

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
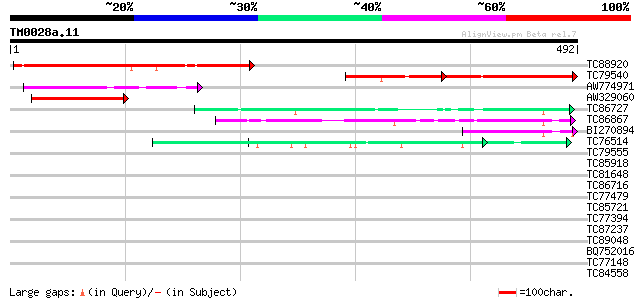
Query= TM0028a.11
(492 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC88920 similar to GP|2829868|gb|AAC00576.1| Unknown protein {Ar... 286 1e-77
TC79540 similar to GP|2829868|gb|AAC00576.1| Unknown protein {Ar... 130 5e-53
AW774971 similar to GP|6016728|gb|A hypothetical protein {Arabid... 109 2e-24
AW329060 similar to GP|6016728|gb|A hypothetical protein {Arabid... 99 5e-21
TC86727 similar to GP|4432859|gb|AAD20707.1| unknown protein {Ar... 63 3e-10
TC86867 similar to GP|9294199|dbj|BAB02101.1 gb|AAF04893.1~gene_... 52 5e-07
BI270894 similar to GP|16209720|gb| At2g35880/F11F19.21 {Arabido... 47 2e-05
TC76514 homologue to PIR|T09648|T09648 nucleolin homolog nuM1 - ... 42 7e-04
TC79555 homologue to GP|10567859|gb|AAG18593.1 L8019.5 {Leishman... 39 0.005
TC85918 similar to GP|13562014|gb|AAK30610.1 fibroin 1 {Plectreu... 39 0.006
TC81648 similar to PIR|T00840|T00840 probable senescence-related... 38 0.010
TC86716 similar to PIR|T14279|T14279 myosin-like protein my5 - c... 37 0.013
TC77479 similar to GP|19423874|gb|AAL87314.1 unknown protein {Ar... 36 0.029
TC85721 similar to SP|P08283|H1_PEA Histone H1. [Garden pea] {Pi... 36 0.038
TC77394 similar to GP|4874305|gb|AAD31367.1| expressed protein {... 36 0.038
TC87237 similar to GP|160409|gb|AAA29651.1|| mature-parasite-inf... 35 0.050
TC89048 weakly similar to PIR|T46215|T46215 hypothetical protein... 35 0.050
BQ752016 homologue to GP|18376006|em probable ATP citrate lyase ... 35 0.066
TC77148 ENOD20 35 0.086
TC84558 weakly similar to SP|Q04634|EF1A_TETPY Elongation factor... 35 0.086
>TC88920 similar to GP|2829868|gb|AAC00576.1| Unknown protein {Arabidopsis
thaliana}, partial (13%)
Length = 828
Score = 286 bits (733), Expect = 1e-77
Identities = 158/218 (72%), Positives = 178/218 (81%), Gaps = 9/218 (4%)
Frame = +2
Query: 4 DQMGETAAASNSALQVSVSFGRFENDSLSWERWSSFSPNKYLEEVEKCATPGSVAQKKAY 63
D+MGET A SNSALQVSVSFGRF+NDSLSWERWSSFSPNKYLEEVEKCATPGSVAQKKAY
Sbjct: 182 DKMGETTA-SNSALQVSVSFGRFDNDSLSWERWSSFSPNKYLEEVEKCATPGSVAQKKAY 358
Query: 64 FEAHYKKIAARKAELLAQEKETEKDSFGSDDQNGVDLSGND------CGTDAEFDVSNAQ 117
FEAHYKKIAARKAELLAQEKE E++SF S++ NG+DLSGN C TD+EF +SN Q
Sbjct: 359 FEAHYKKIAARKAELLAQEKEMERESFRSEENNGIDLSGNGNGNSNACETDSEFGISNTQ 538
Query: 118 GSTTEGVEQ---EISSVGEVSRNLLDALKEEEVAVSRDCYQSSSIEVEESEEQESISHGA 174
GS E ++ E+ VGE+ R+ +D LKEEEVAVS D YQSSS+EV E++E ES SHG+
Sbjct: 539 GSCVEERDEQEIEVIPVGEIDRSHVDDLKEEEVAVSVD-YQSSSVEV-ENKEVESGSHGS 712
Query: 175 EPIDEPEEVVCIKQEENLHVEAEDVKEISHVEYKETEK 212
IDEP + VCIK EE L VEAEDVKEISHV YKETEK
Sbjct: 713 YKIDEPVKDVCIKLEEILDVEAEDVKEISHVVYKETEK 826
>TC79540 similar to GP|2829868|gb|AAC00576.1| Unknown protein {Arabidopsis
thaliana}, partial (7%)
Length = 1003
Score = 130 bits (327), Expect(2) = 5e-53
Identities = 71/118 (60%), Positives = 84/118 (71%)
Frame = +3
Query: 375 GTVSKVPTSSSLRKENGRPNKVESKDTRGGNTVQTTLSRKSDIISGKGKESSRKLDVSSH 434
G + S++LRKENGRP VE + R G+ V+T L+ KSDI S KGK SSRK++ +
Sbjct: 264 GLRQRFQXSTTLRKENGRPATVERMEKRSGSVVRT-LAPKSDIKSEKGKASSRKIEEKPN 440
Query: 435 AKETPRTRLQSKLKEEKEADMKKLKHNFKATPLPAFYGGQKVSKVRPEKVDAKTEKPR 492
K RTRLQ+KLKEEKE +MKKLKH+FKATPLP Y GQKVSK R EK DAKTE R
Sbjct: 441 TKVVERTRLQTKLKEEKEVEMKKLKHDFKATPLPTSYRGQKVSKTRAEKGDAKTESRR 614
Score = 95.9 bits (237), Expect(2) = 5e-53
Identities = 55/95 (57%), Positives = 69/95 (71%), Gaps = 7/95 (7%)
Frame = +1
Query: 292 ITSSEESKKVANKPLHKSLNLEPSNPDPAP-----LP-TLRKSFIMESMGDKDIVKRAFK 345
+TSS E+KK A + LH S++L PSNP+P P +P T+RKS IM+SMGDKDIVKRA
Sbjct: 1 VTSSVENKKAATRSLHLSMSLGPSNPEPVPHTTDPVPHTMRKSLIMDSMGDKDIVKRA-- 174
Query: 346 TFKTFQNNYSQPKSSG-EDRSSVKKQVPSRGTVSK 379
FKTFQ N++QPK+SG ED+SSV KQ +G K
Sbjct: 175 -FKTFQKNFNQPKTSGEEDKSSVIKQASFKGDCVK 276
>AW774971 similar to GP|6016728|gb|A hypothetical protein {Arabidopsis
thaliana}, partial (22%)
Length = 520
Score = 109 bits (273), Expect = 2e-24
Identities = 66/155 (42%), Positives = 92/155 (58%)
Frame = +3
Query: 13 SNSALQVSVSFGRFENDSLSWERWSSFSPNKYLEEVEKCATPGSVAQKKAYFEAHYKKIA 72
+N AL SVSFGRF ++SL+WE+WSSFS N+Y+EE E+ + PGSVAQKKA+FEAHYKK+A
Sbjct: 72 TNHALGQSVSFGRFMSESLAWEKWSSFSHNRYVEEAERYSKPGSVAQKKAFFEAHYKKLA 251
Query: 73 ARKAELLAQEKETEKDSFGSDDQNGVDLSGNDCGTDAEFDVSNAQGSTTEGVEQEISSVG 132
A+KA L +++ + N + ND D++ D S E V +E G
Sbjct: 252 AQKAAALLEQQNN-----AASQNNVTEKEDNDENNDSQKD------SKYEVVVKEDRDDG 398
Query: 133 EVSRNLLDALKEEEVAVSRDCYQSSSIEVEESEEQ 167
+V + LD+ E+ C S S ++EE E Q
Sbjct: 399 KVLSHALDSNMEK-------CIASESNKLEECETQ 482
>AW329060 similar to GP|6016728|gb|A hypothetical protein {Arabidopsis
thaliana}, partial (19%)
Length = 524
Score = 98.6 bits (244), Expect = 5e-21
Identities = 44/84 (52%), Positives = 64/84 (75%)
Frame = +3
Query: 20 SVSFGRFENDSLSWERWSSFSPNKYLEEVEKCATPGSVAQKKAYFEAHYKKIAARKAELL 79
SVSFGRF +SL+WE+WS+FS N+Y+EE E+ + PGSVA+KKA+FEAHYKK+AA+KA L
Sbjct: 120 SVSFGRFTTESLAWEKWSTFSTNRYVEEAERYSKPGSVAEKKAFFEAHYKKLAAQKAAAL 299
Query: 80 AQEKETEKDSFGSDDQNGVDLSGN 103
++++++ D+ VD + N
Sbjct: 300 LEQEKSDSLEMEEHDEAEVDNTNN 371
>TC86727 similar to GP|4432859|gb|AAD20707.1| unknown protein {Arabidopsis
thaliana}, partial (27%)
Length = 1852
Score = 62.8 bits (151), Expect = 3e-10
Identities = 74/335 (22%), Positives = 129/335 (38%), Gaps = 5/335 (1%)
Frame = +2
Query: 161 VEESEEQESISHGAEPIDEPEEVVCIKQEENLHVEAEDVKEISHVEYKETEKASEVEAKD 220
+E+ Q+ + + + + V IKQ + D +S E +E ++E +
Sbjct: 185 LEDGLHQQQLFNSQQDAVDFNVVTQIKQTVLSNGNFND--NVSMEEEEEEGSNGKIEGNN 358
Query: 221 VKLDHPKESKVASVNRESNAAKTKKK---SMLPKSKASQISTPRNSKPASTPSKPLTSAS 277
V + E ++ +S K K S LP + ++++ R +K + S+
Sbjct: 359 VNVSKEVEIEIVDETEKSRTKKDLVKNNNSKLPSPRGLRMTSVRKNKDGKDEEAAVASSV 538
Query: 278 STKKGNSPSLSRKPITSSEESKKVANKPLHKSLNLEPSNPDPAPLPTLRKSFIMESMGDK 337
S S R+P+ + + K + H S AP+ R I + D
Sbjct: 539 SNGTSTFDSHPRQPVNNRAVNDKQTHLSKHSGKTDAASTE--APMEKTRPHLIKKEPLDN 712
Query: 338 DIVKRAFKTFKTFQNNYSQPKSSGEDRSSVKKQVPSRGTVSKVPTSSSLRKENGRPNKVE 397
+P + S PTS E+ +P +V
Sbjct: 713 ---------------------------------LPGKAE-SSFPTS-----EDAKPRRVG 775
Query: 398 SKDTRGGNTVQTTLSRKSDIISGKGKESSRKLDVSSHAKETPRTRLQSKLKEEKEADMKK 457
+ T G S K + + + KE KL+ HAKE + +Q+K KE +EA++K+
Sbjct: 776 TMPTYG-------FSFKCNERAERRKEFYSKLEERIHAKEVEESNIQAKTKESQEAEIKR 934
Query: 458 LKHN--FKATPLPAFYGGQKVSKVRPEKVDAKTEK 490
L+ FKATP+P+FY S+V +K+ K
Sbjct: 935 LRKKLAFKATPMPSFYQEPTPSRVELKKIPTTRAK 1039
>TC86867 similar to GP|9294199|dbj|BAB02101.1
gb|AAF04893.1~gene_id:MXC7.13~similar to unknown protein
{Arabidopsis thaliana}, partial (38%)
Length = 1557
Score = 52.0 bits (123), Expect = 5e-07
Identities = 79/327 (24%), Positives = 138/327 (42%), Gaps = 14/327 (4%)
Frame = +1
Query: 179 EPEEVVCIKQEENLHVEAEDVKEISHVEYKETEKASEVEAKDVKLDHPKESKVASVNRES 238
E ++ IK+ + + V ++ + + S E T+ + E+ +H E+ + ES
Sbjct: 124 EDSDICIIKEPDRVVVYSDGISQDSGHE-TGTDNHNMTES----YEHINETTEHHSSEES 288
Query: 239 NAAKTKKKSMLPKS-KASQISTPRNSKPASTPSKPLTSASSTKKGNSPSLSRKPITSSEE 297
K+ S KA +S R S+ TP + + +++
Sbjct: 289 TKEYEVKECTTEVSVKAPDVSNIRKSEEKLTPDF------------------EGVLNAKS 414
Query: 298 SKKVANKPLHKSLN-LEPSNPDPAPLPTLRKSFIME----------SMGDKDIVKRAFKT 346
SK + HK + ++P+ P P L T R++ I+ G K + K+ +
Sbjct: 415 SKPHKTRGNHKPRDTVKPTVPRPFSLATERRATILTRPTFEEDNKGGHGRKSVNKKNVLS 594
Query: 347 FKTFQNNYSQPKSSGEDRSSVKKQVPSRGTVSKVPTSSSLRKENGRPNKVESKDTRGGNT 406
T + N Q K+ R +K P S S S+ +G V+S +R T
Sbjct: 595 SNTLKQN--QLKTPLVPRKPLK---PDNKKHSDEDDSCSVA--SGTATSVKSFKSRA--T 747
Query: 407 VQTTLSRKSDIISGKGKESSRKLDVSSHAKETPRTRLQSKLKEEKEADMKKLKHN--FKA 464
V + S +S + + KE KL+ A E + + +++ KEEKE +K+L+ + FKA
Sbjct: 748 VASAPSFRSTERAQRRKEFYSKLEEKQQAMEAEKNQNEARSKEEKEEAIKQLRRSLKFKA 927
Query: 465 TPLPAFYGGQKVSKVRPEKVDAKTEKP 491
+P+P+FY + P KVD K P
Sbjct: 928 SPMPSFY-----HEGPPPKVDLKKLPP 993
>BI270894 similar to GP|16209720|gb| At2g35880/F11F19.21 {Arabidopsis
thaliana}, partial (21%)
Length = 652
Score = 46.6 bits (109), Expect = 2e-05
Identities = 35/104 (33%), Positives = 52/104 (49%), Gaps = 5/104 (4%)
Frame = +2
Query: 394 NKVESKDTRGGNTVQTTLSRKSDIISGKGKESSRKLDVSSHAKETPRTRLQSKLKEEKEA 453
N S T G + + S + + + K +E KL+ KE + LQ K KE +EA
Sbjct: 95 NSTTSSLTPGKKSNGSGFSFRLEERAEKRREFFSKLEEKIQEKEAEISNLQEKSKESQEA 274
Query: 454 DMKKLKHN--FKATPLPAFYGGQKVSKVRPEKVDAK---TEKPR 492
++KKL+ FKA P+P+FY K P KV+ K T +P+
Sbjct: 275 EIKKLRKRMTFKAAPMPSFY------KEPPPKVELKKIPTTRPK 388
>TC76514 homologue to PIR|T09648|T09648 nucleolin homolog nuM1 - alfalfa,
partial (61%)
Length = 1288
Score = 41.6 bits (96), Expect = 7e-04
Identities = 63/288 (21%), Positives = 108/288 (36%), Gaps = 8/288 (2%)
Frame = +3
Query: 208 KETEKASEVEAKDVKLDHPKESKVASVNRESNAAKTKKKSMLPKSKASQISTPRNSKPAS 267
K +K ++VK K+ KV V + A K KK ++S+ + A
Sbjct: 168 KSGKKGKRQAEEEVKAVSAKKQKVEEVAAKQKALKVVKKEESSSEESSESEDEQPVVKAP 347
Query: 268 TPSKPLTSASSTKKGNSPSLSRKPITSSEES--------KKVANKPLHKSLNLEPSNPDP 319
PSK + KKGN +P T+SEES ++ KP+ K++ PS
Sbjct: 348 APSKK----TPAKKGNVKKA--QPETTSEESDSDSSSSDEEEVKKPVSKAV---PSKNGS 500
Query: 320 APLPTLRKSFIMESMGDKDIVKRAFKTFKTFQNNYSQPKSSGEDRSSVKKQVPSRGTVSK 379
AP K T + S+ +SS ED+ K VPS+ +
Sbjct: 501 APA----------------------KKVDTSEEEDSE-ESSDEDKKPAAKAVPSKNGSAP 611
Query: 380 VPTSSSLRKENGRPNKVESKDTRGGNTVQTTLSRKSDIISGKGKESSRKLDVSSHAKETP 439
++S ++ + + +D + + S G ++K D S +++
Sbjct: 612 AKKAASDEEDTDESSDEDEEDEK---------PAAKAVPSKNGSVPAKKADTESSDEDS- 761
Query: 440 RTRLQSKLKEEKEADMKKLKHNFKATPLPAFYGGQKVSKVRPEKVDAK 487
+S +E+K+ K K+ T A ++ + E DAK
Sbjct: 762 ----ESSDEEDKKPAAKASKNVSAPTKKAASSSDEESDEESDEDEDAK 893
Score = 41.6 bits (96), Expect = 7e-04
Identities = 67/317 (21%), Positives = 119/317 (37%), Gaps = 27/317 (8%)
Frame = +3
Query: 125 EQEISSVGEVSRNLLDALKEEEVAVSRDCYQSSSIEVEESEEQESISHGAEPIDE-PEEV 183
E+E+ +V + + + +++ +SSS E ESE+++ + P + P +
Sbjct: 198 EEEVKAVSAKKQKVEEVAAKQKALKVVKKEESSSEESSESEDEQPVVKAPAPSKKTPAKK 377
Query: 184 VCIKQEENLHVEAEDVKEISHVEYKETEKA---------SEVEAKDVKLDHPKESKVASV 234
+K+ + E + S + +E +K AK V ++S+ +S
Sbjct: 378 GNVKKAQPETTSEESDSDSSSSDEEEVKKPVSKAVPSKNGSAPAKKVDTSEEEDSEESSD 557
Query: 235 NRESNAAKT--KKKSMLPKSKAS--QISTPRNSKPASTPSKPLTSASSTKKGNSPSLSRK 290
+ AAK K P KA+ + T +S KP A +K G+ P+
Sbjct: 558 EDKKPAAKAVPSKNGSAPAKKAASDEEDTDESSDEDEEDEKPAAKAVPSKNGSVPAKKAD 737
Query: 291 PITS---SEESKKVANKPLHKSLNLEPSNPDPAPLPTLRKSFIMESMGDKD--------I 339
+S SE S + KP K+ + S P + + ES D+D
Sbjct: 738 TESSDEDSESSDEEDKKPAAKA-SKNVSAPTKKAASSSDEESDEESDEDEDAKPVSKPAA 914
Query: 340 VKRAFKTFKTFQNNYSQPKSSGEDRSSVKKQVPSRGTVSKVPTSSSLRKENG--RPNKVE 397
V + K + ++ SS ED+ V + + V K + S +E P+K
Sbjct: 915 VAKKSKKDSSDSDDEDDDSSSDEDKKPVAAKKEDKMNVDKDGSDSDQSEEESEDEPSKTP 1094
Query: 398 SKDTRGGNTVQTTLSRK 414
K + V S K
Sbjct: 1095QKKIKDVEMVDAGKSGK 1145
>TC79555 homologue to GP|10567859|gb|AAG18593.1 L8019.5 {Leishmania major},
partial (1%)
Length = 1410
Score = 38.9 bits (89), Expect = 0.005
Identities = 32/95 (33%), Positives = 43/95 (44%), Gaps = 16/95 (16%)
Frame = +2
Query: 250 PKSKASQISTP-RNSKPASTPSKPLTSASSTKK--GNSPSLSRKPITSSEESKK------ 300
P S S + P R S PA+T P TS S + SPS +R P TSS ++
Sbjct: 488 PSSTRSSPAPPSRPSSPATTTRPPPTSKPSPTQTPSRSPSSARTPSTSSTPTRAPLLSSP 667
Query: 301 -------VANKPLHKSLNLEPSNPDPAPLPTLRKS 328
A +P +SL P P PA LP+ R++
Sbjct: 668 PPPPPFFPARRPGTRSLPKSPPRPRPARLPSARRA 772
>TC85918 similar to GP|13562014|gb|AAK30610.1 fibroin 1 {Plectreurys tristis},
partial (8%)
Length = 4117
Score = 38.5 bits (88), Expect = 0.006
Identities = 103/481 (21%), Positives = 194/481 (39%), Gaps = 35/481 (7%)
Frame = -1
Query: 46 EEVEKCATPGSVAQKKAYFEAHYKKIAARKAELLAQEKETEKDSFGSDDQNGVDLSGNDC 105
EE + P +Q+K+ K IA ++ + A E++T S++ +++ +
Sbjct: 2611 EEPKVTKEPEKESQEKSEEAEQPKAIAIVESTVEANEEKTAAAI--SEETKIIEVEPTET 2438
Query: 106 GTDAEF--DVSNAQGSTTEGVEQEISSVGEVSRNLLDALKEEEVAVSRDCYQSSSIEVEE 163
+ +V+ A+ E V E+ + + LKEE VA + ++ E
Sbjct: 2437 VKEEPMVTEVATAEIVKEEPVVTEVEAT--------ETLKEELVATEVEATETVKEEPAA 2282
Query: 164 SEEQESISHGAEPIDEPEEVVCIKQEENLHVEA---EDVKE---ISHVEYKETEK----A 213
+E + + + EP+ + +EE L EA E VKE + VE ET K A
Sbjct: 2281 TEVKVTETVKDEPVVTEVDPTETVKEEPLATEAEATETVKEEPVATEVEPTETVKEEPVA 2102
Query: 214 SEVEAKDVKLDHPKESKV---ASVNRESNAAKTK-KKSMLPKSKASQISTPRNSKPASTP 269
+EVEA + D P ++V +V E A +T+ +++ + A+++ K S
Sbjct: 2101 TEVEATETVKDEPAATEVEATETVKEEPVATETEATETVKEEPVATEVEATETVKEESV- 1925
Query: 270 SKPLTSASSTKKGNSPSLSRKPITSSEESKKVANKPLHKSLNLEPSNPDPAPLPTLRKSF 329
+T+ + ++ +P+T+ E ++ EP+ + T++
Sbjct: 1924 --------ATEVEATETVKEEPVTTVVEPTEMVKD--------EPAATEFEATETVKDET 1793
Query: 330 IMESMGDKDIVK-RAFKTFKTFQNNYSQPKSSGEDRSS------VKKQVPSRGTVSKVPT 382
I+ + + VK A T +P + D + V +V R V + P
Sbjct: 1792 IVTEVDPTETVKEEAVATEVEITKTVKEPLVTEVDLTESVKVKPVVTEVDPREMVKEEPV 1613
Query: 383 SSSLR-----KENGRPNKVESKDTRGGNTVQTTL--SRKSDIISGKGKESSRKLDVSSHA 435
++ ++ KE +VE +T G V T + ++K + + + SS
Sbjct: 1612 ATEVQATETVKEEPVATEVEPTETVKGEPVVTEVEENQKEPEQHSTERREEEQPNASSIP 1433
Query: 436 KETPRTR----LQSKLKE-EKEADMKKLKHNFKATPLPAFYGGQKVSKVRPEKVDAKTEK 490
+++ T ++ K +E E EA + K N +A P +KV V E VD +
Sbjct: 1432 EQSTETNDVIAVEEKSRELEFEAAILKETKNDEAGPAET----EKVEPVVTE-VDENQSE 1268
Query: 491 P 491
P
Sbjct: 1267 P 1265
Score = 35.4 bits (80), Expect = 0.050
Identities = 63/353 (17%), Positives = 123/353 (33%), Gaps = 2/353 (0%)
Frame = -1
Query: 140 DALKEEEVAVSRDCYQSSSIEVEESEEQESISHGAEPIDEPEEVVCIKQEENLHVEAEDV 199
+ +KEE VA + ++ E +E + + + EP+ E +EE + E E
Sbjct: 2128 ETVKEEPVATEVEATETVKDEPAATEVEATETVKEEPVATETEATETVKEEPVATEVEAT 1949
Query: 200 KEISHVEYKETEKASEVEAKDVKLDHPKESKVASVNRESNAAKTKKKSMLPKSKASQIST 259
+ + KE A+EVEA + + P + V + + K I T
Sbjct: 1948 ETV-----KEESVATEVEATETVKEEPVTTVVEPTEMVKDEPAATEFEATETVKDETIVT 1784
Query: 260 PRNSKPASTPSKPLTSASSTKKGNSPSLSRKPITSSEESKKVANK-PLHKSLNLEPSNPD 318
+ T TK P ++ +T S + K V + + + EP +
Sbjct: 1783 EVDPTETVKEEAVATEVEITKTVKEPLVTEVDLTESVKVKPVVTEVDPREMVKEEPVATE 1604
Query: 319 PAPLPTLRKSFIMESMGDKDIVKRAFKTFKTFQNNYSQPKSSGEDRSSVKKQVPSRGTVS 378
T+++ + + + VK + N +P+ +R ++
Sbjct: 1603 VQATETVKEEPVATEVEPTETVK-GEPVVTEVEENQKEPEQHSTERREEEQ--------- 1454
Query: 379 KVPTSSSLRKENGRPNKVESKDTRGGNTVQTTLSRKSDIISGKGKESSRKLD-VSSHAKE 437
P +SS+ +++ N V + + + K G + K++ V + E
Sbjct: 1453 --PNASSIPEQSTETNDVIAVEEKSRELEFEAAILKETKNDEAGPAETEKVEPVVTEVDE 1280
Query: 438 TPRTRLQSKLKEEKEADMKKLKHNFKATPLPAFYGGQKVSKVRPEKVDAKTEK 490
K+E+E K+ N + T G +V P+ D + K
Sbjct: 1279 NQSEPGIQSFKQEEEVKPKEETENQEQT-------GDRVEVHPPKDSDIEAVK 1142
>TC81648 similar to PIR|T00840|T00840 probable senescence-related protein
[imported] - Arabidopsis thaliana, partial (36%)
Length = 854
Score = 37.7 bits (86), Expect = 0.010
Identities = 34/105 (32%), Positives = 50/105 (47%), Gaps = 3/105 (2%)
Frame = +2
Query: 226 PKESKVASVNRESNAAKTKKKSMLPKSKASQISTPRNSKPASTPSKPLTSASSTKKGNSP 285
P + + ++R + KS P S +S +T N P ++PS S S+ K SP
Sbjct: 167 PASTPLRLLHRRKLPNRFLSKSPEPFSTSSINNTALNLLPVTSPS----SVSAKVKIPSP 334
Query: 286 SLSRKPITSSEESKKVANKPLHKSLNLEPSNPDPAP---LPTLRK 327
S+ P+ S+ K KPL KS+ L S+P P P +PT K
Sbjct: 335 SMLGSPMKSNGLLLKT--KPLLKSMILTTSSPSPLPRVMIPTKMK 463
>TC86716 similar to PIR|T14279|T14279 myosin-like protein my5 - common
sunflower, partial (38%)
Length = 2675
Score = 37.4 bits (85), Expect = 0.013
Identities = 53/226 (23%), Positives = 89/226 (38%), Gaps = 44/226 (19%)
Frame = +2
Query: 136 RNLLDALKEEEVAVSRDCYQSSSIEVEES--------EEQESISHGAEP--------IDE 179
R L+ K +EVA RD + I+VEE+ E + A P I++
Sbjct: 59 RTDLEEEKAQEVAKLRDALHAMQIQVEEANAKVIKEREASQKAIQDAPPVIKETPVIIED 238
Query: 180 PE-------EVVCIKQEENLHVEA-EDVKEI-SHVEYKETEKASEVEAKDVKLDHPK--- 227
E EV C+K+ L EA E+ K + E + E +VE D K D +
Sbjct: 239 TEKINSLLAEVNCLKESLLLEREAKEEAKRAQAETEARSKELFKKVEDSDRKADQLQELV 418
Query: 228 ---ESKVASVNRESNAAKTKKKSMLPKSKASQI--------STPRNSK----PASTPSKP 272
E K+++ E+ + + ++ P +K+ TP N A TPS
Sbjct: 419 QRLEEKISNSESENQVLRQQALAVSPTAKSLAARPRSVIIQRTPENGNALNGEAKTPSDM 598
Query: 273 LTSASSTKKGNSPSLSRKPIT-SSEESKKVANKPLHKSLNLEPSNP 317
+ S+ ++ S +K + +E++ V K + + L P
Sbjct: 599 TLALSNVREPESEGKPQKSLNDKQQENQDVLIKCISQDLGFSEGKP 736
>TC77479 similar to GP|19423874|gb|AAL87314.1 unknown protein {Arabidopsis
thaliana}, partial (21%)
Length = 2241
Score = 36.2 bits (82), Expect = 0.029
Identities = 26/96 (27%), Positives = 42/96 (43%), Gaps = 5/96 (5%)
Frame = +3
Query: 249 LPKSKASQISTPRNSKPAS---TPSKPLTSASSTKKGNSPSLSRKPITSSEESKKVANKP 305
+P ++ +Q+ P N K + PS P + S + K +SP P S S+ ++
Sbjct: 330 MPINQINQMGFPENHKSSGFGIPPSYPNPNTSLSPKPSSPYPQFMPPQSQTHSRSLSQPT 509
Query: 306 LHK--SLNLEPSNPDPAPLPTLRKSFIMESMGDKDI 339
S +L P +P P+P P S G KD+
Sbjct: 510 FLSLDSFSLPPLSPSPSPSPYHLSSNPFSESGSKDV 617
>TC85721 similar to SP|P08283|H1_PEA Histone H1. [Garden pea] {Pisum
sativum}, partial (60%)
Length = 995
Score = 35.8 bits (81), Expect = 0.038
Identities = 34/117 (29%), Positives = 55/117 (46%), Gaps = 2/117 (1%)
Frame = +1
Query: 228 ESKVASVNRESNAAKTKKKSMLPKSKASQISTPRNSKPASTPSKPLTSASSTKKGNSPSL 287
++K A+ + K K KS+ K KA+ + P KPA+T SK + A + K +
Sbjct: 514 KAKPAAKPKAKAVVKPKTKSVTAKPKAAAATKP---KPAATKSKAASKAKTAAKAKPNAK 684
Query: 288 SRKPITSSEESKKV-ANKPLHKSLNLEPSNPDPAPLPTLR-KSFIMESMGDKDIVKR 342
K + KKV A KP K + + AP+ +++ K+ ++S K VKR
Sbjct: 685 VAKTTARTSPGKKVAAAKPAAKKV---AAAKKKAPVKSVKPKTKTVKSPAKKVSVKR 846
>TC77394 similar to GP|4874305|gb|AAD31367.1| expressed protein {Arabidopsis
thaliana}, partial (61%)
Length = 1396
Score = 35.8 bits (81), Expect = 0.038
Identities = 22/107 (20%), Positives = 49/107 (45%)
Frame = +2
Query: 361 GEDRSSVKKQVPSRGTVSKVPTSSSLRKENGRPNKVESKDTRGGNTVQTTLSRKSDIISG 420
GE + + ++ K S +++ NG+PN + ++ ++V R + S
Sbjct: 956 GEKKEITAESKNAKKKKRKDKASKEVKESNGQPNSSDVENAEEDSSVVDVKERLKKVASM 1135
Query: 421 KGKESSRKLDVSSHAKETPRTRLQSKLKEEKEADMKKLKHNFKATPL 467
+ K+SS+++D ++ A ++L K KK K+++ P+
Sbjct: 1136 RKKKSSKEMDAAARAAAQEAAARNARLAAAK----KKEKNHYNQQPV 1264
>TC87237 similar to GP|160409|gb|AAA29651.1|| mature-parasite-infected
erythrocyte surface antigen {Plasmodium falciparum},
partial (2%)
Length = 2007
Score = 35.4 bits (80), Expect = 0.050
Identities = 81/400 (20%), Positives = 152/400 (37%), Gaps = 19/400 (4%)
Frame = +1
Query: 81 QEKETEKDSFGSDDQNGVDLSGNDCGTDAEFDVSNAQGSTTEGVEQEISSVGEVSRN--- 137
+E + K+ SD+Q D N+ + + NA + + QE S+ E +
Sbjct: 511 EEHDKIKEDTSSDNQVQ-DGEKNNEAREENYSGDNASSAVVDNKSQESSNKTEEQFDKKE 687
Query: 138 -----LLDALKEEEVAVSRDCYQSSSIEVEESEEQESISHGAEPIDEPEEVVCIKQE-EN 191
L E S D + + + ESE+ ++ + P E KQE E
Sbjct: 688 KNEFELESQKNSNETTESTDSTITQNSQGNESEKDQAQTENDTPKGSASESDEQKQEQEQ 867
Query: 192 LHVEAEDVKEISHVEYKETEKASEVEAKDVKLDHPKESKVASVNRESNAAKTKKKSMLPK 251
+ +DV +T S D + S+ A+ +E ++A + P
Sbjct: 868 NNTTKDDV---------QTTDTSSQNGNDTTEKQNETSEDANSKKEDSSA----LNTTPN 1008
Query: 252 SKASQISTPRNSKPASTPSKPLTSASSTKKGNSPSLSRKPITSSEESKKVANKPLHKSLN 311
++ S+ + ++T TS+S T+ GN+ K T+ K + +S +
Sbjct: 1009NEDSKSGVAGDQADSTT----TTSSSETQDGNTNHGEYKDTTNENPEKNSGQEGTQESGS 1176
Query: 312 LEPS--NPDPAPLPT-LRKSFIMESMGDKDIVKRAFKTFKTFQNNYSQPKSSGEDRSSVK 368
+ N D A L + S KD A ++ QN+ + +SG+ ++
Sbjct: 1177SSNTFDNKDAASNKVQLTTTSDTSSEQKKDESSSAESKSESSQNDNA---NSGQSNTTSD 1347
Query: 369 KQVPSRGTVSKVPTSS-------SLRKENGRPNKVESKDTRGGNTVQTTLSRKSDIISGK 421
+ S+V TSS S + N N+ +SK+ N T + + ++
Sbjct: 1348ESANDNKDSSQVTTSSENSAEGNSNTENNSDENQNDSKNNENTNDSGNTSNDAN--VNEN 1521
Query: 422 GKESSRKLDVSSHAKETPRTRLQSKLKEEKEADMKKLKHN 461
E++ + S + + ++SK KE E+ K + +N
Sbjct: 1522QNENAAQTKTSENEGDAQNESVESK-KENNESAHKDVDNN 1638
>TC89048 weakly similar to PIR|T46215|T46215 hypothetical protein T8P19.220
- Arabidopsis thaliana, partial (35%)
Length = 1329
Score = 35.4 bits (80), Expect = 0.050
Identities = 39/160 (24%), Positives = 72/160 (44%), Gaps = 8/160 (5%)
Frame = +2
Query: 156 SSSIEVEESEEQESISHGAEPIDEPEEVVCIKQEENLHVEAEDVKEISHVEYK----ETE 211
S + E + + + + + A+ +P+E V + +++ E+++ KEI E E E
Sbjct: 302 SMATEALDDNKPKQVENEAKDEPKPKEEVEVTEQDAEVAESDEKKEIDGHEDSDKDDEDE 481
Query: 212 KASEVEAKDVKLDHPKESKVASVNRESNAAKTKKKSMLPKSKASQISTPRNSKPASTPSK 271
E +A D + + E K V +ES+ K+ + + + R S+P +PSK
Sbjct: 482 GGDEDDADDDEEEDAGEEK-KGVKKESSEKKSPVTPTSDRPTRERKTVERYSEP--SPSK 652
Query: 272 PLTSASS----TKKGNSPSLSRKPITSSEESKKVANKPLH 307
S+SS +KG L P + + SK+ + LH
Sbjct: 653 FGRSSSSKGLIIEKGRGTQLKDIPNVAFKLSKRKPDDNLH 772
>BQ752016 homologue to GP|18376006|em probable ATP citrate lyase subunit 1
{Neurospora crassa}, partial (37%)
Length = 759
Score = 35.0 bits (79), Expect = 0.066
Identities = 26/70 (37%), Positives = 36/70 (51%)
Frame = -2
Query: 259 TPRNSKPASTPSKPLTSASSTKKGNSPSLSRKPITSSEESKKVANKPLHKSLNLEPSNPD 318
+PR SK S+P T+ASS +SPSLS P++ S K S++ PS+P
Sbjct: 515 SPRPSKTCPRSSRPSTTASSRTAPSSPSLS--PLSPRSPSTTRGPKSSVSSVSPLPSSP- 345
Query: 319 PAPLPTLRKS 328
P+P R S
Sbjct: 344 PSPTTVDRSS 315
>TC77148 ENOD20
Length = 1108
Score = 34.7 bits (78), Expect = 0.086
Identities = 28/81 (34%), Positives = 34/81 (41%), Gaps = 6/81 (7%)
Frame = +3
Query: 250 PKSKASQISTPRNSKPASTPSKPLTSASSTKKGNSPSLSRKPITSSEESKKVANKPLHKS 309
P S +P S P STP S SPSLS+ P S ES +A P
Sbjct: 513 PSPSPSPSPSPSPS-PRSTPIPHPRKRSPASPSPSPSLSKSP--SPSESPSLAPSPSDSV 683
Query: 310 LNLEPS------NPDPAPLPT 324
+L PS +P PAP P+
Sbjct: 684 ASLAPSSSPSDESPSPAPSPS 746
>TC84558 weakly similar to SP|Q04634|EF1A_TETPY Elongation factor 1-alpha
(EF-1-alpha) (14-nm filament-associated protein).
{Tetrahymena pyriformis}, partial (42%)
Length = 611
Score = 34.7 bits (78), Expect = 0.086
Identities = 25/86 (29%), Positives = 43/86 (49%), Gaps = 3/86 (3%)
Frame = +3
Query: 206 EYKETEKASEVEAKDV-KLDHPKESKVASVNRESNAAKTKKKSMLPKS--KASQISTPRN 262
E ++ ++ V ++ V KL ES++ S +R K+ K L + S++STPRN
Sbjct: 12 EPSQSAESRPVSSRPVCKLPSLLESRLLSASRSKCTTKSSLKPDLVTTLVSTSRVSTPRN 191
Query: 263 SKPASTPSKPLTSASSTKKGNSPSLS 288
S+ ++P P T+ + P LS
Sbjct: 192 SREETSPPMPRTTQPPKSPTSWPRLS 269
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.302 0.120 0.317
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 11,816,273
Number of Sequences: 36976
Number of extensions: 146222
Number of successful extensions: 939
Number of sequences better than 10.0: 116
Number of HSP's better than 10.0 without gapping: 905
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 925
length of query: 492
length of database: 9,014,727
effective HSP length: 100
effective length of query: 392
effective length of database: 5,317,127
effective search space: 2084313784
effective search space used: 2084313784
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 17 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 43 (21.8 bits)
S2: 60 (27.7 bits)
Lotus: description of TM0028a.11


