

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
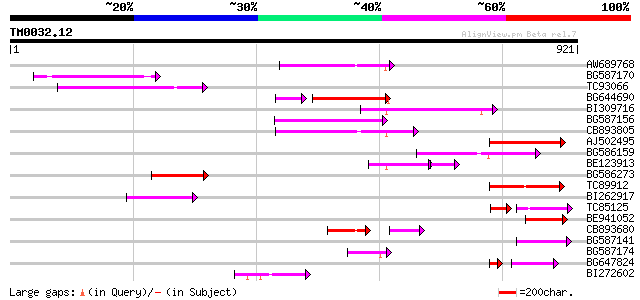
Query= TM0032.12
(921 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
AW689768 weakly similar to GP|10177485|d polyprotein {Arabidopsi... 168 1e-41
BG587170 similar to PIR|F86470|F8 probable retroelement polyprot... 137 1e-32
TC93066 weakly similar to GP|19920130|gb|AAM08562.1 Putative ret... 132 8e-31
BG644690 weakly similar to GP|18542179|gb putative pol protein {... 108 2e-30
BI309716 weakly similar to PIR|G96722|G9 hypothetical protein F2... 127 1e-29
BG587156 similar to PIR|G85055|G8 probable polyprotein [imported... 127 2e-29
CB893805 similar to GP|10177935|d copia-type polyprotein {Arabid... 124 1e-28
AJ502495 weakly similar to GP|18071369|g putative gag-pol polypr... 119 5e-27
BG586159 weakly similar to PIR|T47841|T4 hypothetical protein T2... 108 1e-23
BE123913 weakly similar to GP|22093573|d polyprotein {Oryza sati... 70 2e-18
BG586273 weakly similar to PIR|F86470|F8 probable retroelement p... 90 3e-18
TC89912 weakly similar to PIR|B84512|B84512 probable retroelemen... 87 3e-17
BI262917 weakly similar to GP|19920130|g Putative retroelement {... 82 1e-15
TC85125 weakly similar to SP|P10978|POLX_TOBAC Retrovirus-relate... 63 2e-14
BE941052 weakly similar to PIR|B85188|B85 retrotransposon like p... 74 3e-13
CB893680 weakly similar to GP|1167523|db ORF(AA 1-1338) {Nicotia... 60 1e-12
BG587141 similar to PIR|H86461|H86 hypothetical protein AAF32440... 70 4e-12
BG587174 similar to PIR|A47759|A4775 retrovirus-related reverse ... 65 9e-11
BG647824 weakly similar to PIR|G96722|G96 hypothetical protein F... 55 1e-10
BI272602 GP|11602812|db avenaIII {Gallus gallus}, partial (3%) 62 8e-10
>AW689768 weakly similar to GP|10177485|d polyprotein {Arabidopsis thaliana},
partial (9%)
Length = 675
Score = 168 bits (425), Expect = 1e-41
Identities = 90/197 (45%), Positives = 118/197 (59%), Gaps = 11/197 (5%)
Frame = +1
Query: 439 LSDPNWKFAMQPEFNALIRNNTWELVPRPCDVNVIRYM*IFRHKKQSNGLFEHYKARLVG 498
LSDP W AM+ E+ ALI N TW+LVP P I ++R K+ +G +KARLV
Sbjct: 82 LSDPRWLQAMKTEYKALIDNKTWDLVPLPPHKKAIGCKWVYRVKENPDGSVNKFKARLVA 261
Query: 499 DGWSQIAGVDCDETFNLVVKPATIRTVLSIALSRSWPIH*LDVHNAFLHGDLHETVYMHQ 558
G+SQ G D ETF+ V+KP TIR +L+IA++ W I +D++NAFL+G L E VYM Q
Sbjct: 262 KGFSQTLGCDYTETFSPVIKPVTIRLILTIAITYKWEIQQIDINNAFLNGFLQEEVYMSQ 441
Query: 559 PLGFRDSQHPDYVCRRKKSLYGLKQAPRAWYQRFADYVSSIGFQHSSSDHS--------- 609
P GF ++ + VC+ KSLYGLKQAPRAWY+ GF S D S
Sbjct: 442 PQGF-EAANKSLVCKLNKSLYGLKQAPRAWYEXLTSAQIQFGFTKSRCDPSLLIYNQNGA 618
Query: 610 --YILLYVDDIILVASS 624
Y+ +YVDDI++ SS
Sbjct: 619 CIYLXIYVDDILITGSS 669
>BG587170 similar to PIR|F86470|F8 probable retroelement polyprotein
[imported] - Arabidopsis thaliana, partial (13%)
Length = 718
Score = 137 bits (346), Expect = 1e-32
Identities = 68/206 (33%), Positives = 114/206 (55%)
Frame = -3
Query: 39 VWHNRLGHPTSSALNHLRNNKLIYCDPSRSSTVCDSCVLGKHVRLPFSSSETITLRSFDI 98
+WH RLGHP ALN + + + C++C+LGKH + F + T+ FD+
Sbjct: 584 LWHARLGHPHGRALNLMLPGVVF------ENKNCEACILGKHCKNVFPRTSTVYENCFDL 423
Query: 99 LHSDLWTSPILSTAGHRYYVLVLDDHTDFLWMFPISKKSQVYDIFTTLATLIKTQFSANI 158
+++DLWT+P LS H+Y+V +D+ + + W+ I K +V D F + + A I
Sbjct: 422 IYTDLWTAPSLSRDNHKYFVTFIDEKSKYTWLTLIPSKDRVIDAFKNFQAYVTNHYHAKI 243
Query: 159 KCLQCDNGREYDNDSFRRYCHANGLIFRFSCPHTSSQNGKSERKIRTINNMIRTLLAHSS 218
K L+ DNG EY + +F+ + +G++ + SCP+T QNG ++RK + + + R+L+ ++
Sbjct: 242 KILRSDNGGEYTSYAFKSHLDHHGILHQTSCPYTPQQNGVAKRKNKHLMEVARSLMFQAN 63
Query: 219 VPPSFWHHALQMATYLLNILPRKTLR 244
V A YL+N +P K LR
Sbjct: 62 V---------STACYLINWIPTKVLR 12
>TC93066 weakly similar to GP|19920130|gb|AAM08562.1 Putative retroelement
{Oryza sativa} [Oryza sativa (japonica cultivar-group)],
partial (10%)
Length = 823
Score = 132 bits (331), Expect = 8e-31
Identities = 79/248 (31%), Positives = 124/248 (49%), Gaps = 5/248 (2%)
Frame = +1
Query: 78 GKHVRLPFSSSETITLRSFDILHSDLW-TSPILSTAGHRYYVLVLDDHTDFLWMFPISKK 136
G ++ FS++ T D +HSDLW S + S G RY + ++DD +W++ + K
Sbjct: 43 GNRKKVSFSTATHRTKGILDYIHSDLWGPSKVTSYGGRRYMMTIIDDFPRKVWVYFLRYK 222
Query: 137 SQVYDIFTTLATLIKTQFSANIKCLQCDNGREYDNDSFRRYCHANGLIFRFSCPHTSSQN 196
++ + F L++TQ N+K L DN E+ + F +C +G+ + P QN
Sbjct: 223 NETFPTFKKWRILVETQTGKNVKKLITDN*LEFCSSDFNEFCTNHGIARHKTIPRNPQQN 402
Query: 197 GKSERKIRTINNMIRTLLAHSSV--PPSFWHHALQMATYLLNILPRKTLRNDSPTQRLYH 254
G +ER IRT+ R +L+++ + W A A +L+N P L P
Sbjct: 403 GVAERMIRTLLERARCMLSNAGL*N*RDLWVEAASTACHLVNRSPHSALDFKVPEDIWSG 582
Query: 255 RDPSYSHLRVFGCLCFPFFPSATINKLQPRSSPCVFLGYPMNHRGYK--CYDLSNRKLII 312
YS+LR+FGC P + KL PR+ C+FL Y +GY+ C D ++KLI+
Sbjct: 583 NLVDYSNLRIFGC---PAYALVNDGKLAPRAGECIFLSYASESKGYRLWCSDPKSQKLIL 753
Query: 313 SRHVIFDE 320
SR V F+E
Sbjct: 754 SRDVTFNE 777
>BG644690 weakly similar to GP|18542179|gb putative pol protein {Zea mays},
partial (22%)
Length = 629
Score = 108 bits (271), Expect(2) = 2e-30
Identities = 55/137 (40%), Positives = 86/137 (62%), Gaps = 11/137 (8%)
Frame = -2
Query: 493 KARLVGDGWSQIAGVDCDETFNLVVKPATIRTVLSIALSRSWPIH*LDVHNAFLHGDLHE 552
K++LV G++Q G+D DE F+ V + IR +++ A + ++ +DV +AF++GDL E
Sbjct: 412 KSKLVVQGYNQKEGIDYDEAFSPVARMEAIRILIAFAAFMGFKLYQMDVKSAFINGDLKE 233
Query: 553 TVYMHQPLGFRDSQHPDYVCRRKKSLYGLKQAPRAWYQRFADYVSSIGFQHSSSDHSYIL 612
V++ QP GF D++ P++V R K+LYGLKQAPRAWY+R + ++ GF+ D++ L
Sbjct: 232 EVFVKQPPGFEDAEVPNHVFRLNKTLYGLKQAPRAWYERLSKFLLKNGFKRGKIDNTLFL 53
Query: 613 L-----------YVDDI 618
L YVDDI
Sbjct: 52 LKRE*ELLIIQVYVDDI 2
Score = 42.7 bits (99), Expect(2) = 2e-30
Identities = 22/51 (43%), Positives = 29/51 (56%)
Frame = -3
Query: 432 PCNPKQALSDPNWKFAMQPEFNALIRNNTWELVPRPCDVNVIRYM*IFRHK 482
P N K+AL D +W +MQ E + R+ W LVPRP VI +FR+K
Sbjct: 585 PKNVKEALRDADWINSMQEELHQFERSKVWYLVPRPEGKTVIGTRWVFRNK 433
>BI309716 weakly similar to PIR|G96722|G9 hypothetical protein F20P5.25
[imported] - Arabidopsis thaliana, partial (10%)
Length = 744
Score = 127 bits (320), Expect = 1e-29
Identities = 86/240 (35%), Positives = 116/240 (47%), Gaps = 18/240 (7%)
Frame = +2
Query: 571 VCRRKKSLYGLKQAPRAWYQRFADYVSSIGFQHSSSDHSY-----------ILLYVDDII 619
VC +KS+YGLKQA R WY + ++ + S G+ SSSD S +L+YVDDI+
Sbjct: 20 VCELQKSIYGLKQASRQWYSKLSESLISFGYLQSSSDFSLFTKFKDSSFTTLLVYVDDIV 199
Query: 620 LVASSHDLRKSFMALLAFEFAMKDLGPLSYFLGIAVTRYVGGLFLSQSTYASEIIARAGM 679
L + + L F +KDLG L YFLG+ V R G+ L+Q Y E++ +G
Sbjct: 200 LAGNDISEIQHVKCFLIDRFKIKDLGSLRYFLGLEVARSKQGILLNQRKYTLELLEDSGN 379
Query: 680 ASCNPSATPVDTK*KLSSSSGTPCEDVTLYQSLAGALQYLTFTRPNISYVVQQVCLHMHA 739
+ + TP D KL +S D T Y+ L G L YLT TRP+IS+ VQQ+ +
Sbjct: 380 LAVKSTLTPYDISLKLHNSDSPLYNDETQYRRLIGKLIYLTTTRPDISFAVQQLSQFVSK 559
Query: 740 PHTEHMLAMFVAL*LTVFTCIPPLL----RNLFLIRMLTGGMS---CTRRSTSGYCVFLG 792
P H A L L NL L + TR+S +GY VFLG
Sbjct: 560 PQQVHYQAAIRVLQYLKTAPAKGLFYSATSNLKLSSFADSDWATCPTTRKSVTGYWVFLG 739
>BG587156 similar to PIR|G85055|G8 probable polyprotein [imported] -
Arabidopsis thaliana, partial (17%)
Length = 618
Score = 127 bits (319), Expect = 2e-29
Identities = 70/183 (38%), Positives = 103/183 (56%)
Frame = -1
Query: 431 LPCNPKQALSDPNWKFAMQPEFNALIRNNTWELVPRPCDVNVIRYM*IFRHKKQSNGLFE 490
+P + ++A+ D WK ++ E A+I+N+TW P + IF K +++G E
Sbjct: 561 IPRSYEEAMEDKEWKESVGAEAGAMIKNDTWYESELPKGKKAVSSRWIFTIKYKADGSIE 382
Query: 491 HYKARLVGDGWSQIAGVDCDETFNLVVKPATIRTVLSIALSRSWPIH*LDVHNAFLHGDL 550
K RLV G++ G D ETF V K TIR VLS+A++ W + +DV NAFL G+L
Sbjct: 381 RKKTRLVARGFTLTYGEDYIETFAPVAKLHTIRIVLSLAVNLGWGLWQMDVKNAFLQGEL 202
Query: 551 HETVYMHQPLGFRDSQHPDYVCRRKKSLYGLKQAPRAWYQRFADYVSSIGFQHSSSDHSY 610
+ VYM+ P G V R KK++YGLKQ+PRAWY + + ++ GF+ S DH+
Sbjct: 201 EDEVYMYPPPGLEHLVKRGNVLRLKKAIYGLKQSPRAWYNKLSTTLNGRGFRKSELDHTL 22
Query: 611 ILL 613
L
Sbjct: 21 FTL 13
>CB893805 similar to GP|10177935|d copia-type polyprotein {Arabidopsis
thaliana}, partial (14%)
Length = 778
Score = 124 bits (312), Expect = 1e-28
Identities = 83/248 (33%), Positives = 123/248 (49%), Gaps = 15/248 (6%)
Frame = +3
Query: 432 PCNPKQALSDPNWKFAMQPEFNALIRNNTWELVPRPCDVNVIRYM*IFRHKKQSNGLFEH 491
P ++A+ W+ +M E A RNNTWEL I IF+ K NG E
Sbjct: 42 PTTFEEAVKSEKWRASMNNEMEATERNNTWELTDLRSGAKTIGLKWIFKTKLNENGEIEK 221
Query: 492 YKARLVGDGWSQIAGVDCDETFNLVVKPATIRTVLSIALSRSWP---IH*LDVHNAFLHG 548
YKARLV G+SQ GVD E F V + TIR V+++A I *+ ++ +
Sbjct: 222 YKARLVAKGYSQQYGVDYTEVFAPVARWDTIRMVIALAAQIKRDGVCIS*M*KAHSCMEN 401
Query: 549 DLHETVYMHQPLGFRDSQHPDYVCRRKKSLYGLKQAPRAWYQRFADYVSSIGFQHSSSDH 608
+ + + ++ + R R K++LYGLKQAPRAWY R Y + GF+ +H
Sbjct: 402 *MRKFLLINHRVM*RRV----IS*RVKRALYGLKQAPRAWYSRIEAYFTKEGFEKCPYEH 569
Query: 609 SY------------ILLYVDDIILVASSHDLRKSFMALLAFEFAMKDLGPLSYFLGIAVT 656
+ I LYVDD+I + + ++ + F + EF M DLG + YFLG+ VT
Sbjct: 570 TLFVKLSEGGKILIISLYVDDLIFIGNDENMFEEFKKSMKKEFNMSDLGKMHYFLGVEVT 749
Query: 657 RYVGGLFL 664
+ G+++
Sbjct: 750 QNEKGIYI 773
>AJ502495 weakly similar to GP|18071369|g putative gag-pol polyprotein {Oryza
sativa}, partial (9%)
Length = 542
Score = 119 bits (298), Expect = 5e-27
Identities = 57/124 (45%), Positives = 82/124 (65%)
Frame = +2
Query: 780 TRRSTSGYCVFLGDNLISWSSKRQPTLSHSSAEAEYRGVANVVSESCWIRNLLLELHFPL 839
TR+STSGY LG ISWSSK+QP ++ S+AEAEY + +++ W+R +L +H
Sbjct: 26 TRKSTSGYAFHLGTGAISWSSKKQPVVAFSTAEAEYIASTSCATQTVWLRRILEVMHHEQ 205
Query: 840 SQETLVHCDNVSSIYLSGNPVHHQRTKHIEMDIHFVREKVARGQARILHVPSRHQIADIF 899
+ T ++CDN S+I LS NPV H R+KHI++ H +RE +A + I + P+ +IADIF
Sbjct: 206 NTPTKIYCDNKSAIALSKNPVFHGRSKHIDIQFHKIRELIAEKEVVIEYCPTEEKIADIF 385
Query: 900 TKGL 903
TK L
Sbjct: 386 TKPL 397
>BG586159 weakly similar to PIR|T47841|T4 hypothetical protein T2O9.150 -
Arabidopsis thaliana, partial (11%)
Length = 732
Score = 108 bits (269), Expect = 1e-23
Identities = 69/215 (32%), Positives = 104/215 (48%), Gaps = 13/215 (6%)
Frame = +1
Query: 661 GLFLSQSTYASEIIARAGMASCNPSATPVDTK*KLSSSSGTPCEDVTLYQSLAGALQYLT 720
G+++ Q Y ++++ R GM N S P+ + KL D T Y+ + G L YL
Sbjct: 25 GIYICQRKYVTDLLERFGMEKSNLSRNPIAPRCKLIKDENGVKVDATKYKQIVGCLMYLA 204
Query: 721 FTRPNISYVVQQVCLHMHAPHTEHMLAMFVAL*LTVFTCIPPLLRNLFLIRMLTG----- 775
TRP++ YV+ + M+ P HM A+ L T NL ++ G
Sbjct: 205 ATRPDLMYVLSLISRFMNCPTELHMHAVKRVLRYLNGTI------NLGIMYKRNGSEKLE 366
Query: 776 --------GMSCTRRSTSGYCVFLGDNLISWSSKRQPTLSHSSAEAEYRGVANVVSESCW 827
G R+STSGY L +SWSSK+QP ++ S+ +AE+ A +S W
Sbjct: 367 AYTDSDYAGDLDDRKSTSGYVFMLSSGAVSWSSKKQPVVTLSTTKAEFIAAAFCACQSVW 546
Query: 828 IRNLLLELHFPLSQETLVHCDNVSSIYLSGNPVHH 862
+R +L +L + S ++CDN S+I LS NPV H
Sbjct: 547 MRRVLEKLGYTQSGSITMYCDNNSTIKLSKNPVLH 651
>BE123913 weakly similar to GP|22093573|d polyprotein {Oryza sativa (japonica
cultivar-group)}, partial (8%)
Length = 503
Score = 70.1 bits (170), Expect(2) = 2e-18
Identities = 40/118 (33%), Positives = 55/118 (45%), Gaps = 12/118 (10%)
Frame = +1
Query: 583 QAPRAWYQRFADYVSSIGFQHSSSDHSY------------ILLYVDDIILVASSHDLRKS 630
Q+PR W+ RF V G+ +DH+ +++YVDDI L K
Sbjct: 1 QSPRDWFDRFT*VVKKFGYIQCQTDHAMFIKHSSTVKKAILIVYVDDIFLTGDHGK*IKR 180
Query: 631 FMALLAFEFAMKDLGPLSYFLGIAVTRYVGGLFLSQSTYASEIIARAGMASCNPSATP 688
LLA EF +KDLG L YFLG+ V R+ G +SQ Y +++ M C P
Sbjct: 181 LKNLLAEEFEIKDLGNLKYFLGMEVARWKKGSSISQRKYVLDLLKETRMIGCKTIRDP 354
Score = 41.2 bits (95), Expect(2) = 2e-18
Identities = 22/47 (46%), Positives = 27/47 (56%)
Frame = +2
Query: 684 PSATPVDTK*KLSSSSGTPCEDVTLYQSLAGALQYLTFTRPNISYVV 730
PS TP+D KL + D YQ L G L YL+ TRP+IS+VV
Sbjct: 341 PSETPMDATVKLGTLDNGTLVDKGRYQRLVGKLIYLSHTRPDISFVV 481
>BG586273 weakly similar to PIR|F86470|F8 probable retroelement polyprotein
[imported] - Arabidopsis thaliana, partial (7%)
Length = 705
Score = 90.1 bits (222), Expect = 3e-18
Identities = 40/92 (43%), Positives = 59/92 (63%)
Frame = -2
Query: 231 ATYLLNILPRKTLRNDSPTQRLYHRDPSYSHLRVFGCLCFPFFPSATINKLQPRSSPCVF 290
A YL+N +P + L++ +P + L R PS +++RVFGCLC+ P NKL+ RS +F
Sbjct: 704 ACYLINRIPTRVLKDQAPFEVLNQRKPSLTYMRVFGCLCYVLVPGELRNKLEARSRKAMF 525
Query: 291 LGYPMNHRGYKCYDLSNRKLIISRHVIFDESR 322
+GY +GYKCYD R++++SR V F E R
Sbjct: 524 IGYSTTQKGYKCYDPEARRVLVSRDVKFIEER 429
>TC89912 weakly similar to PIR|B84512|B84512 probable retroelement pol
polyprotein [imported] - Arabidopsis thaliana, partial
(10%)
Length = 814
Score = 87.0 bits (214), Expect = 3e-17
Identities = 42/123 (34%), Positives = 77/123 (62%), Gaps = 1/123 (0%)
Frame = +1
Query: 780 TRRSTSGYCVFLGDNLISWSSKRQPTLSHSSAEAEYRGVANVVSESCWIRNLLLELHFPL 839
TR+S SG+ L ISW + +Q ++ S+ +AEY V ++ W++ ++ EL +
Sbjct: 130 TRKSLSGFVFTLYGTTISWKANQQSVVTLSTTQAEYIAFVEGVKDAIWLKGMIGELG--I 303
Query: 840 SQETL-VHCDNVSSIYLSGNPVHHQRTKHIEMDIHFVREKVARGQARILHVPSRHQIADI 898
+QE + +HCD+ S+I+L+ + V+H+RTKHI++ +HF+R+ + + + + S AD+
Sbjct: 304 TQEYVKIHCDSQSAIHLANHQVYHERTKHIDIRLHFIRDMIESKEIVVEKMASEENPADV 483
Query: 899 FTK 901
FTK
Sbjct: 484 FTK 492
>BI262917 weakly similar to GP|19920130|g Putative retroelement {Oryza
sativa} [Oryza sativa (japonica cultivar-group)],
partial (8%)
Length = 426
Score = 81.6 bits (200), Expect = 1e-15
Identities = 41/114 (35%), Positives = 60/114 (51%)
Frame = +1
Query: 191 HTSSQNGKSERKIRTINNMIRTLLAHSSVPPSFWHHALQMATYLLNILPRKTLRNDSPTQ 250
HT QNG +ER RT+ R +L + + SFW A++ A Y++N P + +P +
Sbjct: 82 HTPQQNGVAERMNRTLLERTRAMLKTAGMAKSFWAEAVKTACYVINRSPSTVIDLKTPME 261
Query: 251 RLYHRDPSYSHLRVFGCLCFPFFPSATINKLQPRSSPCVFLGYPMNHRGYKCYD 304
+ YS L VFGC + + S KL P+S C+FLGY N +GY +D
Sbjct: 262 MWKGKPVDYSSLHVFGCPVYVMYNSQERTKLDPKSRKCIFLGYADNVKGYXLWD 423
>TC85125 weakly similar to SP|P10978|POLX_TOBAC Retrovirus-related Pol
polyprotein from transposon TNT 1-94 [Contains: Protease
(EC 3.4.23.-);, partial (7%)
Length = 705
Score = 62.8 bits (151), Expect(2) = 2e-14
Identities = 34/91 (37%), Positives = 51/91 (55%)
Frame = +3
Query: 824 ESCWIRNLLLELHFPLSQETLVHCDNVSSIYLSGNPVHHQRTKHIEMDIHFVREKVARGQ 883
E+ W++ L+ EL Q T V+CD+ S+++++ NP H RTKHI + HFVRE V G
Sbjct: 249 EAIWMQRLMEELGHKQEQIT-VYCDSQSALHIARNPAFHSRTKHIGIQYHFVREVVEEGS 425
Query: 884 ARILHVPSRHQIADIFTKGLPRVLFDDFRSS 914
+ + + +AD TK + F RSS
Sbjct: 426 VDMQKIHTNDNLADAMTKSINTDKFIWCRSS 518
Score = 35.4 bits (80), Expect(2) = 2e-14
Identities = 16/35 (45%), Positives = 22/35 (62%)
Frame = +1
Query: 781 RRSTSGYCVFLGDNLISWSSKRQPTLSHSSAEAEY 815
R+ST+GY L +SW SK Q ++ S+ EAEY
Sbjct: 118 RKSTTGYVFTLAGGAVSWLSKLQTVVALSTTEAEY 222
>BE941052 weakly similar to PIR|B85188|B85 retrotransposon like protein
[imported] - Arabidopsis thaliana, partial (4%)
Length = 480
Score = 73.6 bits (179), Expect = 3e-13
Identities = 33/68 (48%), Positives = 48/68 (70%)
Frame = +2
Query: 838 PLSQETLVHCDNVSSIYLSGNPVHHQRTKHIEMDIHFVREKVARGQARILHVPSRHQIAD 897
P+SQ L+ CD +S+ YL+ NPV+H R KHI +DIHFVR+ V +G+ ++ HV + Q+AD
Sbjct: 2 PISQSGLLRCDYLSATYLTHNPVYHSRMKHISIDIHFVRDLVQQGKLKVQHVCTVDQLAD 181
Query: 898 IFTKGLPR 905
TK L +
Sbjct: 182 CLTKPLSK 205
>CB893680 weakly similar to GP|1167523|db ORF(AA 1-1338) {Nicotiana tabacum},
partial (7%)
Length = 780
Score = 60.5 bits (145), Expect(2) = 1e-12
Identities = 36/71 (50%), Positives = 44/71 (61%)
Frame = -2
Query: 516 VVKPATIRTVLSIALSRSWPIH*LDVHNAFLHGDLHETVYMHQPLGFRDSQHPDYVCRRK 575
+VK TI +LSI + + *LDV AFL GDL E +YMHQP GF + V + K
Sbjct: 545 IVKLNTIMFLLSIVAIENLYLE*LDVKTAFLRGDLVEDIYMHQPEGF-S*EVGKMVGKLK 369
Query: 576 KSLYGLKQAPR 586
KS+YGLKQ PR
Sbjct: 368 KSMYGLKQGPR 336
Score = 31.2 bits (69), Expect(2) = 1e-12
Identities = 19/59 (32%), Positives = 31/59 (52%), Gaps = 2/59 (3%)
Frame = -3
Query: 618 IILVASSHDLRKSFMALLAFEFAMKDLGPLSYFLG--IAVTRYVGGLFLSQSTYASEII 674
+++V S+ D K+ + E MKDLGP +G I + + G L LSQ Y + ++
Sbjct: 271 LLVVGSNIDEIKNLKTRFSKEIDMKDLGPAKKIIGMQIMIDKQKGVL*LSQVEYITRVL 95
>BG587141 similar to PIR|H86461|H86 hypothetical protein AAF32440.1
[imported] - Arabidopsis thaliana, partial (20%)
Length = 731
Score = 70.1 bits (170), Expect = 4e-12
Identities = 35/89 (39%), Positives = 51/89 (56%)
Frame = +3
Query: 824 ESCWIRNLLLELHFPLSQETLVHCDNVSSIYLSGNPVHHQRTKHIEMDIHFVREKVARGQ 883
++ W+++LL E+ + +E ++ DN S I L+ NPV H R HI HF+RE V GQ
Sbjct: 135 QAMWLQDLLSEVTWEPCEEVVIRIDNQSVIALTRNPVFHGRGNHIHKRYHFIRECVENGQ 314
Query: 884 ARILHVPSRHQIADIFTKGLPRVLFDDFR 912
+ HVP A I TK L R++F + R
Sbjct: 315 VEVEHVPGEKHRAYI*TKALGRIIFREIR 401
>BG587174 similar to PIR|A47759|A4775 retrovirus-related reverse
transcriptase homolog - rape retrotransposon copia-like
(fragment), partial (84%)
Length = 249
Score = 65.5 bits (158), Expect = 9e-11
Identities = 35/83 (42%), Positives = 48/83 (57%), Gaps = 12/83 (14%)
Frame = -1
Query: 549 DLHETVYMHQPLGFRDSQHPDYVCRRKKSLYGLKQAPRAWYQRFADYVSSIG-------- 600
+L E +YM QP GF D+VC+ +KSLYGLKQ+PR WY+RF Y SS
Sbjct: 249 ELEEKIYMTQPEGFLFPGKEDHVCKLRKSLYGLKQSPRQWYKRFDSYRSSWATTGVLMTV 70
Query: 601 ----FQHSSSDHSYILLYVDDII 619
+ S + Y++LYVDD++
Sbjct: 69 VST*TR*RMSRYIYLVLYVDDML 1
>BG647824 weakly similar to PIR|G96722|G96 hypothetical protein F20P5.25
[imported] - Arabidopsis thaliana, partial (5%)
Length = 721
Score = 54.7 bits (130), Expect(2) = 1e-10
Identities = 28/78 (35%), Positives = 45/78 (56%), Gaps = 1/78 (1%)
Frame = -3
Query: 815 YRGVANVVSESCWIRNLLLELHFPLSQETLVHCDNVSSI-YLSGNPVHHQRTKHIEMDIH 873
YR + + + E W+ LL +L F + +++CDN S+ +++ N +RTKHIE+D H
Sbjct: 374 YRSI*STICEIKWLTYLLNDLKFTFIKPAMLYCDNQSAARHIAANSSFLERTKHIELDCH 195
Query: 874 FVREKVARGQARILHVPS 891
VR K+ ILH+ S
Sbjct: 194 IVRVKLQLKLFHILHILS 141
Score = 30.4 bits (67), Expect(2) = 1e-10
Identities = 12/21 (57%), Positives = 16/21 (76%)
Frame = -2
Query: 780 TRRSTSGYCVFLGDNLISWSS 800
T +S S +C+FLGD+LI W S
Sbjct: 465 T*KSISYFCIFLGDSLICWKS 403
>BI272602 GP|11602812|db avenaIII {Gallus gallus}, partial (3%)
Length = 488
Score = 62.4 bits (150), Expect = 8e-10
Identities = 49/142 (34%), Positives = 65/142 (45%), Gaps = 18/142 (12%)
Frame = +1
Query: 365 PSSPTDSTMPSPAPTSSPTS----------SAPLPLPPVPPTP-PTRTMTTH-------S 406
PSSP + + SSP S ++P P PP PPT P TT+ S
Sbjct: 25 PSSPNSNEVAQNIQISSPASPTSNELANITTSPTPPPPPPPTARPHNNHTTNVGGIITRS 204
Query: 407 MHGISKPKKPFSLSVSIDDPSISPLPCNPKQALSDPNWKFAMQPEFNALIRNNTWELVPR 466
+ I K K +L V P P N QAL D +W+ MQ EF+AL NNTW+LV R
Sbjct: 205 QNNIVKTIKKLNLHVR---PLCPIEPSNITQALRDHDWRSIMQDEFDALQNNNTWDLVSR 375
Query: 467 PCDVNVIRYM*IFRHKKQSNGL 488
N++ F ++ +S L
Sbjct: 376 SFAQNLVGCKLGFSYQTKSGWL 441
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.325 0.138 0.439
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 35,918,132
Number of Sequences: 36976
Number of extensions: 677338
Number of successful extensions: 13369
Number of sequences better than 10.0: 622
Number of HSP's better than 10.0 without gapping: 6083
Number of HSP's successfully gapped in prelim test: 308
Number of HSP's that attempted gapping in prelim test: 1255
Number of HSP's gapped (non-prelim): 9141
length of query: 921
length of database: 9,014,727
effective HSP length: 105
effective length of query: 816
effective length of database: 5,132,247
effective search space: 4187913552
effective search space used: 4187913552
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 63 (28.9 bits)
Lotus: description of TM0032.12


