

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0026.15
(145 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
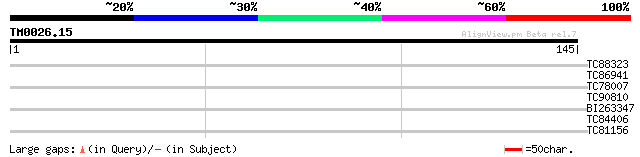
Score E
Sequences producing significant alignments: (bits) Value
TC88323 similar to PIR|T08917|T08917 auxin response factor 9 - A... 28 0.99
TC86941 similar to GP|15294290|gb|AAK95322.1 AT5g02240/T7H20_290... 27 3.8
TC78007 PIR|T09261|T09261 JUN kinase-activation-domain-binding p... 26 4.9
TC90810 similar to GP|7339693|dbj|BAA92898.1 ESTs C72979(E2587) ... 26 4.9
BI263347 25 8.4
TC84406 similar to GP|12039327|gb|AAG46115.1 putative sugar tran... 25 8.4
TC81156 similar to PIR|T45754|T45754 hypothetical protein F24M12... 25 8.4
>TC88323 similar to PIR|T08917|T08917 auxin response factor 9 - Arabidopsis
thaliana, partial (21%)
Length = 1232
Score = 28.5 bits (62), Expect = 0.99
Identities = 13/37 (35%), Positives = 23/37 (62%), Gaps = 6/37 (16%)
Frame = +3
Query: 9 HSWEIILHDEDRDRAWVNDE------HQVRRIYVGSA 39
++WEI+ D++RD V D+ + VRRI++ S+
Sbjct: 912 NTWEIVFTDDERDMMLVGDDPWPEFCNMVRRIFICSS 1022
>TC86941 similar to GP|15294290|gb|AAK95322.1 AT5g02240/T7H20_290
{Arabidopsis thaliana}, partial (98%)
Length = 1121
Score = 26.6 bits (57), Expect = 3.8
Identities = 15/46 (32%), Positives = 27/46 (58%)
Frame = -3
Query: 2 TILRRTGHSWEIILHDEDRDRAWVNDEHQVRRIYVGSAAVRSFDQR 47
+I+R+ H+ I+H E+ + V+ E++ + +VGS A FD R
Sbjct: 915 SIVRQ*SHT---IIHHENPQKREVSWENKALKSFVGSPAPSGFDAR 787
>TC78007 PIR|T09261|T09261 JUN kinase-activation-domain-binding protein
homolog - alfalfa, partial (97%)
Length = 1628
Score = 26.2 bits (56), Expect = 4.9
Identities = 8/22 (36%), Positives = 14/22 (63%)
Frame = +2
Query: 20 RDRAWVNDEHQVRRIYVGSAAV 41
RD+ W ND H +R+ + + A+
Sbjct: 404 RDKPWANDPHYFKRVKISALAL 469
>TC90810 similar to GP|7339693|dbj|BAA92898.1 ESTs C72979(E2587)
AU082675(E2587) correspond to a region of the predicted
gene.~hypothetical, partial (37%)
Length = 737
Score = 26.2 bits (56), Expect = 4.9
Identities = 10/23 (43%), Positives = 15/23 (64%), Gaps = 3/23 (13%)
Frame = +3
Query: 5 RRTGHS---WEIILHDEDRDRAW 24
R TGH W +++ +DRD+AW
Sbjct: 267 RHTGHVLNLWLLVVSTDDRDKAW 335
>BI263347
Length = 582
Score = 25.4 bits (54), Expect = 8.4
Identities = 12/22 (54%), Positives = 15/22 (67%)
Frame = +3
Query: 71 LPIVVGSHQLLNMKSEPEKDPL 92
LP+ V S +LL M SE K+PL
Sbjct: 303 LPVQVASRKLLQMISERVKEPL 368
>TC84406 similar to GP|12039327|gb|AAG46115.1 putative sugar transporter
{Oryza sativa}, partial (13%)
Length = 753
Score = 25.4 bits (54), Expect = 8.4
Identities = 13/40 (32%), Positives = 22/40 (54%)
Frame = -1
Query: 79 QLLNMKSEPEKDPLELVPDDENEHYVDSKLMSKPMEEEEL 118
Q+LN ++PL L + EH ++ L +P+EE E+
Sbjct: 600 QILN*CLNLIEEPLLLPAQESEEHSIEIHLCQQPVEEFEV 481
>TC81156 similar to PIR|T45754|T45754 hypothetical protein F24M12.270 -
Arabidopsis thaliana, partial (30%)
Length = 1204
Score = 25.4 bits (54), Expect = 8.4
Identities = 26/81 (32%), Positives = 38/81 (46%), Gaps = 7/81 (8%)
Frame = +1
Query: 62 LPYAPRSFELPIVVGSH---QLLNMKS-EPEKD---PLELVPDDENEHYVDSKLMSKPME 114
LP SF VV Q+LN +S E EK+ PL L D++E DSK+ S +
Sbjct: 475 LPLVSLSFSPSSVVQEQLIPQVLNQESVESEKNVLSPLLLDFSDKSEVTEDSKMGSTEDD 654
Query: 115 EEELPPLEGELWNWEDPPDDE 135
+ + ++ +D DDE
Sbjct: 655 GDSIWSIQVNASTHDDEDDDE 717
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.317 0.137 0.434
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 5,094,006
Number of Sequences: 36976
Number of extensions: 69620
Number of successful extensions: 351
Number of sequences better than 10.0: 14
Number of HSP's better than 10.0 without gapping: 350
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 351
length of query: 145
length of database: 9,014,727
effective HSP length: 87
effective length of query: 58
effective length of database: 5,797,815
effective search space: 336273270
effective search space used: 336273270
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 53 (25.0 bits)
Lotus: description of TM0026.15


