

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0033b.5
(88 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
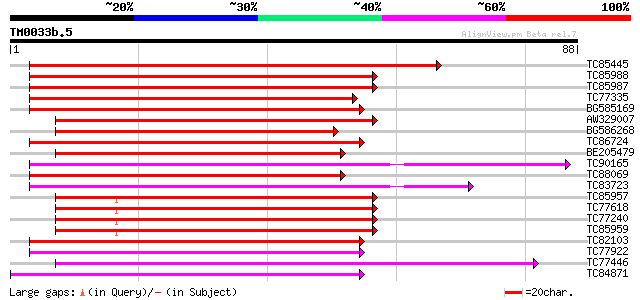
Sequences producing significant alignments: (bits) Value
TC85445 homologue to GP|1370180|emb|CAA98167.1 RAB5B {Lotus japo... 75 3e-15
TC85988 homologue to PIR|S49225|S49225 guanine nucleotide regula... 72 5e-14
TC85987 homologue to GP|1370178|emb|CAA98166.1 RAB5A {Lotus japo... 68 7e-13
TC77335 homologue to GP|1370176|emb|CAA98165.1 RAB2A {Lotus japo... 56 2e-09
BG585169 similar to GP|1370152|emb RAB11F {Lotus japonicus}, par... 55 4e-09
AW329007 weakly similar to GP|12834379|dbj data source:SPTR sou... 50 1e-07
BG586268 homologue to PIR|S71585|S71 GTP-binding protein GB2 - A... 50 2e-07
TC86724 homologue to PIR|T06445|T06445 GTP-binding protein - gar... 50 2e-07
BE205479 homologue to PIR|T06443|T06 GTP-binding protein - garde... 50 2e-07
TC90165 similar to PIR|H84610|H84610 probable GTP-binding protei... 50 2e-07
TC88069 homologue to GP|1370154|emb|CAA98183.1 RAB11G {Lotus jap... 48 7e-07
TC83723 homologue to PIR|T01588|T01588 GTP-binding protein At2g4... 46 2e-06
TC85957 homologue to PIR|S57471|S57471 GTP-binding protein GTP6 ... 45 4e-06
TC77618 homologue to PIR|S57478|S57478 GTP-binding protein GTP13... 45 5e-06
TC77240 homologue to GP|1370196|emb|CAA98175.1 RAB8D {Lotus japo... 45 6e-06
TC85959 homologue to PIR|S57471|S57471 GTP-binding protein GTP6 ... 45 6e-06
TC82103 similar to GP|1370144|emb|CAA98178.1 RAB11B {Lotus japon... 44 8e-06
TC77922 homologue to PIR|T06448|T06448 GTP-binding protein - gar... 44 8e-06
TC77446 homologue to GP|303750|dbj|BAA02116.1| GTP-binding prote... 44 8e-06
TC84871 similar to GP|1370172|emb|CAA98163.1 RAB1X {Lotus japoni... 43 2e-05
>TC85445 homologue to GP|1370180|emb|CAA98167.1 RAB5B {Lotus japonicus},
partial (98%)
Length = 1147
Score = 75.5 bits (184), Expect = 3e-15
Identities = 35/64 (54%), Positives = 49/64 (75%)
Frame = +1
Query: 4 LVMALVANKSDLEPKREVEIEEGGQFAQENGMFYMETSAKTVENINDLFFEIGRGFESPG 63
+VMALV NK+DL+ KREV +E+G +A++NGMF++ETSAKT +NIN+LF EI + P
Sbjct: 601 IVMALVGNKADLQEKREVAVEDGMDYAEKNGMFFIETSAKTADNINELFEEIAKRLPRPS 780
Query: 64 RSSY 67
+ Y
Sbjct: 781 VT*Y 792
>TC85988 homologue to PIR|S49225|S49225 guanine nucleotide regulatory
protein - fava bean, complete
Length = 955
Score = 71.6 bits (174), Expect = 5e-14
Identities = 32/54 (59%), Positives = 43/54 (79%)
Frame = +1
Query: 4 LVMALVANKSDLEPKREVEIEEGGQFAQENGMFYMETSAKTVENINDLFFEIGR 57
+VMAL NKSDLE KR+V EE +A+ENG+F+METSAK+ N+ND+F+EI +
Sbjct: 475 MVMALAGNKSDLEDKRKVTAEEARVYAEENGLFFMETSAKSAANVNDVFYEIAK 636
>TC85987 homologue to GP|1370178|emb|CAA98166.1 RAB5A {Lotus japonicus},
complete
Length = 1028
Score = 67.8 bits (164), Expect = 7e-13
Identities = 29/54 (53%), Positives = 42/54 (77%)
Frame = +2
Query: 4 LVMALVANKSDLEPKREVEIEEGGQFAQENGMFYMETSAKTVENINDLFFEIGR 57
+V+A NK+DLE KR+V EE +A+ENG+F++ETSAKT N+ND+F+EI +
Sbjct: 530 MVVAFAGNKADLEDKRKVTAEEARVYAEENGLFFIETSAKTAANVNDIFYEIAK 691
>TC77335 homologue to GP|1370176|emb|CAA98165.1 RAB2A {Lotus japonicus},
complete
Length = 1121
Score = 56.2 bits (134), Expect = 2e-09
Identities = 24/51 (47%), Positives = 35/51 (68%)
Frame = +3
Query: 4 LVMALVANKSDLEPKREVEIEEGGQFAQENGMFYMETSAKTVENINDLFFE 54
+ + L+ NK DL +R V EEG QFA+ENG+ +ME SAKT +N+ + F +
Sbjct: 564 MTIMLIGNKCDLAHRRAVSTEEGEQFAKENGLIFMEASAKTAQNVEEAFIK 716
>BG585169 similar to GP|1370152|emb RAB11F {Lotus japonicus}, partial (83%)
Length = 668
Score = 55.5 bits (132), Expect = 4e-09
Identities = 28/52 (53%), Positives = 36/52 (68%)
Frame = +3
Query: 4 LVMALVANKSDLEPKREVEIEEGGQFAQENGMFYMETSAKTVENINDLFFEI 55
+V+ LVANKSDL REVE EEG FA++ G+ +METSA N+ D F E+
Sbjct: 471 MVIILVANKSDLSQSREVEKEEGKGFAEKEGLCFMETSALQNLNVEDAFLEM 626
>AW329007 weakly similar to GP|12834379|dbj data source:SPTR source
key:P10536 evidence:ISS~homolog to RAS-RELATED PROTEIN
RAB-1B~putative, partial (42%)
Length = 482
Score = 50.4 bits (119), Expect = 1e-07
Identities = 25/50 (50%), Positives = 32/50 (64%)
Frame = +2
Query: 8 LVANKSDLEPKREVEIEEGGQFAQENGMFYMETSAKTVENINDLFFEIGR 57
LV NKSDLE R+VEI EG Q +E M + ETSAK N+ ++F + R
Sbjct: 275 LVGNKSDLEKDRQVEINEGRQLGKERDMPFFETSAKDGSNVTEVFSLVAR 424
>BG586268 homologue to PIR|S71585|S71 GTP-binding protein GB2 - Arabidopsis
thaliana, partial (49%)
Length = 565
Score = 50.1 bits (118), Expect = 2e-07
Identities = 21/44 (47%), Positives = 32/44 (72%)
Frame = +1
Query: 8 LVANKSDLEPKREVEIEEGGQFAQENGMFYMETSAKTVENINDL 51
L+ NK DL KR V EEG QFA+E+G+ ++E SA+T +N+ ++
Sbjct: 127 LIGNKCDLSHKRAVSKEEGQQFAKEHGLLFLEASARTAQNVEEV 258
>TC86724 homologue to PIR|T06445|T06445 GTP-binding protein - garden pea,
complete
Length = 1037
Score = 50.1 bits (118), Expect = 2e-07
Identities = 22/52 (42%), Positives = 35/52 (67%)
Frame = +3
Query: 4 LVMALVANKSDLEPKREVEIEEGGQFAQENGMFYMETSAKTVENINDLFFEI 55
+V+ LV NKSDLE +R+V E+ +FA++ G+F++ETSA N+ F +
Sbjct: 540 IVIILVGNKSDLENQRDVPTEDAKEFAEKEGLFFLETSALQATNVEASFMTV 695
>BE205479 homologue to PIR|T06443|T06 GTP-binding protein - garden pea,
partial (77%)
Length = 499
Score = 49.7 bits (117), Expect = 2e-07
Identities = 25/45 (55%), Positives = 29/45 (63%)
Frame = +2
Query: 8 LVANKSDLEPKREVEIEEGGQFAQENGMFYMETSAKTVENINDLF 52
LV NK DLE REV IEEG A+E G+F+METSA N+ F
Sbjct: 314 LVGNKCDLENIREVSIEEGKALAEEEGLFFMETSALDSTNVQTAF 448
>TC90165 similar to PIR|H84610|H84610 probable GTP-binding protein
[imported] - Arabidopsis thaliana, partial (62%)
Length = 742
Score = 49.7 bits (117), Expect = 2e-07
Identities = 30/84 (35%), Positives = 46/84 (54%)
Frame = +2
Query: 4 LVMALVANKSDLEPKREVEIEEGGQFAQENGMFYMETSAKTVENINDLFFEIGRGFESPG 63
+++ LV NK+DL KR+ IEEG ++E G+ ++ETSAK NI LF +I PG
Sbjct: 113 VIIVLVGNKTDLVEKRQGSIEEGDAKSKELGIMFIETSAKAGFNIKPLFRKIASAL--PG 286
Query: 64 RSSYAPTKEWASAGLSSMPPVSCS 87
+ + T + ++ P V S
Sbjct: 287 METLSSTNKEDMVDVNLKPTVHSS 358
>TC88069 homologue to GP|1370154|emb|CAA98183.1 RAB11G {Lotus japonicus},
partial (79%)
Length = 606
Score = 47.8 bits (112), Expect = 7e-07
Identities = 23/49 (46%), Positives = 30/49 (60%)
Frame = +2
Query: 4 LVMALVANKSDLEPKREVEIEEGGQFAQENGMFYMETSAKTVENINDLF 52
+ M LV NK DL+ R+V IEEG A+ G+F+METSA N+ F
Sbjct: 425 VAMMLVGNKCDLDNIRDVSIEEGKSLAESEGLFFMETSALDSTNVKTAF 571
>TC83723 homologue to PIR|T01588|T01588 GTP-binding protein At2g44610 -
Arabidopsis thaliana, partial (67%)
Length = 574
Score = 46.2 bits (108), Expect = 2e-06
Identities = 26/69 (37%), Positives = 40/69 (57%)
Frame = +3
Query: 4 LVMALVANKSDLEPKREVEIEEGGQFAQENGMFYMETSAKTVENINDLFFEIGRGFESPG 63
+++ LV NK+DL KR+V EEG ++E + ++E SAK NI LF +I PG
Sbjct: 165 VIVVLVGNKTDLVEKRQVSTEEGEAKSRELNVMFIEASAKAGFNIKALFRKIAAAL--PG 338
Query: 64 RSSYAPTKE 72
+ + TK+
Sbjct: 339 METLSTTKQ 365
>TC85957 homologue to PIR|S57471|S57471 GTP-binding protein GTP6 - garden
pea, complete
Length = 1148
Score = 45.4 bits (106), Expect = 4e-06
Identities = 24/51 (47%), Positives = 33/51 (64%), Gaps = 1/51 (1%)
Frame = +3
Query: 8 LVANKSDL-EPKREVEIEEGGQFAQENGMFYMETSAKTVENINDLFFEIGR 57
LV NK+D+ E KR V +G A E G+ + ETSAKT N++++FF I R
Sbjct: 579 LVGNKADMDESKRAVPTSKGQALADEYGIKFFETSAKTNMNVDEVFFSIAR 731
>TC77618 homologue to PIR|S57478|S57478 GTP-binding protein GTP13 - garden
pea, complete
Length = 1136
Score = 45.1 bits (105), Expect = 5e-06
Identities = 24/51 (47%), Positives = 32/51 (62%), Gaps = 1/51 (1%)
Frame = +2
Query: 8 LVANKSDL-EPKREVEIEEGGQFAQENGMFYMETSAKTVENINDLFFEIGR 57
LV NK+D+ E KR V +G A E G+ + ETSAKT N+ ++FF I R
Sbjct: 590 LVGNKADMDESKRAVPTSKGQALADEYGIKFFETSAKTNMNVEEVFFSIAR 742
>TC77240 homologue to GP|1370196|emb|CAA98175.1 RAB8D {Lotus japonicus},
complete
Length = 1148
Score = 44.7 bits (104), Expect = 6e-06
Identities = 24/51 (47%), Positives = 32/51 (62%), Gaps = 1/51 (1%)
Frame = +1
Query: 8 LVANKSDL-EPKREVEIEEGGQFAQENGMFYMETSAKTVENINDLFFEIGR 57
LV NK+D+ E KR V +G A E G+ + ETSAKT N+ ++FF I R
Sbjct: 595 LVGNKADMDESKRAVPTSKGQALADEYGIKFFETSAKTNLNVEEVFFSIAR 747
>TC85959 homologue to PIR|S57471|S57471 GTP-binding protein GTP6 - garden
pea, complete
Length = 886
Score = 44.7 bits (104), Expect = 6e-06
Identities = 24/51 (47%), Positives = 32/51 (62%), Gaps = 1/51 (1%)
Frame = +3
Query: 8 LVANKSDL-EPKREVEIEEGGQFAQENGMFYMETSAKTVENINDLFFEIGR 57
LV NK+D+ E KR V +G A E G+ + ETSAKT N+ ++FF I R
Sbjct: 444 LVGNKADMDESKRAVPTSKGQALADEYGIKFFETSAKTNLNVEEVFFSIAR 596
>TC82103 similar to GP|1370144|emb|CAA98178.1 RAB11B {Lotus japonicus},
partial (80%)
Length = 1513
Score = 44.3 bits (103), Expect = 8e-06
Identities = 21/52 (40%), Positives = 32/52 (61%)
Frame = +1
Query: 4 LVMALVANKSDLEPKREVEIEEGGQFAQENGMFYMETSAKTVENINDLFFEI 55
+V+ LV NK DL R V E+ +FA++ G+F+ ETSA T N+ F ++
Sbjct: 532 IVIMLVGNKGDLVDLRMVPTEDAVEFAEDQGLFFSETSALTGVNVEGAFMKL 687
>TC77922 homologue to PIR|T06448|T06448 GTP-binding protein - garden pea,
complete
Length = 1182
Score = 44.3 bits (103), Expect = 8e-06
Identities = 20/52 (38%), Positives = 31/52 (59%)
Frame = +2
Query: 4 LVMALVANKSDLEPKREVEIEEGGQFAQENGMFYMETSAKTVENINDLFFEI 55
+V+ L+ NKSDL V E+G FA+ +++METSA N+ + F E+
Sbjct: 518 IVVMLIGNKSDLRHLVAVPTEDGKSFAERESLYFMETSALEATNVENAFTEV 673
>TC77446 homologue to GP|303750|dbj|BAA02116.1| GTP-binding protein {Pisum
sativum}, complete
Length = 1080
Score = 44.3 bits (103), Expect = 8e-06
Identities = 26/75 (34%), Positives = 36/75 (47%)
Frame = +3
Query: 8 LVANKSDLEPKREVEIEEGGQFAQENGMFYMETSAKTVENINDLFFEIGRGFESPGRSSY 67
LV NKSDL K+ V E FA E G+ +METSAK N+ F + ++ S
Sbjct: 468 LVGNKSDLSDKKVVSSETAKAFADEIGIPFMETSAKNASNVEQAFMAMAAEIKNRMASQP 647
Query: 68 APTKEWASAGLSSMP 82
A + A+ + P
Sbjct: 648 ANSARPATVQIRGQP 692
>TC84871 similar to GP|1370172|emb|CAA98163.1 RAB1X {Lotus japonicus},
partial (87%)
Length = 727
Score = 43.1 bits (100), Expect = 2e-05
Identities = 24/55 (43%), Positives = 32/55 (57%)
Frame = +3
Query: 1 SQILVMALVANKSDLEPKREVEIEEGGQFAQENGMFYMETSAKTVENINDLFFEI 55
+Q + LV NK D E +R V EEG A+E G + E SAKT EN++ F E+
Sbjct: 447 NQDCMKMLVGNKVDRESERTVSTEEGLALAKEFGCSFFECSAKTRENVDKCFEEL 611
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.311 0.129 0.361
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 2,291,639
Number of Sequences: 36976
Number of extensions: 22824
Number of successful extensions: 122
Number of sequences better than 10.0: 65
Number of HSP's better than 10.0 without gapping: 122
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 122
length of query: 88
length of database: 9,014,727
effective HSP length: 64
effective length of query: 24
effective length of database: 6,648,263
effective search space: 159558312
effective search space used: 159558312
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.8 bits)
S2: 51 (24.3 bits)
Lotus: description of TM0033b.5


