

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
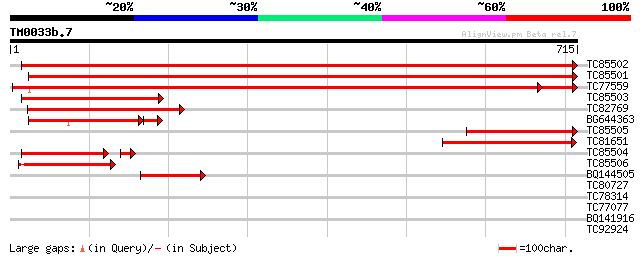
Query= TM0033b.7
(715 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC85502 homologue to SP|P27990|PALY_MEDSA Phenylalanine ammonia-... 1238 0.0
TC85501 homologue to SP|P45732|PALY_STYHU Phenylalanine ammonia-... 1234 0.0
TC77559 homologue to SP|Q9SMK9|PAL2_CICAR Phenylalanine ammonia-... 1062 0.0
TC85503 homologue to SP|P27990|PALY_MEDSA Phenylalanine ammonia-... 275 3e-74
TC82769 similar to SP|P19143|PAL3_PHAVU Phenylalanine ammonia-ly... 273 2e-73
BG644363 homologue to SP|P26600|PAL5 Phenylalanine ammonia-lyase... 232 9e-70
TC85505 homologue to SP|P27990|PALY_MEDSA Phenylalanine ammonia-... 250 1e-66
TC81651 similar to SP|P19143|PAL3_PHAVU Phenylalanine ammonia-ly... 223 3e-58
TC85504 homologue to SP|P27990|PALY_MEDSA Phenylalanine ammonia-... 177 1e-48
TC85506 similar to GP|7208616|gb|AAF40224.1| phenylalanine ammon... 178 6e-45
BQ144505 similar to SP|P45734|PALY_ Phenylalanine ammonia-lyase ... 114 1e-25
TC80727 similar to GP|15982870|gb|AAL09782.1 At1g21880/T26F17_5 ... 32 0.65
TC78314 weakly similar to PIR|H86356|H86356 probable UDP-glucose... 29 7.2
TC77077 similar to GP|9293909|dbj|BAB01812.1 sugar transporter p... 28 9.4
BQ141916 similar to GP|19172379|gb| mannitol transporter {Apium ... 28 9.4
TC92924 28 9.4
>TC85502 homologue to SP|P27990|PALY_MEDSA Phenylalanine ammonia-lyase (EC
4.3.1.5). [Alfalfa] {Medicago sativa}, complete
Length = 2415
Score = 1238 bits (3203), Expect = 0.0
Identities = 623/700 (89%), Positives = 659/700 (94%)
Frame = +2
Query: 16 NTVKTAATDPLSWGVAAESMKGSHLDEVKRMVAEYRKAVVKLGGETLTIAQVAAIAANHQ 75
N +K DPL+WGVAAE+MKGSHLDEVKRMV EYRK VV+LGGETLTI+QVAAIAA+
Sbjct: 179 NNMKVNEADPLNWGVAAEAMKGSHLDEVKRMVEEYRKPVVRLGGETLTISQVAAIAAHDH 358
Query: 76 GVSVELCESARAGVKASSDWVMNSMNNGTDSYGVTTGFGATSHRRTKNGNALQLELIRFL 135
GV VEL ESARAGV+ASSDWVM SMN GTDSYGVTTGFGATSHRRTK G ALQ ELIRFL
Sbjct: 359 GVQVELSESARAGVEASSDWVMESMNKGTDSYGVTTGFGATSHRRTKQGGALQKELIRFL 538
Query: 136 NAGIFGNGTESTHTLPQPATRAAMLVRINTLLQGYSGIRFEILEAITKLINNNITPCLPL 195
NAGIFGNGTES+HTLP ATRAAMLVRINTLLQGYSGIRFEILEAIT L+NNN+TPCLPL
Sbjct: 539 NAGIFGNGTESSHTLPHTATRAAMLVRINTLLQGYSGIRFEILEAITNLLNNNVTPCLPL 718
Query: 196 RGTVTASGDLVPLSYIAGLLTGRPNSKAVGPSGEVVNAKDAFQLAGIDSGFFELQPKEGL 255
RGT+TASGDLVPLSYIAGLLTGRPNSKA GPSGE++NAK+AF LAGI++ FFELQPKEGL
Sbjct: 719 RGTITASGDLVPLSYIAGLLTGRPNSKAHGPSGEILNAKEAFALAGINAEFFELQPKEGL 898
Query: 256 ALVNGTAVGSGLASIVLFDANILAILAEVLSAIFAEVMQGKPEFTDHLTHKLKHHPGQIE 315
ALVNGTAVGSGLASIVLF+ANILA+L+EVLSAIFAEVMQGKPEFTDHLTHKLKHHPGQIE
Sbjct: 899 ALVNGTAVGSGLASIVLFEANILAVLSEVLSAIFAEVMQGKPEFTDHLTHKLKHHPGQIE 1078
Query: 316 AAAIMEHILDGSSYMKAAKKLHEVDPLQKPKQDRYALRTSPQWLGPLIEVIRFSTKSIER 375
AAAIMEHILDGS+Y+KA KKLHE+DPLQKPKQDRYALRTSPQWLGPL+EVIRFSTKSIER
Sbjct: 1079AAAIMEHILDGSAYVKADKKLHEMDPLQKPKQDRYALRTSPQWLGPLVEVIRFSTKSIER 1258
Query: 376 EINSVNDNPLIDVSRNKALHGGNFQGTPIGVSMDNTRLALAAIGKLMFAQFTELVDDHYN 435
EINSVNDNPLIDVSRNKALHGGNFQGTPIGVSMDNTRLALA+IGKLMFAQF+ELV+D YN
Sbjct: 1259EINSVNDNPLIDVSRNKALHGGNFQGTPIGVSMDNTRLALASIGKLMFAQFSELVNDFYN 1438
Query: 436 NGLPSNLTASRNPSLDYGFKGAEIAMASYCSELQYLANPVTTHVQSAEQHNQDVNSLGLI 495
NGLPSNL+ASRNPSLDYGFKGAEIAMASYCSELQYLANPVTTHVQSAEQHNQDVNSLGLI
Sbjct: 1439NGLPSNLSASRNPSLDYGFKGAEIAMASYCSELQYLANPVTTHVQSAEQHNQDVNSLGLI 1618
Query: 496 SSRKTNEAIEILKLMSSTFLIALCQAIDLRHLEENLKYSVKSTVSQVAKRTLTTGVNGEL 555
SSRKT EAIEIL+LMSSTFLIALCQAIDLRHLEENLK SVK+TVSQVAK+TLT GVNGEL
Sbjct: 1619SSRKTYEAIEILQLMSSTFLIALCQAIDLRHLEENLKNSVKNTVSQVAKKTLTIGVNGEL 1798
Query: 556 HPSRFCEKDLLKVVDRETLFAYIDDPCLATYPLMQKLRQVLVDHALVNGEYEKDSKTSIF 615
HPSRFCEKDLLKVVDRE +FAYIDDPC ATYPL QKLRQVLVDHALVNGE EK+ TSIF
Sbjct: 1799HPSRFCEKDLLKVVDREHVFAYIDDPCSATYPLSQKLRQVLVDHALVNGESEKNLNTSIF 1978
Query: 616 QKIATFEDELKSLLPKEVESARAAYESGNPAMPNKINECRSYPLYKFVRKELGTELLTGE 675
QKIATFE+ELKSLLPKEVESAR AYESGNP +PNKIN CRSYPLYKFVR+ELGT LLTGE
Sbjct: 1979QKIATFEEELKSLLPKEVESARTAYESGNPTIPNKINGCRSYPLYKFVREELGTGLLTGE 2158
Query: 676 KTRSPGEECDKLFTAICQGKIIDPLLECLGEWNGAPLPIC 715
SPGE CDKLFTA+CQGKIIDPLLECLGEWNGAPLPIC
Sbjct: 2159NVISPGEVCDKLFTAMCQGKIIDPLLECLGEWNGAPLPIC 2278
>TC85501 homologue to SP|P45732|PALY_STYHU Phenylalanine ammonia-lyase (EC
4.3.1.5). [Townsville stylo] {Stylosanthes humilis},
partial (96%)
Length = 2567
Score = 1234 bits (3192), Expect = 0.0
Identities = 618/692 (89%), Positives = 655/692 (94%)
Frame = +3
Query: 24 DPLSWGVAAESMKGSHLDEVKRMVAEYRKAVVKLGGETLTIAQVAAIAANHQGVSVELCE 83
DPL+WG AAES+ GSHLDEVKRMV EYR +VK+GGETLTIAQVA IA++ GV VEL E
Sbjct: 192 DPLNWGAAAESLTGSHLDEVKRMVEEYRNPLVKIGGETLTIAQVAGIASHDSGVRVELSE 371
Query: 84 SARAGVKASSDWVMNSMNNGTDSYGVTTGFGATSHRRTKNGNALQLELIRFLNAGIFGNG 143
SARAGVKASSDWVM+SMN GTDSYGVTTGFGATSHRRTK G ALQ ELIRFLNAGIFGNG
Sbjct: 372 SARAGVKASSDWVMDSMNKGTDSYGVTTGFGATSHRRTKQGGALQKELIRFLNAGIFGNG 551
Query: 144 TESTHTLPQPATRAAMLVRINTLLQGYSGIRFEILEAITKLINNNITPCLPLRGTVTASG 203
TES TLP ATRAAMLVRINTLLQGYSGIRFEILEAITKL+NNNITPCLPLRGT+TASG
Sbjct: 552 TESNCTLPHTATRAAMLVRINTLLQGYSGIRFEILEAITKLLNNNITPCLPLRGTITASG 731
Query: 204 DLVPLSYIAGLLTGRPNSKAVGPSGEVVNAKDAFQLAGIDSGFFELQPKEGLALVNGTAV 263
DLVPLSYIAGLLTGRPNSKAVGPSGE++NAK+AFQLAGI S FFELQPKEGLALVNGTAV
Sbjct: 732 DLVPLSYIAGLLTGRPNSKAVGPSGEILNAKEAFQLAGIGSDFFELQPKEGLALVNGTAV 911
Query: 264 GSGLASIVLFDANILAILAEVLSAIFAEVMQGKPEFTDHLTHKLKHHPGQIEAAAIMEHI 323
GSGLASIVLF+AN+LA+L+EV+SAIFAEVMQGKPEFTDHLTHKLKHHPGQIEAAAIMEHI
Sbjct: 912 GSGLASIVLFEANVLAVLSEVMSAIFAEVMQGKPEFTDHLTHKLKHHPGQIEAAAIMEHI 1091
Query: 324 LDGSSYMKAAKKLHEVDPLQKPKQDRYALRTSPQWLGPLIEVIRFSTKSIEREINSVNDN 383
LDGS+Y+KAAKKLHE DPLQKPKQDRYALRTSPQWLGPLIEVIRFSTKSIEREINSVNDN
Sbjct: 1092LDGSAYVKAAKKLHETDPLQKPKQDRYALRTSPQWLGPLIEVIRFSTKSIEREINSVNDN 1271
Query: 384 PLIDVSRNKALHGGNFQGTPIGVSMDNTRLALAAIGKLMFAQFTELVDDHYNNGLPSNLT 443
PLIDVSRNKA+HGGNFQGTPIGVSMDNTRLALA+IGKLMFAQF+ELV+D YNNGLPSNLT
Sbjct: 1272PLIDVSRNKAIHGGNFQGTPIGVSMDNTRLALASIGKLMFAQFSELVNDFYNNGLPSNLT 1451
Query: 444 ASRNPSLDYGFKGAEIAMASYCSELQYLANPVTTHVQSAEQHNQDVNSLGLISSRKTNEA 503
ASRNPSLDYGFKG+EIAMASYCSELQYLANPVTTHVQSAEQHNQDVNSLGLISSRKTNEA
Sbjct: 1452ASRNPSLDYGFKGSEIAMASYCSELQYLANPVTTHVQSAEQHNQDVNSLGLISSRKTNEA 1631
Query: 504 IEILKLMSSTFLIALCQAIDLRHLEENLKYSVKSTVSQVAKRTLTTGVNGELHPSRFCEK 563
IEILKLMSSTFLIALCQAIDLRHLEENL+ +VK+TVSQVAKRTLTTGVNGELHPSRFCEK
Sbjct: 1632IEILKLMSSTFLIALCQAIDLRHLEENLRNTVKNTVSQVAKRTLTTGVNGELHPSRFCEK 1811
Query: 564 DLLKVVDRETLFAYIDDPCLATYPLMQKLRQVLVDHALVNGEYEKDSKTSIFQKIATFED 623
DLLKVVDRE +FAY DDPCLATYPLMQKLRQVLVDHALVN E EK+S TSIFQKIATFED
Sbjct: 1812DLLKVVDREYVFAYADDPCLATYPLMQKLRQVLVDHALVNTEGEKNSNTSIFQKIATFED 1991
Query: 624 ELKSLLPKEVESARAAYESGNPAMPNKINECRSYPLYKFVRKELGTELLTGEKTRSPGEE 683
ELK++LPKEVESAR AYE+G + NKI ECRSYPLYKFVR+ELGT LLTGEK SPGEE
Sbjct: 1992ELKAILPKEVESARTAYENGQSGISNKIKECRSYPLYKFVREELGTALLTGEKVISPGEE 2171
Query: 684 CDKLFTAICQGKIIDPLLECLGEWNGAPLPIC 715
CDKLFTA+CQGKI+DPL+ECLGEWNGAPLPIC
Sbjct: 2172CDKLFTAMCQGKIVDPLMECLGEWNGAPLPIC 2267
>TC77559 homologue to SP|Q9SMK9|PAL2_CICAR Phenylalanine ammonia-lyase 2 (EC
4.3.1.5). [Chickpea Garbanzo] {Cicer arietinum},
partial (96%)
Length = 2406
Score = 1062 bits (2746), Expect(2) = 0.0
Identities = 530/671 (78%), Positives = 597/671 (87%), Gaps = 3/671 (0%)
Frame = +3
Query: 4 NTNATQNGSIFCNTVKTAAT---DPLSWGVAAESMKGSHLDEVKRMVAEYRKAVVKLGGE 60
N+N + S+ T + DPL+WG+AA+SMKGSHLDEVKRMV EYRK VV LGG+
Sbjct: 90 NSNGSNGSSMNLRNGGTKTSNNNDPLNWGIAADSMKGSHLDEVKRMVEEYRKPVVPLGGK 269
Query: 61 TLTIAQVAAIAANHQGVSVELCESARAGVKASSDWVMNSMNNGTDSYGVTTGFGATSHRR 120
LTIAQVAA+A GV+VEL E AR+GVKASSDWV++SMN GTDSYGVTTGFGATSHRR
Sbjct: 270 GLTIAQVAAVATYSTGVAVELAEEARSGVKASSDWVVDSMNKGTDSYGVTTGFGATSHRR 449
Query: 121 TKNGNALQLELIRFLNAGIFGNGTESTHTLPQPATRAAMLVRINTLLQGYSGIRFEILEA 180
T G+ALQ ELIRFLNAGIFGNGTE++ TLP ATRA MLVRINTLLQGYSGIRFEI+EA
Sbjct: 450 TNKGSALQSELIRFLNAGIFGNGTEASQTLPPTATRAGMLVRINTLLQGYSGIRFEIMEA 629
Query: 181 ITKLINNNITPCLPLRGTVTASGDLVPLSYIAGLLTGRPNSKAVGPSGEVVNAKDAFQLA 240
I K +N+NITPCLPLRGT+TASGDL+PLSY+AGLL GRPNSK++GP+G+V+NA++AFQLA
Sbjct: 630 IAKFLNHNITPCLPLRGTITASGDLIPLSYVAGLLIGRPNSKSIGPNGQVLNAQEAFQLA 809
Query: 241 GIDSGFFELQPKEGLALVNGTAVGSGLASIVLFDANILAILAEVLSAIFAEVMQGKPEFT 300
GI++GFFELQPKEGLALVNGTAVGSGLAS+VLFD N+L +L+E+LSAIFAEVM GKP+FT
Sbjct: 810 GIETGFFELQPKEGLALVNGTAVGSGLASLVLFDTNLLVVLSEILSAIFAEVMLGKPQFT 989
Query: 301 DHLTHKLKHHPGQIEAAAIMEHILDGSSYMKAAKKLHEVDPLQKPKQDRYALRTSPQWLG 360
DHL HK+K HPGQIEAAAIMEHILDGS Y KAA+K+HE++PLQKPKQDRYA+RTSPQWLG
Sbjct: 990 DHLIHKVKLHPGQIEAAAIMEHILDGSYYGKAAQKVHEINPLQKPKQDRYAIRTSPQWLG 1169
Query: 361 PLIEVIRFSTKSIEREINSVNDNPLIDVSRNKALHGGNFQGTPIGVSMDNTRLALAAIGK 420
P IEVIR++TK IEREINSVNDNPLIDVSR+KALHGGNFQGTPIGVSMDNTRLA+AAIGK
Sbjct: 1170PQIEVIRYATKMIEREINSVNDNPLIDVSRDKALHGGNFQGTPIGVSMDNTRLAIAAIGK 1349
Query: 421 LMFAQFTELVDDHYNNGLPSNLTASRNPSLDYGFKGAEIAMASYCSELQYLANPVTTHVQ 480
LMFAQFTELV+D YNNGLPS+LTASRNPSLDYGFKGAE+AMASYCSELQYLA+PVTTHVQ
Sbjct: 1350LMFAQFTELVNDIYNNGLPSSLTASRNPSLDYGFKGAEVAMASYCSELQYLASPVTTHVQ 1529
Query: 481 SAEQHNQDVNSLGLISSRKTNEAIEILKLMSSTFLIALCQAIDLRHLEENLKYSVKSTVS 540
SAEQHNQDVNSLGLIS+RKT EA+EI KLMSSTFL+ALCQAIDLRH+EEN K VK+TVS
Sbjct: 1530SAEQHNQDVNSLGLISARKTAEAVEIWKLMSSTFLVALCQAIDLRHIEENFKSVVKNTVS 1709
Query: 541 QVAKRTLTTGVNGELHPSRFCEKDLLKVVDRETLFAYIDDPCLATYPLMQKLRQVLVDHA 600
QVAKR LT GVNGELHPSRFCEKDLL VV+ E +F YIDDPC YPLMQKLR VLVDHA
Sbjct: 1710QVAKRILTVGVNGELHPSRFCEKDLLNVVEGEYVFTYIDDPCSPIYPLMQKLRYVLVDHA 1889
Query: 601 LVNGEYEKDSKTSIFQKIATFEDELKSLLPKEVESARAAYESGNPAMPNKINECRSYPLY 660
L NG+ E +S TSIFQKI FE+ELK+LLPKEVE+AR E+GNPA+PN I ECRSYPLY
Sbjct: 1890LQNGDKEANSSTSIFQKIGAFEEELKALLPKEVENARVEIENGNPAVPNMIKECRSYPLY 2069
Query: 661 KFVRKELGTEL 671
KF+R+ LGT L
Sbjct: 2070KFMRETLGTSL 2102
Score = 82.0 bits (201), Expect(2) = 0.0
Identities = 33/48 (68%), Positives = 40/48 (82%)
Frame = +1
Query: 668 GTELLTGEKTRSPGEECDKLFTAICQGKIIDPLLECLGEWNGAPLPIC 715
G LTGEK +SPGE+CDK+FTA+C G+ IDP+L+CL EWNG PLPIC
Sbjct: 2092 GQVCLTGEKIKSPGEDCDKVFTAMCDGRFIDPMLDCLKEWNGVPLPIC 2235
>TC85503 homologue to SP|P27990|PALY_MEDSA Phenylalanine ammonia-lyase (EC
4.3.1.5). [Alfalfa] {Medicago sativa}, partial (27%)
Length = 724
Score = 275 bits (704), Expect = 3e-74
Identities = 141/178 (79%), Positives = 151/178 (84%)
Frame = +3
Query: 16 NTVKTAATDPLSWGVAAESMKGSHLDEVKRMVAEYRKAVVKLGGETLTIAQVAAIAANHQ 75
N +K DPL+WGVAAE+MKGSHLDEVKRMV EYRK VV+LGGETLTI+QVAAIAA+
Sbjct: 189 NNMKVNEADPLNWGVAAEAMKGSHLDEVKRMVEEYRKPVVRLGGETLTISQVAAIAAHDH 368
Query: 76 GVSVELCESARAGVKASSDWVMNSMNNGTDSYGVTTGFGATSHRRTKNGNALQLELIRFL 135
GV VEL ESARAGV+ASSDWVM SMN GTDSYGVTTGFGATSHRRTK G ALQ ELIRFL
Sbjct: 369 GVQVELSESARAGVEASSDWVMESMNKGTDSYGVTTGFGATSHRRTKQGGALQKELIRFL 548
Query: 136 NAGIFGNGTESTHTLPQPATRAAMLVRINTLLQGYSGIRFEILEAITKLINNNITPCL 193
NAGIFG G+ES+HTLP ATRAAMLVRINTLLQGYSGI FEILEAIT L+ CL
Sbjct: 549 NAGIFGKGSESSHTLPHTATRAAMLVRINTLLQGYSGIXFEILEAITNLLTTTSLHCL 722
>TC82769 similar to SP|P19143|PAL3_PHAVU Phenylalanine ammonia-lyase class
III (EC 4.3.1.5). [Kidney bean French bean] {Phaseolus
vulgaris}, partial (27%)
Length = 652
Score = 273 bits (697), Expect = 2e-73
Identities = 139/199 (69%), Positives = 163/199 (81%), Gaps = 1/199 (0%)
Frame = +2
Query: 23 TDPLSWGVAAESMKGSHLDEVKRMVAEYRKAVVKLGGE-TLTIAQVAAIAANHQGVSVEL 81
+DPL+W AA S+KGSHLDEVKRM+AEY+K V+ LGG TLTI+QVAA++ + V VEL
Sbjct: 56 SDPLNWNSAANSLKGSHLDEVKRMLAEYKKPVICLGGVGTLTISQVAAVSNSSSHVKVEL 235
Query: 82 CESARAGVKASSDWVMNSMNNGTDSYGVTTGFGATSHRRTKNGNALQLELIRFLNAGIFG 141
ESARAGV+AS DW+ ++ GT YGVTTGFGA SHRRT+ G ALQ E++RFLN IFG
Sbjct: 236 SESARAGVEASCDWISENIVKGTPIYGVTTGFGAASHRRTEQGFALQKEMVRFLNCAIFG 415
Query: 142 NGTESTHTLPQPATRAAMLVRINTLLQGYSGIRFEILEAITKLINNNITPCLPLRGTVTA 201
+E +HTLP ATRAAMLVR+NTLLQGYSGIRFEIL AITKL+N+N+TP LPLRGTVTA
Sbjct: 416 RESELSHTLPSSATRAAMLVRVNTLLQGYSGIRFEILXAITKLLNHNVTPILPLRGTVTA 595
Query: 202 SGDLVPLSYIAGLLTGRPN 220
SGDL+PLSYIA LLTG N
Sbjct: 596 SGDLIPLSYIAALLTGXRN 652
>BG644363 homologue to SP|P26600|PAL5 Phenylalanine ammonia-lyase (EC
4.3.1.5) (PAL). [Tomato] {Lycopersicon esculentum},
partial (26%)
Length = 687
Score = 232 bits (591), Expect(2) = 9e-70
Identities = 118/148 (79%), Positives = 131/148 (87%), Gaps = 3/148 (2%)
Frame = +3
Query: 24 DPLSWGVAAESMKGSHLDEVKRMVAEYRKAVVKLGGETLTIAQVAAIAA---NHQGVSVE 80
DPL+W +AAES++GSHLDEVK+MV E+RK +VKLGGETLT+AQVA+IA GV VE
Sbjct: 171 DPLNWEMAAESLRGSHLDEVKKMVDEFRKPIVKLGGETLTVAQVASIANADNKSNGVKVE 350
Query: 81 LCESARAGVKASSDWVMNSMNNGTDSYGVTTGFGATSHRRTKNGNALQLELIRFLNAGIF 140
L ESARAGVKASSDWVM+SM+ GTDSYGVTTGFGATSHRRTKNG ALQ ELIRFLNAG+F
Sbjct: 351 LSESARAGVKASSDWVMDSMSKGTDSYGVTTGFGATSHRRTKNGGALQKELIRFLNAGVF 530
Query: 141 GNGTESTHTLPQPATRAAMLVRINTLLQ 168
GNGTES+HTLP ATRAAMLVRINTLLQ
Sbjct: 531 GNGTESSHTLPHSATRAAMLVRINTLLQ 614
Score = 50.8 bits (120), Expect(2) = 9e-70
Identities = 23/24 (95%), Positives = 24/24 (99%)
Frame = +2
Query: 169 GYSGIRFEILEAITKLINNNITPC 192
GYSGIRFEILEAITKLIN+NITPC
Sbjct: 614 GYSGIRFEILEAITKLINSNITPC 685
>TC85505 homologue to SP|P27990|PALY_MEDSA Phenylalanine ammonia-lyase (EC
4.3.1.5). [Alfalfa] {Medicago sativa}, partial (19%)
Length = 652
Score = 250 bits (639), Expect = 1e-66
Identities = 121/139 (87%), Positives = 126/139 (90%)
Frame = +1
Query: 577 YIDDPCLATYPLMQKLRQVLVDHALVNGEYEKDSKTSIFQKIATFEDELKSLLPKEVESA 636
YIDDPC ATYPL QKLRQVLVDHALVNGE EK+ TSIFQKIATFE+ELKSLLPKEVESA
Sbjct: 1 YIDDPCSATYPLSQKLRQVLVDHALVNGESEKNLNTSIFQKIATFEEELKSLLPKEVESA 180
Query: 637 RAAYESGNPAMPNKINECRSYPLYKFVRKELGTELLTGEKTRSPGEECDKLFTAICQGKI 696
R AYESGNP +PNKIN CRSYPLYKFVR+ELGT LLTGE SPGE CDKLFTA+CQGKI
Sbjct: 181 RTAYESGNPTIPNKINGCRSYPLYKFVREELGTGLLTGENVISPGEVCDKLFTAMCQGKI 360
Query: 697 IDPLLECLGEWNGAPLPIC 715
IDPLLECLGEWNGAPLPIC
Sbjct: 361 IDPLLECLGEWNGAPLPIC 417
>TC81651 similar to SP|P19143|PAL3_PHAVU Phenylalanine ammonia-lyase class
III (EC 4.3.1.5). [Kidney bean French bean] {Phaseolus
vulgaris}, partial (23%)
Length = 708
Score = 223 bits (567), Expect = 3e-58
Identities = 107/169 (63%), Positives = 131/169 (77%)
Frame = +2
Query: 546 TLTTGVNGELHPSRFCEKDLLKVVDRETLFAYIDDPCLATYPLMQKLRQVLVDHALVNGE 605
TL E+ P R CE++LLKVVDRE +F+YIDDP TYPLM KL+QVL +HA ++
Sbjct: 2 TLIIDDREEIDPFRHCEENLLKVVDREYVFSYIDDPFNVTYPLMPKLKQVLYEHAHISAI 181
Query: 606 YEKDSKTSIFQKIATFEDELKSLLPKEVESARAAYESGNPAMPNKINECRSYPLYKFVRK 665
KD+K+S F+KI FEDELKSLLPKEVESAR A+E GN +PN+I ECRSYPLYKFVR+
Sbjct: 182 NNKDAKSSTFEKIGAFEDELKSLLPKEVESARVAFEKGNSEIPNRIKECRSYPLYKFVRE 361
Query: 666 ELGTELLTGEKTRSPGEECDKLFTAICQGKIIDPLLECLGEWNGAPLPI 714
EL LLTGEK +P EE +K+FTA+CQ KI+DP+LECLG+W G P+PI
Sbjct: 362 ELKIGLLTGEKDVTPDEEFEKVFTAMCQAKIVDPILECLGDWKGVPIPI 508
>TC85504 homologue to SP|P27990|PALY_MEDSA Phenylalanine ammonia-lyase (EC
4.3.1.5). [Alfalfa] {Medicago sativa}, partial (22%)
Length = 654
Score = 177 bits (448), Expect(2) = 1e-48
Identities = 88/109 (80%), Positives = 95/109 (86%)
Frame = +2
Query: 16 NTVKTAATDPLSWGVAAESMKGSHLDEVKRMVAEYRKAVVKLGGETLTIAQVAAIAANHQ 75
N +K DPL+WGVAAE+MKGSHLDEVKRMV EYRK VV+LGGETLTI+QVAAIAA+
Sbjct: 215 NNMKVNEADPLNWGVAAEAMKGSHLDEVKRMVEEYRKPVVRLGGETLTISQVAAIAAHDH 394
Query: 76 GVSVELCESARAGVKASSDWVMNSMNNGTDSYGVTTGFGATSHRRTKNG 124
GV VEL ESARAGV+ASSDWVM SMN GTDSYGVTTGFGATSHRRTK G
Sbjct: 395 GVQVELSESARAGVEASSDWVMESMNKGTDSYGVTTGFGATSHRRTKQG 541
Score = 35.4 bits (80), Expect(2) = 1e-48
Identities = 15/19 (78%), Positives = 16/19 (83%)
Frame = +1
Query: 140 FGNGTESTHTLPQPATRAA 158
FGNGTES HTLP ATRA+
Sbjct: 589 FGNGTESNHTLPHTATRAS 645
Score = 32.3 bits (72), Expect = 0.65
Identities = 18/27 (66%), Positives = 18/27 (66%), Gaps = 1/27 (3%)
Frame = +3
Query: 115 ATSHRRTKN-GNALQLELIRFLNAGIF 140
A H N G ALQ ELIRFLNAGIF
Sbjct: 510 APPHTAVPNKGGALQKELIRFLNAGIF 590
>TC85506 similar to GP|7208616|gb|AAF40224.1| phenylalanine ammonia-lyase 2
{Rubus idaeus}, partial (15%)
Length = 1025
Score = 178 bits (452), Expect = 6e-45
Identities = 92/122 (75%), Positives = 100/122 (81%)
Frame = +3
Query: 12 SIFCNTVKTAATDPLSWGVAAESMKGSHLDEVKRMVAEYRKAVVKLGGETLTIAQVAAIA 71
S FC + DPL+WG AAES+ GSHLDEVKRMV EYR +VK+GGETLTIAQVA IA
Sbjct: 9 STFC--LSKNGGDPLNWGAAAESLTGSHLDEVKRMVEEYRNPLVKIGGETLTIAQVAGIA 182
Query: 72 ANHQGVSVELCESARAGVKASSDWVMNSMNNGTDSYGVTTGFGATSHRRTKNGNALQLEL 131
++ GV VEL ESARAGVKASSDWVM+SMN GTDSYGVTTGFGATSHRRTK G ALQ EL
Sbjct: 183 SHDSGVRVELSESARAGVKASSDWVMDSMNKGTDSYGVTTGFGATSHRRTKQGGALQKEL 362
Query: 132 IR 133
IR
Sbjct: 363 IR 368
>BQ144505 similar to SP|P45734|PALY_ Phenylalanine ammonia-lyase (EC
4.3.1.5). [Subterranean clover] {Trifolium
subterraneum}, partial (11%)
Length = 863
Score = 114 bits (285), Expect = 1e-25
Identities = 56/82 (68%), Positives = 65/82 (78%)
Frame = +2
Query: 166 LLQGYSGIRFEILEAITKLINNNITPCLPLRGTVTASGDLVPLSYIAGLLTGRPNSKAVG 225
LLQGYSGI FEILEAIT L+NNN+TPC PL GT+TAS D VPLS+IAGLLTG PN K G
Sbjct: 11 LLQGYSGITFEILEAITNLLNNNVTPCSPLAGTITASRDSVPLSHIAGLLTGIPNCKPHG 190
Query: 226 PSGEVVNAKDAFQLAGIDSGFF 247
P G+++NAK F L GI++ F
Sbjct: 191 PCGQILNAKHPFALTGINAESF 256
>TC80727 similar to GP|15982870|gb|AAL09782.1 At1g21880/T26F17_5
{Arabidopsis thaliana}, partial (69%)
Length = 1019
Score = 32.3 bits (72), Expect = 0.65
Identities = 31/121 (25%), Positives = 50/121 (40%), Gaps = 8/121 (6%)
Frame = +3
Query: 319 IMEHILDGSSYMKAAKKLHEVDPLQKPKQDRYALRTSPQWLGPLIEVIRFSTKSIE--RE 376
+ HIL Y+K + +D ++K Y +R S L + + I S + RE
Sbjct: 321 VEHHILPSKLYLKIPIQCSCIDGIRKSVSTNYKIRPS-DTLSSIADSIYGGLVSSDQLRE 497
Query: 377 INSVNDNPLIDVSRNKAL------HGGNFQGTPIGVSMDNTRLALAAIGKLMFAQFTELV 430
NSV D ++DV +N + G+ G P + M L ++ + FT L
Sbjct: 498 ANSVTDPNVLDVGQNLVVPLPCTCFNGSDNGLP-AIYMSYVVQPLDSLNNIAARYFTTLT 674
Query: 431 D 431
D
Sbjct: 675 D 677
>TC78314 weakly similar to PIR|H86356|H86356 probable UDP-glucose
glucosyltransferase [imported] - Arabidopsis thaliana,
partial (33%)
Length = 1163
Score = 28.9 bits (63), Expect = 7.2
Identities = 21/59 (35%), Positives = 30/59 (50%)
Frame = +2
Query: 383 NPLIDVSRNKALHGGNFQGTPIGVSMDNTRLALAAIGKLMFAQFTELVDDHYNNGLPSN 441
NPLI ++ K LH F T + ++ RL L + G F FT+ + +GLPSN
Sbjct: 194 NPLIKLA--KLLHLRGFHITFVNTEYNHKRL-LKSRGPNAFVGFTDFTFEAIPDGLPSN 361
>TC77077 similar to GP|9293909|dbj|BAB01812.1 sugar transporter protein
{Arabidopsis thaliana}, partial (85%)
Length = 1752
Score = 28.5 bits (62), Expect = 9.4
Identities = 11/21 (52%), Positives = 16/21 (75%)
Frame = +1
Query: 331 KAAKKLHEVDPLQKPKQDRYA 351
KA K L + DP +KPK+++YA
Sbjct: 49 KAQKNLQDFDPQKKPKRNKYA 111
>BQ141916 similar to GP|19172379|gb| mannitol transporter {Apium graveolens
var. dulce}, partial (9%)
Length = 824
Score = 28.5 bits (62), Expect = 9.4
Identities = 11/21 (52%), Positives = 16/21 (75%)
Frame = +1
Query: 331 KAAKKLHEVDPLQKPKQDRYA 351
KA K L + DP +KPK+++YA
Sbjct: 49 KAQKNLQDFDPQKKPKRNKYA 111
>TC92924
Length = 713
Score = 28.5 bits (62), Expect = 9.4
Identities = 24/62 (38%), Positives = 33/62 (52%), Gaps = 1/62 (1%)
Frame = +2
Query: 527 LEENLKYSVKSTVSQVAKRTLTTGVNGELHPSRFCEKDLLKVVDRETL-FAYIDDPCLAT 585
L+EN KYS KST++ K+ + G EL ++ K+ +E AY D PCL T
Sbjct: 365 LKENSKYS-KSTLT--LKQIASDGSCPELESAK------QKLYKKEVFELAYQDLPCLLT 517
Query: 586 YP 587
YP
Sbjct: 518 YP 523
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.316 0.133 0.379
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 18,485,682
Number of Sequences: 36976
Number of extensions: 220408
Number of successful extensions: 1065
Number of sequences better than 10.0: 32
Number of HSP's better than 10.0 without gapping: 1056
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1061
length of query: 715
length of database: 9,014,727
effective HSP length: 103
effective length of query: 612
effective length of database: 5,206,199
effective search space: 3186193788
effective search space used: 3186193788
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 62 (28.5 bits)
Lotus: description of TM0033b.7


