
Identities = 70/167 (41%), Positives = 98/167 (57%), Gaps = 5/167 (2%)
Frame = -1
Query: 187 EQQVLESNPLLEAFGNARTVRND---NSSRFGKFVEIQFDSNGRISGAAIRTYLLERSRV 243
E+Q+L + P+++ R ++D +SSRFGK++EI FD I GA IRTYLLERSR+
Sbjct: 525 EEQILATTPIMKL----RKRKDDAQ*HSSRFGKYIEIMFDEKTNIIGAKIRTYLLERSRL 358
Query: 244 VQITDPERNYHCFYQLCAFETDAEKYELG--HPSHFHYLNQSKIYELDGVSNVEEYVRTR 301
V ERNYH FYQL A +D E+ EL F YLNQ +DGV + E+ T+
Sbjct: 357 VFQPLKERNYHIFYQLVAGASDKERQELSLLPIEEFEYLNQGNCPTIDGVDDKSEFEATK 178
Query: 302 RAMNIVGISHEDQEAIFRTLAAILHLGNIEFSPGKEYDSSVIKDEKS 348
++ +G+S Q IF+ LA +LHLGN++ + DS + E S
Sbjct: 177 GSLKTIGVSEAQQTEIFKLLAGLLHLGNVKIGASRN-DSVLAPTEPS 40
>TC82357 similar to GP|7243765|gb|AAF43440.1| unconventional myosin XI
{Vallisneria gigantea}, partial (8%)
Length = 618
Score = 116 bits (290), Expect = 8e-26
Identities = 68/206 (33%), Positives = 114/206 (55%), Gaps = 2/206 (0%)
Frame = +3
Query: 1040 LEDENHVLRQKALS--APLKSNRQGFAKSLSERYSNAVASRTERKPIFESPTPTKLIPTF 1097
+E NHVL++++LS +P+K+ + + +SE+ N E TP K
Sbjct: 6 IEFANHVLQKQSLSINSPVKTAVENLSTPVSEKLENGHHVAEEPYDADTYVTPVKQFVA- 182
Query: 1098 TPGMSDSRRSKLTAERHQDNCEFLSRCIKENLGFKNGKPLAAPIIYKCLLHWHAFESERT 1157
SD + + +E H + + L C+ +N+GF +GKP+AA IYKCLLHW +FE+ER+
Sbjct: 183 ---ESDVKLKRSCSELHHGSFDSLVNCVSKNIGFNHGKPIAAFTIYKCLLHWKSFEAERS 353
Query: 1158 AIFDYIIEGINEVLKARDDDDVLPYWLSNTSALLCLLQRNLRSNGFLTTTGQRYTGSAGL 1217
++FD +I+ I ++ +DD+ ++ YWLSNTSALL LL+++L++ T +
Sbjct: 354 SVFDRLIQMIGSAIEDQDDNALMAYWLSNTSALLFLLEQSLKTGTSTNATPNGKPPNPTS 533
Query: 1218 ASRTVHVSYLVSTSINHSCGPKSPLK 1243
+ S+L S S + P S ++
Sbjct: 534 LFGRMTKSFLSSPSSANLASPSSVVR 611
>BE323202 similar to PIR|S51823|S51 myosin heavy chain ATM2 - Arabidopsis
thaliana (fragment), partial (7%)
Length = 354
Score = 84.7 bits (208), Expect = 2e-16
Identities = 39/75 (52%), Positives = 53/75 (70%)
Frame = +2
Query: 611 QLQALMETLNSTEPHYVRCVKPNSLNRPQMFENGSVIHQLRCGGVLEAVRISLAGYPTRR 670
QL LM L ST PH++RC+KPN+ P +++N V+ QLRC G+LEAVRIS AGYPTR
Sbjct: 110 QLFKLMHQLESTTPHFIRCIKPNTKKLPGIYDNELVLQQLRCCGLLEAVRISRAGYPTRI 289
Query: 671 TYSEFVDRFGLIALE 685
+ +F R+G++ E
Sbjct: 290 KHQDFSRRYGILLSE 334
>BQ751519 similar to GP|940860|emb|C MYO2 {Saccharomyces cerevisiae}, partial
(7%)
Length = 794
Score = 66.2 bits (160), Expect = 9e-11
Identities = 37/111 (33%), Positives = 62/111 (55%), Gaps = 10/111 (9%)
Frame = +2
Query: 2 SFRMGSKVWVQDRDQAWVAAEVLASDGGGNRVQLV--TDSGKK--------VLASPEKLC 51
++ +G++ W D + WVA+EV+ GN+V+LV +SG++ L + +
Sbjct: 419 NYDVGTRAWQPDATEGWVASEVVNKTEEGNKVKLVFKLESGEEKTIEVTAAALQNGDSSL 598
Query: 52 PRDADEEEHGGVEDMTRLAYLNEPGVLYNLKRRYTLNDIYTYTGSILIAVN 102
P + +D+T L++LNEP VL ++ RY +IYTY+G +LIA N
Sbjct: 599 PPLMNPTMLEASDDLTNLSHLNEPAVLQAIRLRYAQKEIYTYSGIVLIAAN 751
>TC91398 similar to GP|9828627|gb|AAG00250.1| F1N21.13 {Arabidopsis
thaliana}, partial (17%)
Length = 880
Score = 45.8 bits (107), Expect = 1e-04
Identities = 30/91 (32%), Positives = 44/91 (47%), Gaps = 2/91 (2%)
Frame = +1
Query: 737 AARCIQRQLRTFIARRVFIAVRAAAVCLQACCRGY--IGQKMYASKRETAAAISVQKYIR 794
AA IQ R R+ A+ CL G IG + +S+ +AA+S+QK R
Sbjct: 319 AAARIQAAFRAHSFRKQMEREAASTTCLNGYVTGLGGIGGYVRSSRDYHSAALSIQKKYR 498
Query: 795 MWLRRKTYMKLCSSATIIQSCVRGFMTRQRF 825
W RK Y+ IQ+ VRG+ TR+++
Sbjct: 499 GWKVRKEYLAFRQKVVTIQAHVRGYQTRRQY 591
>TC84423 similar to PIR|F84730|F84730 probable myosin heavy chain [imported] -
Arabidopsis thaliana, partial (10%)
Length = 787
Score = 40.8 bits (94), Expect = 0.004
Identities = 43/184 (23%), Positives = 78/184 (42%), Gaps = 2/184 (1%)
Frame = +2
Query: 861 QCLWRRRQAKRELRRLKQEANETGALRLAKSKLEKQLDELTWRLHLEKKIRVSNEDAKQI 920
Q L + +++++ L+ NE+GA+ S+ +L+ H+E + E Q+
Sbjct: 176 QALSNNSELEQKVKSLEDLHNESGAVAATASQRSLELEG-----HIEATNAAAEEAKSQL 340
Query: 921 -EISKLQKMIEALNLELDAAKLATINECNKNAVLQNQFELSIKEKSALKRELVA-VDEIR 978
E+ E N+EL+ + N + N E + E S L A + E
Sbjct: 341 RELETRFIAAEQKNVELE-------QQLNLVQLKANDAERDVTEFSEKISHLDAKLKEAE 499
Query: 979 KENAMLKVSLDAFEKKCTALEVELINAQKGRDETIEKLREFEQKCSQLEQNVKSLEEKML 1038
+E +L L K + LE +L + + + E+L+ ++KCS+ E E+
Sbjct: 500 EEKNLLNSLLQEHMDKLSQLESDLNQSTQKNSQLEEELKIVKEKCSEHEDRATMNNERSR 679
Query: 1039 SLED 1042
LED
Sbjct: 680 ELED 691
Score = 32.0 bits (71), Expect = 1.9
Identities = 30/109 (27%), Positives = 49/109 (44%), Gaps = 12/109 (11%)
Frame = +2
Query: 962 KEKSALKRELVAVDEIRKENAMLKVSLDAFEKKCTALEVELINAQKGRDETIEKLREFEQ 1021
+E + L E ++E ++ L V+ F++ C+ LE +L + + +T L +
Sbjct: 17 EELTKLNAEKKGLEETVED---LTVNAKHFKELCSDLEEKLKLSDESFSKTDSLLSQALS 187
Query: 1022 KCSQLEQNVKSLEE------------KMLSLEDENHVLRQKALSAPLKS 1058
S+LEQ VKSLE+ SLE E H+ A + KS
Sbjct: 188 NNSELEQKVKSLEDLHNESGAVAATASQRSLELEGHIEATNAAAEEAKS 334
>TC88896 weakly similar to GP|6728968|gb|AAF26966.1| unknown protein
{Arabidopsis thaliana}, partial (27%)
Length = 1097
Score = 40.4 bits (93), Expect = 0.005
Identities = 42/198 (21%), Positives = 87/198 (43%), Gaps = 6/198 (3%)
Frame = +2
Query: 921 EISKLQKMIEALNLELDAAKLA------TINECNKNAVLQNQFELSIKEKSALKRELVAV 974
++++ + +IE LN+E +AAK+A ++EC K + E+ ++E + L+R
Sbjct: 68 KLAEKETLIEQLNVESEAAKMAESYARSVLDECRKKV---EELEMKVEEANQLERSA--- 229
Query: 975 DEIRKENAMLKVSLDAFEKKCTALEVELINAQKGRDETIEKLREFEQKCSQLEQNVKSLE 1034
+SL+ K+ L +A+ EKL E + +++ E
Sbjct: 230 ----------SLSLETATKQLEGKNELLHDAESEISSLKEKLGMLEMTVGRQRGDLEDAE 379
Query: 1035 EKMLSLEDENHVLRQKALSAPLKSNRQGFAKSLSERYSNAVASRTERKPIFESPTPTKLI 1094
+L+ ++EN + +K S L+S + +K ++ +N S + + + E KLI
Sbjct: 380 RCLLAAKEENIEMSKKIES--LESEIETVSKEKAQALNNEKLSASSVQTLLEE--KNKLI 547
Query: 1095 PTFTPGMSDSRRSKLTAE 1112
+ ++KL +
Sbjct: 548 NELEICRDEEEKTKLAMD 601
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.320 0.135 0.391
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 44,718,476
Number of Sequences: 36976
Number of extensions: 622627
Number of successful extensions: 3226
Number of sequences better than 10.0: 68
Number of HSP's better than 10.0 without gapping: 3162
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 3204
length of query: 1532
length of database: 9,014,727
effective HSP length: 109
effective length of query: 1423
effective length of database: 4,984,343
effective search space: 7092720089
effective search space used: 7092720089
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 65 (29.6 bits)
Lotus: description of TM0034a.6


