

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0029b.1
(1377 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
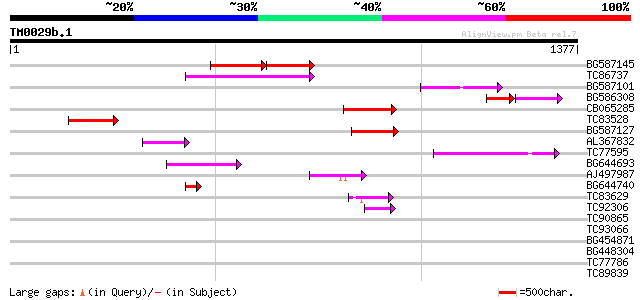
Score E
Sequences producing significant alignments: (bits) Value
BG587145 similar to PIR|H86337|H8 protein F5M15.26 [imported] - ... 171 2e-70
TC86737 weakly similar to GP|6683624|dbj|BAA89272.1 Pol {Alterna... 150 3e-36
BG587101 similar to GP|6691191|gb F7F22.15 {Arabidopsis thaliana... 120 4e-27
BG586308 weakly similar to PIR|F84528|F8 probable retroelement p... 77 2e-26
CB065285 weakly similar to PIR|A84500|A84 probable retroelement ... 100 5e-21
TC83528 96 1e-19
BG587127 weakly similar to PIR|H84506|H84 probable retroelement ... 92 1e-18
AL367832 weakly similar to GP|14091845|gb Putative retroelement ... 84 3e-16
TC77595 weakly similar to PIR|T18350|T18350 probable pol polypro... 84 5e-16
BG644693 weakly similar to GP|18767374|g Putative 22 kDa kafirin... 81 2e-15
AJ497987 weakly similar to GP|9927273|dbj Similar to Arabidopsis... 67 4e-11
BG644740 similar to PIR|A84460|A84 probable retroelement pol pol... 48 2e-05
TC83629 weakly similar to GP|6692261|gb|AAF24611.1| putative RNa... 46 1e-04
TC92306 45 3e-04
TC90865 homologue to GP|9049326|dbj|BAA99336.1 orf129b {Beta vul... 42 0.002
TC93066 weakly similar to GP|19920130|gb|AAM08562.1 Putative ret... 38 0.031
BG454871 weakly similar to GP|10140673|g putative gag-pol polypr... 36 0.090
BG448304 34 0.34
TC77786 34 0.34
TC89839 homologue to SP|P23466|CYAA_SACKL Adenylate cyclase (EC ... 33 1.00
>BG587145 similar to PIR|H86337|H8 protein F5M15.26 [imported] - Arabidopsis
thaliana, partial (13%)
Length = 763
Score = 171 bits (433), Expect(2) = 2e-70
Identities = 78/136 (57%), Positives = 106/136 (77%)
Frame = +2
Query: 488 MHPADEDKTAFMTARVNYCYRTMPFGLKNAGATYQRLMDRVFARQVGRNMEVYVDDMIVK 547
MHP D +KTAF+T R YCY+ MPFGLKNAG+TYQRL++R+FA ++G MEVY+DDM+VK
Sbjct: 2 MHPDDLEKTAFITDRGTYCYKVMPFGLKNAGSTYQRLVNRMFADKLGNTMEVYIDDMLVK 181
Query: 548 SVRGLDHHQDLEEAFGEIRKHSMRLNPEKCSFGVQGGKFLGFMITSRGIEINPEKCKAIQ 607
S+R DH L+E F + ++ M+LNP KC+FGV G+FLG+++T +GIE+NP++ AI
Sbjct: 182 SLRATDHLNHLKE*FKTLDEYIMKLNPAKCTFGVTSGEFLGYIVTQQGIEVNPKQITAIL 361
Query: 608 QMKSPSNVKEVQRLTG 623
+ SP N +EVQRLTG
Sbjct: 362 DLPSPKNSREVQRLTG 409
Score = 114 bits (286), Expect(2) = 2e-70
Identities = 55/118 (46%), Positives = 81/118 (68%)
Frame = +3
Query: 623 GRIAALSRFLPKSGDRSFPFFKCLRKNVVFEWTAECEEAFVRLKELLSSPPILSKPIQGH 682
GRIAAL+RF+ +S D+ PF+K L N F W +CEEAF +LK+ L++PP+LSKP G
Sbjct: 408 GRIAALNRFISRSTDKCLPFYKLLCGNKRFVWDEKCEEAFEQLKQYLTTPPVLSKPEAGD 587
Query: 683 PLHLYFAVSDSALSSVMLQEIDGEHRIVYFVSHTLQGAEVRYQKIEKAALAVLVTARR 740
L LY A+S +A+SSV+++E GE + +++ S + E RY +EK A AV+ +AR+
Sbjct: 588 TLSLYIAISSTAVSSVLIREDRGEQKPIFYTSKRMTDPETRYPTLEKMAFAVITSARK 761
>TC86737 weakly similar to GP|6683624|dbj|BAA89272.1 Pol {Alternaria
alternata}, partial (21%)
Length = 1540
Score = 150 bits (379), Expect = 3e-36
Identities = 98/317 (30%), Positives = 151/317 (46%), Gaps = 3/317 (0%)
Frame = +1
Query: 426 ANVVMVKKANGKWRMCVDYTDLNKACPKDSYPLPSIDSLVDGASGNELLSLMDAYSGYHQ 485
A V+ V+K G R CVDY LN KD YPLP I + +G + +D + +H+
Sbjct: 577 APVLFVRKPGGGIRFCVDYRALNAITKKDRYPLPLISETLRRVAGARWFTKLDVVAAFHK 756
Query: 486 IRMHPADEDKTAFMTARVNYCYRTMPFGLKNAGATYQRLMDRVFARQVGRNMEVYVDDMI 545
+R+ D++KTAF T + + PFGL A AT+QR +++ + + Y+DD++
Sbjct: 757 MRIKDEDQEKTAFRTRYGLFEWIVCPFGLTGAPATFQRYINKTLHEFLDDFVTAYIDDVL 936
Query: 546 VKSVRG-LDHHQDLEEAFGEIRKHSMRLNPEKCSFGVQGGKFLGFMITS-RGIEINPEKC 603
+ + DH + + + L+P+KC F V K++GF++T+ +G+ +P K
Sbjct: 937 IYTTGSKKDHEAQVRRVLRRLADAGLSLDPKKCEFSVTTVKYVGFILTAGKGVSCDPLKL 1116
Query: 604 KAIQQMKSPSNVKEVQRLTGRIAALSRFLPKSGDRSFPFFKCLRKNVVFEWTAECEEAFV 663
AI+ P +VK + G F+P + + P + RK+ F W AE E AF
Sbjct: 1117AAIRDWLPPGSVKGARSFLGFCNYYKDFIPGYSEITEPLTRLTRKDFPFRWGAEQEAAFT 1296
Query: 664 RLKELLSSPPILSKPIQGHPLHLYFAVSDSALSSVMLQEI-DGEHRIVYFVSHTLQGAEV 722
+LK L + P+L + S AL V+ QE G V F S L AE
Sbjct: 1297KLKRLFAEEPVLRMFDPEAVTTVETDCSGFALGGVLTQEDGTGAAHPVAFHSQRLSPAEY 1476
Query: 723 RYQKIEKAALAVLVTAR 739
Y +K LAV R
Sbjct: 1477NYPIHDKELLAVWACLR 1527
>BG587101 similar to GP|6691191|gb F7F22.15 {Arabidopsis thaliana}, partial
(10%)
Length = 624
Score = 120 bits (301), Expect = 4e-27
Identities = 62/202 (30%), Positives = 106/202 (51%), Gaps = 3/202 (1%)
Frame = +2
Query: 997 REASHYTLIDGHLYRRGFSTPLLKCVSPEKYEAIMSEVHEGVCASHIGGRSLACKVLRAG 1056
++ Y + LY++ ++CV+ E+ I+ H A H K+ +AG
Sbjct: 5 KDVRRYLWDEPFLYKQCADNIYIRCVAEEEIPGILFHCHGSNYAGHFAVSKTVSKIQQAG 184
Query: 1057 FYWPTLRKDCMDFVKQCKECQVFADLS---KAPPKELVTMSAPWPFAMWGVDLVGPFPTA 1113
F+WPT+ KD F+ +C CQ ++S + P ++ + F +WG+D +GPFP++
Sbjct: 185 FWWPTMFKDAHSFISKCDPCQRQGNIS*RNEMPQNFILEVEV---FDVWGIDFMGPFPSS 355
Query: 1114 RAQMKFILVAVDYFTKWIEAEPLAKITSAKIVNFYWKRIVCRFGIPRAIVSDNGTQFSSS 1173
K+ILVAVDY +KW+EA + +V + I RFG+PR ++SD G+ F +
Sbjct: 356 YNN-KYILVAVDYVSKWVEAIASPTNDATVVVKMFKSVIFPRFGVPRVVISDGGSHFINK 532
Query: 1174 QTREFCKEMGIQMRFASVEHPQ 1195
+ K+ G++ + A+ HPQ
Sbjct: 533 VFEKLLKKNGVRHKVATAYHPQ 598
>BG586308 weakly similar to PIR|F84528|F8 probable retroelement pol polyprotein
[imported] - Arabidopsis thaliana, partial (7%)
Length = 686
Score = 77.4 bits (189), Expect(2) = 2e-26
Identities = 34/70 (48%), Positives = 51/70 (72%)
Frame = -2
Query: 1157 GIPRAIVSDNGTQFSSSQTREFCKEMGIQMRFASVEHPQTNGQVESANRVILRGLRRRLA 1216
G+P IV+DNG+ F S++ REFC+ I++ AS +PQ+NGQ E++N++I+ GL++RL
Sbjct: 682 GLPYEIVTDNGSHFISNKFREFCERWRIRLNTASPRYPQSNGQAEASNKIIIDGLKKRLD 503
Query: 1217 EAKGAWLDEL 1226
KG W DEL
Sbjct: 502 LKKGCWADEL 473
Score = 61.2 bits (147), Expect(2) = 2e-26
Identities = 38/119 (31%), Positives = 64/119 (52%), Gaps = 6/119 (5%)
Frame = -3
Query: 1229 VLWSYNTTEQSTTRETPFRMTYGVDAMLPVEIDNFTW-RTR-PGFEEENQANMAVELDLL 1286
VLWS+ T + T+ TPF M + V+AM P E++ + R+R P E N + L+ +
Sbjct: 465 VLWSHRTNPRGATKSTPFSMAHRVEAMAPAEVNVTSL*RSRMPQNIELNNDRLFNALETI 286
Query: 1287 SETRDEAHIRETAMKQRVAAKFNSRVRVRDMQVGDLVL----KWRSGAPGNKLTPNWEG 1341
E RD+A +R + ++ + +N V+ + +++GD+VL + KL NWEG
Sbjct: 285 EERRDQALLRIQNYQHQIESYYNKTVKSQPLKLGDIVLCKVFENTKELNAGKLGTNWEG 109
>CB065285 weakly similar to PIR|A84500|A84 probable retroelement gag/pol
polyprotein [imported] - Arabidopsis thaliana, partial
(2%)
Length = 592
Score = 100 bits (248), Expect = 5e-21
Identities = 53/128 (41%), Positives = 79/128 (61%)
Frame = -2
Query: 812 DTQWTLFVDGSSNSSGSGAGVALEGPGELVLEQSLKFEFKATNNQAEYEALIAGLKLARE 871
+++W L DG+ N+ G G G + P + + + F+ TNN AEYEA I G++ A +
Sbjct: 543 NSKWGLVFDGAVNAYGKGIGAVIVSPQGHYIPFTARILFECTNNMAEYEACIFGIEEAID 364
Query: 872 VKIRSLLIRTDSQLVENQVKGTFQVKDPNLIKYLERVRYLMTLFQEVVVEYVPRAENQRA 931
++I+ L I DS LV NQ+KG ++ NLI Y + R L+T F +V + ++PR ENQ A
Sbjct: 363 MRIKHLDIYGDSALVINQIKGEWETHHANLIPYRDYARRLLTYFTKVELHHIPRDENQMA 184
Query: 932 DALAKLAS 939
DALA L+S
Sbjct: 183 DALATLSS 160
>TC83528
Length = 555
Score = 95.5 bits (236), Expect = 1e-19
Identities = 52/125 (41%), Positives = 81/125 (64%), Gaps = 4/125 (3%)
Frame = +3
Query: 144 PIVVQLRMNS-FNIRRVLLDQGSSADIIYGDAFDKLGLTDNDLTPYAGTLVGFAGEQVMV 202
P+V++L++N+ F++ RV ++ S DI+Y AF K+ L ++ L P G L G G+ + V
Sbjct: 180 PMVIKLQINNNFSVLRVFVNPMSKVDILYWSAFLKMKLQESMLKPCQGFLKGTFGKGLPV 359
Query: 203 RGYIDLDTIFGEDECARVLKVRYLVLQ---VVASYNVIIGRNTLNRLCAVISTAHLAVKY 259
+GYIDLDT FG+ E + +KVRY V++ V+ YNV++G +L L AV+S A +KY
Sbjct: 360 KGYIDLDTTFGKGENTKTIKVRYFVVESPPSVSIYNVVLGWPSLKDLKAVLSVAEFTIKY 539
Query: 260 PLSSG 264
P+ G
Sbjct: 540 PVGDG 554
>BG587127 weakly similar to PIR|H84506|H84 probable retroelement pol
polyprotein [imported] - Arabidopsis thaliana, partial
(13%)
Length = 415
Score = 92.0 bits (227), Expect = 1e-18
Identities = 48/113 (42%), Positives = 71/113 (62%)
Frame = +3
Query: 831 GVALEGPGELVLEQSLKFEFKATNNQAEYEALIAGLKLAREVKIRSLLIRTDSQLVENQV 890
G+ L P +L+QS + EF A+NN+ YEALIAG++LA +KIR++ DSQLV +Q
Sbjct: 3 GIRLTSPTNEILKQSFRLEFHASNNETNYEALIAGVRLAHGLKIRNIHAYCDSQLVASQF 182
Query: 891 KGTFQVKDPNLIKYLERVRYLMTLFQEVVVEYVPRAENQRADALAKLASTRKP 943
G ++ +D + YL+ V+ L + +PR+EN +ADALA LAS+ P
Sbjct: 183 SGEYEARDELMDTYLKLVQKLAQKLDYFALTRIPRSENVQADALAALASSSDP 341
>AL367832 weakly similar to GP|14091845|gb Putative retroelement {Oryza
sativa}, partial (2%)
Length = 384
Score = 84.3 bits (207), Expect = 3e-16
Identities = 45/118 (38%), Positives = 66/118 (55%), Gaps = 4/118 (3%)
Frame = -1
Query: 323 EETKALKFGD----RTLKIGTRLTEEQETRLTKLLGENLDLFAWSCKDMPGIDPNFICHR 378
EE + + G R +K+G L E + ++ +LL E LD+FA S +DMPG+DP + HR
Sbjct: 354 EEIELINLGTEENKREIKVGAALEEGVKRKIFQLLREYLDIFACSYEDMPGLDPKIVEHR 175
Query: 379 LALNPSVKPVSQLRRRLGGDKGKAVQQEVDKLLAAEFIREVKYPTWLANVVMVKKANG 436
+ P PV RR D ++ EV K + A F+ V+YP W+AN+V V K +G
Sbjct: 174 IPTKPECPPVR*KLRRTHPDMALKIKSEVQKQIDAGFLMTVEYPEWVANIVPVPKKDG 1
>TC77595 weakly similar to PIR|T18350|T18350 probable pol polyprotein - rice
blast fungus gypsy retroelement (fragment), partial (14%)
Length = 1708
Score = 83.6 bits (205), Expect = 5e-16
Identities = 79/315 (25%), Positives = 131/315 (41%), Gaps = 9/315 (2%)
Frame = +2
Query: 1030 IMSEVHEGVCASHIGGRSLACKVLRAGFYWPTLRKDCMDFVKQCKECQVFADLSKAPPKE 1089
++ E H+ A H GR+ +++ F+WP + FV+ C C +A
Sbjct: 179 LVQESHDSTAAGH-PGRNGTLEIVSRKFFWPGQSQTVRRFVRNCDVCGGIHIWRQAKRGF 355
Query: 1090 LVTMSAPWPF-AMWGVDLVGPFPTARAQ-MKFILVAVDYFTKWIEAEPLAKITSAKIVNF 1147
L + P + +D + P R + +++ V VD +K + E + + +
Sbjct: 356 LKPLPVPNRLHSDLSMDFITSLPPTRGRGSQYLWVIVDRLSKSVTLEEMDTMEAEACAQR 535
Query: 1148 YWKRIVCRFGIPRAIVSDNGTQFSSSQTREFCKEMGIQMRFASVEHPQTNGQVESANRVI 1207
+ G+P++IVSD G+ + REFC+ G+ ++ HPQT+G E N+ I
Sbjct: 536 FLSCHYRFHGMPQSIVSDRGSNWVGRFWREFCRLTGVTQLLSTSYHPQTDGGTERWNQEI 715
Query: 1208 LRGLRRRLAEAKGAWLDELPAVLWSYNTTEQSTTRETPFRMTYG--VDAMLPVEIDNFTW 1265
LR + ++ W D LP V + S+ TPF + +G VD + VE
Sbjct: 716 QAVLRAYVCWSQDNWGDLLPTVQLALRNRHNSSIGATPFFVEHGYHVDPIPTVE------ 877
Query: 1266 RTRPGFEEENQANMAVELDLLSETRDEAHIRETAMKQRVAAKFNSRVRVRD-MQVGDL-- 1322
G E +A + + + + A +QR A N R D QVGD
Sbjct: 878 -DTGGVVSEGEAAAQLLVKRMKDVTGFIQAEIVAAQQRSEASANKRRCPADRYQVGDKVW 1054
Query: 1323 --VLKWRSGAPGNKL 1335
V ++S P KL
Sbjct: 1055 LNVSNYKSPRPSKKL 1099
>BG644693 weakly similar to GP|18767374|g Putative 22 kDa kafirin cluster;
Ty3-Gypsy type {Oryza sativa}, partial (15%)
Length = 716
Score = 81.3 bits (199), Expect = 2e-15
Identities = 58/181 (32%), Positives = 87/181 (48%)
Frame = +2
Query: 381 LNPSVKPVSQLRRRLGGDKGKAVQQEVDKLLAAEFIREVKYPTWLANVVMVKKANGKWRM 440
L P++ P+ R+ K K ++ ++ LL FI+ YP + V+ +KK +G RM
Sbjct: 53 LLPNMNPI*IPSYRINPLKLKVLKLQLKDLLEKGFIQPSIYP*GVV-VLFLKKKDGFLRM 229
Query: 441 CVDYTDLNKACPKDSYPLPSIDSLVDGASGNELLSLMDAYSGYHQIRMHPADEDKTAFMT 500
+DY LN K YPLP ID L D G++ +D G HQ R+ D KTAF
Sbjct: 230 SIDYPQLNNVNIKIKYPLPLIDELFDNLQGSKWFFKIDLRLG*HQHRVIGEDVPKTAFRI 409
Query: 501 ARVNYCYRTMPFGLKNAGATYQRLMDRVFARQVGRNMEVYVDDMIVKSVRGLDHHQDLEE 560
+Y M FG N + LM+RVF + + V+ +D+++ S +H L
Sbjct: 410 RYGHYEILVMSFG*TNPPMAFMELMNRVFQDYLDSLVIVFSNDILIYSKNENEHENHLRL 589
Query: 561 A 561
A
Sbjct: 590 A 592
>AJ497987 weakly similar to GP|9927273|dbj Similar to Arabidopsis thaliana
chromosome II BAC F26H6; putative retroelement pol
polyprotein, partial (1%)
Length = 636
Score = 67.4 bits (163), Expect = 4e-11
Identities = 46/167 (27%), Positives = 74/167 (43%), Gaps = 29/167 (17%)
Frame = -2
Query: 729 KAALAVLVTARRLRPYFQSFPVKVRTDL-PLRQVLQKPDLSGRLVAWSVELSEYGLQYDK 787
K A+ A+RLR Y + + + + P++ + +KP L+GR+ W + LSEY ++Y
Sbjct: 635 KTCCALAWAAKRLRHYMINHTTWLVSKMDPIKYIFEKPALTGRIARWQMLLSEYDIEYRS 456
Query: 788 RGTVGAQSLADFVV-----ELTPDRFERVDTQ-----------------------WTLFV 819
+ + LAD + + P +F+ D + W L
Sbjct: 455 QKAIKGSILADHLAHQPLEDYRPIKFDFPDEEIMYLKMKDCDEPLFGEGPDPDSVWGLIF 276
Query: 820 DGSSNSSGSGAGVALEGPGELVLEQSLKFEFKATNNQAEYEALIAGL 866
DG+ N G+G G L P + + + F TNN AEYEA I G+
Sbjct: 275 DGAVNVYGNGIGAVLLTPKGAHIPFTARLRFDCTNNIAEYEACIMGI 135
>BG644740 similar to PIR|A84460|A84 probable retroelement pol polyprotein
[imported] - Arabidopsis thaliana, partial (4%)
Length = 754
Score = 48.1 bits (113), Expect = 2e-05
Identities = 20/41 (48%), Positives = 28/41 (67%)
Frame = -1
Query: 426 ANVVMVKKANGKWRMCVDYTDLNKACPKDSYPLPSIDSLVD 466
A ++ V+K +G +RMC+DY NK K+ YPLP ID+L D
Sbjct: 238 AALLFVRKKDGYFRMCIDYRQFNKVTTKNKYPLPRIDNLFD 116
>TC83629 weakly similar to GP|6692261|gb|AAF24611.1| putative RNase H
{Arabidopsis thaliana}, partial (21%)
Length = 1071
Score = 45.8 bits (107), Expect = 1e-04
Identities = 34/113 (30%), Positives = 52/113 (45%), Gaps = 5/113 (4%)
Frame = +2
Query: 824 NSSGSGAGVALEGPGELVLEQSLKFEFKA-----TNNQAEYEALIAGLKLAREVKIRSLL 878
N +GAG L L + SL + F+ T AEY AL+ GLK A + +
Sbjct: 587 NRGPAGAGALL-----LAEDGSLLYGFRQGLGHQTKESAEYRALLLGLKHASMKGFKYVT 751
Query: 879 IRTDSQLVENQVKGTFQVKDPNLIKYLERVRYLMTLFQEVVVEYVPRAENQRA 931
+ DS+LV NQ+ +++KD +L K L F ++++ R N A
Sbjct: 752 AKGDSELVINQILDPWKIKDEHLKKLCAEALELSDNFHSFRIQHISRERNYGA 910
>TC92306
Length = 521
Score = 44.7 bits (104), Expect = 3e-04
Identities = 28/73 (38%), Positives = 37/73 (50%)
Frame = +2
Query: 863 IAGLKLAREVKIRSLLIRTDSQLVENQVKGTFQVKDPNLIKYLERVRYLMTLFQEVVVEY 922
I GL AR + +R D Q V Q G+++V +PNL L + F+ V VE+
Sbjct: 2 ILGLNEARNQGYEHVHVRGDFQXVCKQFXGSWKVNNPNLRNLCNXAVELKSNFKSVSVEH 181
Query: 923 VPRAENQRADALA 935
VPR N ADA A
Sbjct: 182 VPRGXNXAADAQA 220
>TC90865 homologue to GP|9049326|dbj|BAA99336.1 orf129b {Beta vulgaris},
partial (16%)
Length = 940
Score = 41.6 bits (96), Expect = 0.002
Identities = 22/55 (40%), Positives = 33/55 (60%), Gaps = 1/55 (1%)
Frame = -3
Query: 155 NIRRVLLDQGSSADIIYGDAFDKLGLTDNDLTPYAG-TLVGFAGEQVMVRGYIDL 208
++R+++LD GSS DI+Y + F L L + +L Y G L GF G GY++L
Sbjct: 938 DVRKIILD*GSSCDIMYSNMF*TLQLLEKNLISYVGFDLQGFNGATTRP*GYVEL 774
>TC93066 weakly similar to GP|19920130|gb|AAM08562.1 Putative retroelement
{Oryza sativa} [Oryza sativa (japonica cultivar-group)],
partial (10%)
Length = 823
Score = 37.7 bits (86), Expect = 0.031
Identities = 27/107 (25%), Positives = 42/107 (39%), Gaps = 24/107 (22%)
Frame = +1
Query: 1160 RAIVSDNGTQFSSSQTREFCKEMGIQMRFASVEHPQTNGQVESANRVILRGLRRRLAEA- 1218
+ +++DN +F SS EFC GI +PQ NG E R +L R L+ A
Sbjct: 289 KKLITDN*LEFCSSDFNEFCTNHGIARHKTIPRNPQQNGVAERMIRTLLERARCMLSNAG 468
Query: 1219 ----KGAWLD-------------------ELPAVLWSYNTTEQSTTR 1242
+ W++ ++P +WS N + S R
Sbjct: 469 L*N*RDLWVEAASTACHLVNRSPHSALDFKVPEDIWSGNLVDYSNLR 609
>BG454871 weakly similar to GP|10140673|g putative gag-pol polyprotein {Oryza
sativa (japonica cultivar-group)}, partial (7%)
Length = 674
Score = 36.2 bits (82), Expect = 0.090
Identities = 28/113 (24%), Positives = 49/113 (42%)
Frame = +2
Query: 1176 REFCKEMGIQMRFASVEHPQTNGQVESANRVILRGLRRRLAEAKGAWLDELPAVLWSYNT 1235
++ K G + +S HP ++GQ E+ N+ LR + W P + YNT
Sbjct: 44 KQLFKLHGTILTMSSAYHP*SDGQSEALNKGXEMYLRCLMFTDPLKWSKAFPWAEYWYNT 223
Query: 1236 TEQSTTRETPFRMTYGVDAMLPVEIDNFTWRTRPGFEEENQANMAVELDLLSE 1288
+ + TPF+ YG D + + + T + Q+ +A +LLS+
Sbjct: 224 SYNISAAMTPFKALYGRDLSMLIRSKGSSKDT-----ADLQSQLAQREELLSQ 367
>BG448304
Length = 637
Score = 34.3 bits (77), Expect = 0.34
Identities = 28/102 (27%), Positives = 44/102 (42%), Gaps = 8/102 (7%)
Frame = +2
Query: 1 WCEFHRSAGHDTDDCRTLQREIDKLIRAGYQGNRQGQWRNGGDQNKAHKREEERADTKGK 60
+C FH+ H TDDC L+ I+ LI+ G N +N +++ + T K
Sbjct: 110 YCRFHKCHRHLTDDCIHLKDTIEILIQRGR--------LNQLTKNPEPEKQTVKLITDKK 265
Query: 61 KKQESAAIATKGADDTFAQH--------SGPPVGTINTIAGG 94
K A++ + D+ F +H + P T N I GG
Sbjct: 266 NKDIVVAMSVEQLDE-FVEHVDITPYSCTWEPFPTANVITGG 388
>TC77786
Length = 1094
Score = 34.3 bits (77), Expect = 0.34
Identities = 13/26 (50%), Positives = 19/26 (73%)
Frame = +3
Query: 2 CEFHRSAGHDTDDCRTLQREIDKLIR 27
C +HRS GHDT++C L+ I+ LI+
Sbjct: 825 CCYHRSRGHDTEECFQLREAIEDLIK 902
>TC89839 homologue to SP|P23466|CYAA_SACKL Adenylate cyclase (EC 4.6.1.1)
(ATP pyrophosphate-lyase) (Adenylyl cyclase). [Yeast],
partial (0%)
Length = 665
Score = 32.7 bits (73), Expect = 1.00
Identities = 14/26 (53%), Positives = 18/26 (68%), Gaps = 1/26 (3%)
Frame = +3
Query: 2 CEFHRSA-GHDTDDCRTLQREIDKLI 26
C FH+ A GHDT+ C L+ E+ KLI
Sbjct: 501 CVFHQGAPGHDTEQCYPLKEEVQKLI 578
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.320 0.136 0.406
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 39,252,476
Number of Sequences: 36976
Number of extensions: 514052
Number of successful extensions: 2346
Number of sequences better than 10.0: 52
Number of HSP's better than 10.0 without gapping: 2290
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 2339
length of query: 1377
length of database: 9,014,727
effective HSP length: 108
effective length of query: 1269
effective length of database: 5,021,319
effective search space: 6372053811
effective search space used: 6372053811
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 64 (29.3 bits)
Lotus: description of TM0029b.1


