

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0026.14
(597 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
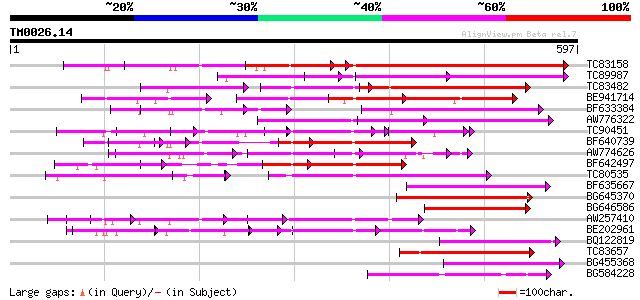
TC83158 weakly similar to PIR|H86191|H86191 hypothetical protein... 253 2e-67
TC89987 weakly similar to PIR|D86443|D86443 probable PPR-repeat ... 170 1e-54
TC83482 weakly similar to GP|9758453|dbj|BAB08982.1 selenium-bin... 159 4e-39
BE941714 weakly similar to GP|11994734|dbj selenium-binding prot... 156 2e-38
BF633384 similar to PIR|T46179|T46 hypothetical protein T8H10.30... 154 1e-37
AW776322 weakly similar to PIR|H84859|H84 hypothetical protein A... 144 1e-34
TC90451 similar to PIR|H84508|H84508 hypothetical protein At2g13... 97 1e-33
BF640739 weakly similar to GP|7363286|dbj Similar to Arabidopsis... 126 2e-29
AW774626 similar to PIR|T47909|T47 hypothetical protein T20K12.7... 122 5e-28
BF642497 weakly similar to PIR|H96713|H96 hypothetical protein T... 117 1e-26
TC80535 weakly similar to GP|14335136|gb|AAK59848.1 At2g15690/F9... 117 2e-26
BF635667 weakly similar to PIR|C85359|C853 hypothetical protein ... 114 1e-25
BG645370 weakly similar to PIR|T00405|T004 hypothetical protein ... 109 3e-24
BG646586 weakly similar to GP|10177683|dbj selenium-binding prot... 102 5e-22
AW257410 weakly similar to GP|10177533|db selenium-binding prote... 96 4e-20
BE202961 weakly similar to GP|6016735|gb|A hypothetical protein ... 96 5e-20
BQ122819 similar to PIR|C84748|C84 hypothetical protein At2g3368... 94 1e-19
TC83657 weakly similar to GP|11994382|dbj|BAB02341. gb|AAB82628.... 94 1e-19
BG455368 weakly similar to GP|9759524|dbj selenium-binding prote... 93 3e-19
BG584228 weakly similar to PIR|F71411|F71 hypothetical protein -... 81 1e-15
>TC83158 weakly similar to PIR|H86191|H86191 hypothetical protein [imported]
- Arabidopsis thaliana, partial (42%)
Length = 1194
Score = 253 bits (645), Expect = 2e-67
Identities = 128/340 (37%), Positives = 217/340 (63%)
Frame = +2
Query: 249 GELNRAEKLFAELPQKDVITWTSMMTGYAQHGLSEEALKMFTKMQANGGLKPNNGTFVTV 308
G+++ A KLF +LP K+V++WT ++ G+ + EEAL+ F +MQ G+ P+ T + +
Sbjct: 2 GDVDDALKLFDKLPVKNVVSWTVVIGGFVKKECYEEALECFREMQL-AGVVPDFVTVIAI 178
Query: 309 LGACSGLASLTEGQQIHQLISKTGFQENTRVVSALINMYSKCGELHIARKIFDDGLLRQR 368
+ AC+ L +L G +H+L+ K F++N +V+++LI+MY++CG + +AR++FD + QR
Sbjct: 179 ISACANLGALGLGLWVHRLVMKKEFRDNVKVLNSLIDMYARCGCIELARQVFDG--MSQR 352
Query: 369 DLISWNGMIAAYAHHGYGNEAINLFNKMQELGFQANDVTYVELLTACSHAGLVDEGIQYF 428
+L+SWN +I +A +G ++A++ F M++ G + N V+Y LTACSHAGL+DEG++ F
Sbjct: 353 NLVSWNSIIVGFAVNGLADKALSFFRSMKKEGLEPNGVSYTSALTACSHAGLIDEGLKIF 532
Query: 429 DKLLKNRSIQVKEDHYACLVDLCGRAGRLKEAFYIIEGLGVKLSLSVWGPLLAGCNVHGN 488
+ ++ + +HY CLVDL RAGRLKEA+ +I+ + + + V G LLA C G+
Sbjct: 533 ADIKRDHRNSPRIEHYGCLVDLYSRAGRLKEAWDVIKKMPMMPNEVVLGSLLAACRTQGD 712
Query: 489 ADIGKLVAKKILKVEHENAGTYSLLSNMYASVGKWKEAANVRMKMKDKGLKKQPGCSWIE 548
++ + V K +++ Y L SN+YA+VGKW A+ VR +MK++GL+K S IE
Sbjct: 713 VELAEKVMKYQVELYPGGDSNYVLFSNIYAAVGKWDGASKVRREMKERGLQKNLAFSSIE 892
Query: 549 VGNTVQVFVVGDKSHSQSEMLEYLLLGLHTKMKKFGDILD 588
+ + + FV GDK H +++ + L L ++ +G + D
Sbjct: 893 IDSGIHKFVSGDKYHEENDYIYSALELLSFELHLYGYVPD 1012
Score = 91.7 bits (226), Expect = 7e-19
Identities = 65/247 (26%), Positives = 124/247 (49%), Gaps = 9/247 (3%)
Frame = +2
Query: 122 IEEAESLFYEMPERNELSWNIMIGGYGQNGQIEKALDLFRRMPKRNLVSWNI----IIKA 177
+++A LF ++P +N +SW ++IGG+ + E+AL+ FR M +V + II A
Sbjct: 8 VDDALKLFDKLPVKNVVSWTVVIGGFVKKECYEEALECFREMQLAGVVPDFVTVIAIISA 187
Query: 178 LARCGRIEDAQW-HLIRCRKGMMPLRNVVSWNAMITGYAQNRRLDEALELFERMPERDMA 236
A G + W H + +K NV N++I YA+ ++ A ++F+ M +R++
Sbjct: 188 CANLGALGLGLWVHRLVMKKEFRD--NVKVLNSLIDMYARCGCIELARQVFDGMSQRNLV 361
Query: 237 SWNAMLTGFFQNGELNRAEKLFAELPQKDV----ITWTSMMTGYAQHGLSEEALKMFTKM 292
SWN+++ GF NG ++A F + ++ + +++TS +T + GL +E LK+F +
Sbjct: 362 SWNSIIVGFAVNGLADKALSFFRSMKKEGLEPNGVSYTSALTACSHAGLIDEGLKIFADI 541
Query: 293 QANGGLKPNNGTFVTVLGACSGLASLTEGQQIHQLISKTGFQENTRVVSALINMYSKCGE 352
+ + P + ++ S L E + I K N V+ +L+ G+
Sbjct: 542 KRDHRNSPRIEHYGCLVDLYSRAGRLKEAWDV---IKKMPMMPNEVVLGSLLAACRTQGD 712
Query: 353 LHIARKI 359
+ +A K+
Sbjct: 713 VELAEKV 733
Score = 77.0 bits (188), Expect = 2e-14
Identities = 80/333 (24%), Positives = 137/333 (41%), Gaps = 46/333 (13%)
Frame = +2
Query: 57 GRTDHARKLFDQMPQRDKHLWTTMIDGYIKCGAIKEARKLFDQ-------PDA------- 102
G D A KLFD++P ++ WT +I G++K +EA + F + PD
Sbjct: 2 GDVDDALKLFDKLPVKNVVSWTVVIGGFVKKECYEEALECFREMQLAGVVPDFVTVIAII 181
Query: 103 ------------------------KKSVVTWTTMVNGYVKLNQIEEAESLFYEMPERNEL 138
+ +V ++++ Y + IE A +F M +RN +
Sbjct: 182 SACANLGALGLGLWVHRLVMKKEFRDNVKVLNSLIDMYARCGCIELARQVFDGMSQRNLV 361
Query: 139 SWNIMIGGYGQNGQIEKALDLFRRMPKRNL----VSWNIIIKALARCGRIEDAQWHLIRC 194
SWN +I G+ NG +KAL FR M K L VS+ + A + G I++
Sbjct: 362 SWNSIIVGFAVNGLADKALSFFRSMKKEGLEPNGVSYTSALTACSHAGLIDEGLKIFADI 541
Query: 195 RKGMMPLRNVVSWNAMITGYAQNRRLDEALELFERMPER-DMASWNAMLTGFFQNGELNR 253
++ + + ++ Y++ RL EA ++ ++MP + ++L G++
Sbjct: 542 KRDHRNSPRIEHYGCLVDLYSRAGRLKEAWDVIKKMPMMPNEVVLGSLLAACRTQGDVEL 721
Query: 254 AEKLF---AELPQKDVITWTSMMTGYAQHGLSEEALKMFTKMQANGGLKPNNGTFVTVLG 310
AEK+ EL + YA G + A K+ +M+ G K N F ++
Sbjct: 722 AEKVMKYQVELYPGGDSNYVLFSNIYAAVGKWDGASKVRREMKERGLQK--NLAFSSI-E 892
Query: 311 ACSGLASLTEGQQIHQLISKTGFQENTRVVSAL 343
SG+ G + H +EN + SAL
Sbjct: 893 IDSGIHKFVSGDKYH--------EENDYIYSAL 967
>TC89987 weakly similar to PIR|D86443|D86443 probable PPR-repeat protein
[imported] - Arabidopsis thaliana, partial (62%)
Length = 1559
Score = 170 bits (430), Expect(2) = 1e-54
Identities = 86/224 (38%), Positives = 133/224 (58%)
Frame = +2
Query: 365 LRQRDLISWNGMIAAYAHHGYGNEAINLFNKMQELGFQANDVTYVELLTACSHAGLVDEG 424
+ +++ S+ MI+ A HG G EA+ +F++M E G +DV YV + +ACSHAGLV+EG
Sbjct: 593 MSEKNRYSYTVMISGLAIHGRGKEALKVFSEMIEEGLAPDDVVYVGVFSACSHAGLVEEG 772
Query: 425 IQYFDKLLKNRSIQVKEDHYACLVDLCGRAGRLKEAFYIIEGLGVKLSLSVWGPLLAGCN 484
+Q F + I+ HY C+VDL GR G LKEA+ +I+ + +K + +W LL+ C
Sbjct: 773 LQCFKSMQFEHKIEPTVQHYGCMVDLLGRFGMLKEAYELIKSMSIKPNDVIWRSLLSACK 952
Query: 485 VHGNADIGKLVAKKILKVEHENAGTYSLLSNMYASVGKWKEAANVRMKMKDKGLKKQPGC 544
VH N +IGK+ A+ + + N+G Y +L+NMYA KW + A +R K+ ++ L + PG
Sbjct: 953 VHHNLEIGKIAAENLFMLNQNNSGDYLVLANMYAKAQKWDDVAKIRTKLAERNLVQTPGF 1132
Query: 545 SWIEVGNTVQVFVVGDKSHSQSEMLEYLLLGLHTKMKKFGDILD 588
S IE V FV DKS Q ++ ++ + ++K G I D
Sbjct: 1133SLIEAKRKVYKFVSQDKSIPQWNIIYEMIHQMEWQVKFEGYIPD 1264
Score = 77.8 bits (190), Expect = 1e-14
Identities = 52/156 (33%), Positives = 85/156 (54%), Gaps = 1/156 (0%)
Frame = +1
Query: 311 ACSGLASLTEGQQIHQLISKTGFQENTRVVSALINMYSKCGELHIARKIFDDGLLRQRDL 370
ACS L + EG Q+H + K G + + V ++LINMY KCGE+ A +F+ + ++ +
Sbjct: 130 ACSLLGVVDEGIQVHGHVFKMGLEGDVIVQNSLINMYGKCGEIKNACDVFNG--MDEKSV 303
Query: 371 ISWNGMIAAYAHHGYGNEAINLFNKMQELG-FQANDVTYVELLTACSHAGLVDEGIQYFD 429
SW+ +I A+A NE + L KM G + + T V +L+AC+H G D G
Sbjct: 304 ASWSAIIGAHACVEMWNECLMLLGKMSSEGRCRVEESTLVNVLSACTHLGSPDLGKCIHG 483
Query: 430 KLLKNRSIQVKEDHYACLVDLCGRAGRLKEAFYIIE 465
LL+N S ++ L+D+ ++G L++ F I+
Sbjct: 484 ILLRNIS-ELNVVVKTSLIDMYVKSGCLEKRFTRIQ 588
Score = 62.0 bits (149), Expect(2) = 1e-54
Identities = 38/136 (27%), Positives = 72/136 (52%), Gaps = 4/136 (2%)
Frame = +1
Query: 220 LDEALEL----FERMPERDMASWNAMLTGFFQNGELNRAEKLFAELPQKDVITWTSMMTG 275
+DE +++ F+ E D+ N+++ + + GE+ A +F + +K V +W++++
Sbjct: 151 VDEGIQVHGHVFKMGLEGDVIVQNSLINMYGKCGEIKNACDVFNGMDEKSVASWSAIIGA 330
Query: 276 YAQHGLSEEALKMFTKMQANGGLKPNNGTFVTVLGACSGLASLTEGQQIHQLISKTGFQE 335
+A + E L + KM + G + T V VL AC+ L S G+ IH ++ + +
Sbjct: 331 HACVEMWNECLMLLGKMSSEGRCRVEESTLVNVLSACTHLGSPDLGKCIHGILLRNISEL 510
Query: 336 NTRVVSALINMYSKCG 351
N V ++LI+MY K G
Sbjct: 511 NVVVKTSLIDMYVKSG 558
Score = 35.4 bits (80), Expect = 0.063
Identities = 16/39 (41%), Positives = 26/39 (66%)
Frame = +1
Query: 141 NIMIGGYGQNGQIEKALDLFRRMPKRNLVSWNIIIKALA 179
N +I YG+ G+I+ A D+F M ++++ SW+ II A A
Sbjct: 220 NSLINMYGKCGEIKNACDVFNGMDEKSVASWSAIIGAHA 336
Score = 28.1 bits (61), Expect = 10.0
Identities = 21/94 (22%), Positives = 45/94 (47%), Gaps = 4/94 (4%)
Frame = +2
Query: 80 MIDGYIKCGAIKEARKLFDQPDAKKSVVTWTTMVNGYVKLNQIE----EAESLFYEMPER 135
M+D + G +KEA +L K + V W ++++ + +E AE+LF + +
Sbjct: 839 MVDLLGRFGMLKEAYELIKSMSIKPNDVIWRSLLSACKVHHNLEIGKIAAENLFM-LNQN 1015
Query: 136 NELSWNIMIGGYGQNGQIEKALDLFRRMPKRNLV 169
N + ++ Y + + + + ++ +RNLV
Sbjct: 1016NSGDYLVLANMYAKAQKWDDVAKIRTKLAERNLV 1117
>TC83482 weakly similar to GP|9758453|dbj|BAB08982.1 selenium-binding
protein-like {Arabidopsis thaliana}, partial (10%)
Length = 895
Score = 159 bits (401), Expect = 4e-39
Identities = 73/180 (40%), Positives = 116/180 (63%)
Frame = +2
Query: 369 DLISWNGMIAAYAHHGYGNEAINLFNKMQELGFQANDVTYVELLTACSHAGLVDEGIQYF 428
DL+SWN +I YA HG A+ F++M+ +G ++VT+V +L+AC HAGLV+EG ++F
Sbjct: 2 DLVSWNAIIGGYASHGLATRALEEFDRMKVVG-TPDEVTFVNVLSACVHAGLVEEGEKHF 178
Query: 429 DKLLKNRSIQVKEDHYACLVDLCGRAGRLKEAFYIIEGLGVKLSLSVWGPLLAGCNVHGN 488
+L IQ + +HY+C+VDL GRAGR EA +I+ + + + +WG LLA C +H N
Sbjct: 179 TDMLTKYGIQAEMEHYSCMVDLYGRAGRFDEAENLIKNMPFEPDVVLWGALLAACGLHSN 358
Query: 489 ADIGKLVAKKILKVEHENAGTYSLLSNMYASVGKWKEAANVRMKMKDKGLKKQPGCSWIE 548
++G+ A++I ++E + +YS+LS + G W +R MK++G+KKQ SW+E
Sbjct: 359 LELGEYAAERIRRLESSHPVSYSVLSKIQGEKGVWSSVNELRDTMKERGIKKQTAISWVE 538
Score = 66.6 bits (161), Expect = 3e-11
Identities = 35/120 (29%), Positives = 65/120 (54%), Gaps = 1/120 (0%)
Frame = +2
Query: 265 DVITWTSMMTGYAQHGLSEEALKMFTKMQANGGLKPNNGTFVTVLGACSGLASLTEGQQ- 323
D+++W +++ GYA HGL+ AL+ F +M+ G P+ TFV VL AC + EG++
Sbjct: 2 DLVSWNAIIGGYASHGLATRALEEFDRMKVVG--TPDEVTFVNVLSACVHAGLVEEGEKH 175
Query: 324 IHQLISKTGFQENTRVVSALINMYSKCGELHIARKIFDDGLLRQRDLISWNGMIAAYAHH 383
+++K G Q S ++++Y + G A + + + + D++ W ++AA H
Sbjct: 176 FTDMLTKYGIQAEMEHYSCMVDLYGRAGRFDEAENLIKN-MPFEPDVVLWGALLAACGLH 352
Score = 52.0 bits (123), Expect = 6e-07
Identities = 36/120 (30%), Positives = 57/120 (47%), Gaps = 6/120 (5%)
Frame = +2
Query: 138 LSWNIMIGGYGQNGQIEKALDLFRRMP---KRNLVSWNIIIKALARCGRIEDAQWHLIRC 194
+SWN +IGGY +G +AL+ F RM + V++ ++ A G +E+ + H
Sbjct: 8 VSWNAIIGGYASHGLATRALEEFDRMKVVGTPDEVTFVNVLSACVHAGLVEEGEKHFTDM 187
Query: 195 RKGMMPLRNVVSWNAMITGYAQNRRLDEALELFERMP-ERDMASWNAMLT--GFFQNGEL 251
+ ++ M+ Y + R DEA L + MP E D+ W A+L G N EL
Sbjct: 188 LTKYGIQAEMEHYSCMVDLYGRAGRFDEAENLIKNMPFEPDVVLWGALLAACGLHSNLEL 367
Score = 40.4 bits (93), Expect = 0.002
Identities = 35/136 (25%), Positives = 64/136 (46%), Gaps = 8/136 (5%)
Frame = +2
Query: 203 NVVSWNAMITGYAQNRRLDEALELFERMP---ERDMASWNAMLTGFFQNGELNRAEKLFA 259
++VSWNA+I GYA + ALE F+RM D ++ +L+ G + EK F
Sbjct: 2 DLVSWNAIIGGYASHGLATRALEEFDRMKVVGTPDEVTFVNVLSACVHAGLVEEGEKHFT 181
Query: 260 EL-----PQKDVITWTSMMTGYAQHGLSEEALKMFTKMQANGGLKPNNGTFVTVLGACSG 314
++ Q ++ ++ M+ Y + G +EA + M +P+ + +L AC
Sbjct: 182 DMLTKYGIQAEMEHYSCMVDLYGRAGRFDEAENLIKNMP----FEPDVVLWGALLAACGL 349
Query: 315 LASLTEGQQIHQLISK 330
++L G+ + I +
Sbjct: 350 HSNLELGEYAAERIRR 397
Score = 30.8 bits (68), Expect = 1.5
Identities = 30/138 (21%), Positives = 61/138 (43%), Gaps = 9/138 (6%)
Frame = +2
Query: 167 NLVSWNIIIKALARCGRIEDAQWHLIRCRKGMMPLRNVVSWNAMITGYAQNRRLDEALEL 226
+LVSWN II A G A R + P + V++ +++ ++E +
Sbjct: 2 DLVSWNAIIGGYASHGLATRALEEFDRMKVVGTP--DEVTFVNVLSACVHAGLVEEGEKH 175
Query: 227 FERM-----PERDMASWNAMLTGFFQNGELNRAEKLFAELP-QKDVITWTSMMTGYAQHG 280
F M + +M ++ M+ + + G + AE L +P + DV+ W +++ H
Sbjct: 176 FTDMLTKYGIQAEMEHYSCMVDLYGRAGRFDEAENLIKNMPFEPDVVLWGALLAACGLHS 355
Query: 281 ---LSEEALKMFTKMQAN 295
L E A + +++++
Sbjct: 356 NLELGEYAAERIRRLESS 409
>BE941714 weakly similar to GP|11994734|dbj selenium-binding protein-like
{Arabidopsis thaliana}, partial (10%)
Length = 619
Score = 156 bits (394), Expect = 2e-38
Identities = 78/205 (38%), Positives = 130/205 (63%), Gaps = 6/205 (2%)
Frame = +2
Query: 336 NTRVVSALINMYSKCGELHIARKIFDDGLLRQRDLISWNGMIAAYAHHGYGNEAINLFNK 395
+ R+ ++L++MY+KCG L +A+++FD + RD++ WN MI+ A HG G A+ LF
Sbjct: 11 SVRLSTSLLDMYAKCGNLELAKRLFDS--MNMRDVVCWNAMISGMAMHGDGKGALKLFYD 184
Query: 396 MQELGFQANDVTYVELLTACSHAGLVDEGIQYFDKLLKNRSIQVKEDHYACLVDLCGRAG 455
M+++G + +D+T++ + TACS++G+ EG+ DK+ +I K +HY CLVDL RAG
Sbjct: 185 MEKVGVKPDDITFIAVFTACSYSGMAYEGLMLLDKMCSVYNIVPKSEHYGCLVDLLSRAG 364
Query: 456 RLKEAFYIIEGL-----GVKLSLSVWGPLLAGCNVHGNADIGKLVAKKILKVE-HENAGT 509
+EA +I + G + +L+ W L+ C HG + +L A+K+L+++ H ++G
Sbjct: 365 LFEEAMVMIRKITNSWNGSEETLA-WRAFLSACCNHGETQLAELAAEKVLQLDNHIHSGV 541
Query: 510 YSLLSNMYASVGKWKEAANVRMKMK 534
Y LLSN+YA+ GK +A V+ MK
Sbjct: 542 YVLLSNLYAASGKHNDARRVKDMMK 616
Score = 66.2 bits (160), Expect = 3e-11
Identities = 46/184 (25%), Positives = 87/184 (47%), Gaps = 4/184 (2%)
Frame = +2
Query: 240 AMLTGFFQNGELNRAEKLFAELPQKDVITWTSMMTGYAQHGLSEEALKMFTKMQANGGLK 299
++L + + G L A++LF + +DV+ W +M++G A HG + ALK+F M+ G+K
Sbjct: 29 SLLDMYAKCGNLELAKRLFDSMNMRDVVCWNAMISGMAMHGDGKGALKLFYDME-KVGVK 205
Query: 300 PNNGTFVTVLGACSGLASLTEG-QQIHQLISKTGFQENTRVVSALINMYSKCG---ELHI 355
P++ TF+ V ACS EG + ++ S + L+++ S+ G E +
Sbjct: 206 PDDITFIAVFTACSYSGMAYEGLMLLDKMCSVYNIVPKSEHYGCLVDLLSRAGLFEEAMV 385
Query: 356 ARKIFDDGLLRQRDLISWNGMIAAYAHHGYGNEAINLFNKMQELGFQANDVTYVELLTAC 415
+ + + ++W ++A +HG A K+ +L + YV L
Sbjct: 386 MIRKITNSWNGSEETLAWRAFLSACCNHGETQLAELAAEKVLQLDNHIHSGVYVLLSNLY 565
Query: 416 SHAG 419
+ +G
Sbjct: 566 AASG 577
Score = 47.4 bits (111), Expect = 2e-05
Identities = 32/148 (21%), Positives = 69/148 (46%), Gaps = 11/148 (7%)
Frame = +2
Query: 76 LWTTMIDGYIKCGAIKEARKLFDQPDAKKSVVTWTTMVNGYVKLNQIEEAESLFYEMP-- 133
L T+++D Y KCG ++ A++LFD + + VV W M++G + A LFY+M
Sbjct: 20 LSTSLLDMYAKCGNLELAKRLFDSMN-MRDVVCWNAMISGMAMHGDGKGALKLFYDMEKV 196
Query: 134 --ERNELSWNIMIGGYGQNGQ-------IEKALDLFRRMPKRNLVSWNIIIKALARCGRI 184
+ +++++ + +G ++K ++ +PK + ++ L+R G
Sbjct: 197 GVKPDDITFIAVFTACSYSGMAYEGLMLLDKMCSVYNIVPKSE--HYGCLVDLLSRAGLF 370
Query: 185 EDAQWHLIRCRKGMMPLRNVVSWNAMIT 212
E+A + + ++W A ++
Sbjct: 371 EEAMVMIRKITNSWNGSEETLAWRAFLS 454
Score = 41.2 bits (95), Expect = 0.001
Identities = 45/199 (22%), Positives = 88/199 (43%), Gaps = 11/199 (5%)
Frame = +2
Query: 174 IIKALARCGRIEDAQWHLIRCRKGMMPLRNVVSWNAMITGYAQNRRLDEALELFERMP-- 231
++ A+CG +E L + M +R+VV WNAMI+G A + AL+LF M
Sbjct: 32 LLDMYAKCGNLE-----LAKRLFDSMNMRDVVCWNAMISGMAMHGDGKGALKLFYDMEKV 196
Query: 232 --ERDMASWNAMLTGFFQNGE-------LNRAEKLFAELPQKDVITWTSMMTGYAQHGLS 282
+ D ++ A+ T +G L++ ++ +P+ + + ++ ++ GL
Sbjct: 197 GVKPDDITFIAVFTACSYSGMAYEGLMLLDKMCSVYNIVPKSE--HYGCLVDLLSRAGLF 370
Query: 283 EEALKMFTKMQANGGLKPNNGTFVTVLGACSGLASLTEGQQIHQLISKTGFQENTRVVSA 342
EEA+ M K+ + + L AC + + + + ++ V
Sbjct: 371 EEAMVMIRKITNSWNGSEETLAWRAFLSACCNHGETQLAELAAEKVLQLDNHIHSGVYVL 550
Query: 343 LINMYSKCGELHIARKIFD 361
L N+Y+ G+ + AR++ D
Sbjct: 551 LSNLYAASGKHNDARRVKD 607
Score = 36.2 bits (82), Expect = 0.037
Identities = 32/128 (25%), Positives = 56/128 (43%), Gaps = 10/128 (7%)
Frame = +2
Query: 55 KEGRTDHARKLFDQMPQRDKHLWTTMIDGYIKCGAIKEARKLF---DQPDAKKSVVTWTT 111
K G + A++LFD M RD W MI G G K A KLF ++ K +T+
Sbjct: 50 KCGNLELAKRLFDSMNMRDVVCWNAMISGMAMHGDGKGALKLFYDMEKVGVKPDDITFIA 229
Query: 112 MVN-------GYVKLNQIEEAESLFYEMPERNELSWNIMIGGYGQNGQIEKALDLFRRMP 164
+ Y L +++ S++ +P+ + ++ + G E+A+ + R++
Sbjct: 230 VFTACSYSGMAYEGLMLLDKMCSVYNIVPKSEH--YGCLVDLLSRAGLFEEAMVMIRKIT 403
Query: 165 KRNLVSWN 172
SWN
Sbjct: 404 N----SWN 415
>BF633384 similar to PIR|T46179|T46 hypothetical protein T8H10.30 -
Arabidopsis thaliana, partial (22%)
Length = 595
Score = 154 bits (388), Expect = 1e-37
Identities = 77/198 (38%), Positives = 117/198 (58%), Gaps = 6/198 (3%)
Frame = +1
Query: 371 ISWNGMIAAYAHHGYGNEAINLFNKMQELG-----FQANDVTYVELLTACSHAGLVDEGI 425
I+WN +I AY HG G EA+ LF +M E G + N+VTY+ + + SH+G+VDEG+
Sbjct: 1 ITWNVLIMAYGMHGKGEEALKLFRRMVEEGDNNREIRPNEVTYIAIFASLSHSGMVDEGL 180
Query: 426 QYFDKLLKNRSIQVKEDHYACLVDLCGRAGRLKEAFYIIEGLGVKLS-LSVWGPLLAGCN 484
F + I+ DHYACLVDL GR+G+++EA+ +I+ + + + W LL C
Sbjct: 181 NLFYTMKAKHGIEPTSDHYACLVDLLGRSGQIEEAYNLIKTMPSNMKKVDAWSSLLGACK 360
Query: 485 VHGNADIGKLVAKKILKVEHENAGTYSLLSNMYASVGKWKEAANVRMKMKDKGLKKQPGC 544
+H N +IG++ AK + ++ A Y LLSN+Y+S G W +A +VR KMK+KG++K
Sbjct: 361 IHQNLEIGEIAAKNLFVLDPNVASYYVLLSNIYSSAGLWDQAIDVRKKMKEKGVRKNLDV 540
Query: 545 SWIEVGNTVQVFVVGDKS 562
+ + F GD S
Sbjct: 541 VGLNMAMRFTSFWQGDVS 594
Score = 53.5 bits (127), Expect = 2e-07
Identities = 43/177 (24%), Positives = 94/177 (52%), Gaps = 18/177 (10%)
Frame = +1
Query: 138 LSWNIMIGGYGQNGQIEKALDLFRRMPKR---------NLVSWNIIIKALARCGRIEDA- 187
++WN++I YG +G+ E+AL LFRRM + N V++ I +L+ G +++
Sbjct: 1 ITWNVLIMAYGMHGKGEEALKLFRRMVEEGDNNREIRPNEVTYIAIFASLSHSGMVDEGL 180
Query: 188 -QWHLIRCRKGMMPLRNVVSWNAMITGYAQNRRLDEALELFERMPE--RDMASWNAMLTG 244
++ ++ + G+ P + + ++ ++ +++EA L + MP + + +W+++L
Sbjct: 181 NLFYTMKAKHGIEPTSD--HYACLVDLLGRSGQIEEAYNLIKTMPSNMKKVDAWSSLLGA 354
Query: 245 --FFQNGELNR--AEKLFAELPQKDVITWTSMMTG-YAQHGLSEEALKMFTKMQANG 296
QN E+ A+ LF P +V ++ +++ Y+ GL ++A+ + KM+ G
Sbjct: 355 CKIHQNLEIGEIAAKNLFVLDP--NVASYYVLLSNIYSSAGLWDQAIDVRKKMKEKG 519
Score = 42.4 bits (98), Expect = 5e-04
Identities = 37/163 (22%), Positives = 77/163 (46%), Gaps = 18/163 (11%)
Frame = +1
Query: 107 VTWTTMVNGYVKLNQIEEAESLFYEMPER---------NELSWNIMIGGYGQNGQIEKAL 157
+TW ++ Y + EEA LF M E NE+++ + +G +++ L
Sbjct: 1 ITWNVLIMAYGMHGKGEEALKLFRRMVEEGDNNREIRPNEVTYIAIFASLSHSGMVDEGL 180
Query: 158 DLFRRMPKRNLVS-----WNIIIKALARCGRIEDAQWHLIRCRKGMMPLRNVVSWNAMIT 212
+LF M ++ + + ++ L R G+IE+A ++LI+ M + V +W++++
Sbjct: 181 NLFYTMKAKHGIEPTSDHYACLVDLLGRSGQIEEA-YNLIKTMPSNM--KKVDAWSSLLG 351
Query: 213 GYAQNRRLD----EALELFERMPERDMASWNAMLTGFFQNGEL 251
++ L+ A LF P ++AS+ +L+ + + L
Sbjct: 352 ACKIHQNLEIGEIAAKNLFVLDP--NVASYYVLLSNIYSSAGL 474
>AW776322 weakly similar to PIR|H84859|H84 hypothetical protein At2g42920
[imported] - Arabidopsis thaliana, partial (27%)
Length = 633
Score = 144 bits (362), Expect = 1e-34
Identities = 69/207 (33%), Positives = 122/207 (58%), Gaps = 1/207 (0%)
Frame = +1
Query: 367 QRDLISWNGMIAAYAHHGYGNEAINLFNKMQELGFQAND-VTYVELLTACSHAGLVDEGI 425
+R L WN +I A +G+ EA F+K++ D V+++ +LTAC H G +++
Sbjct: 13 RRGLSCWNSIIIGLAMNGHEREAFEFFSKLESSKLLKPDSVSFIGVLTACKHLGAINKAR 192
Query: 426 QYFDKLLKNRSIQVKEDHYACLVDLCGRAGRLKEAFYIIEGLGVKLSLSVWGPLLAGCNV 485
YF+ ++ I+ HY C+VD+ G+AG L+EA +I+G+ +K +WG LL+ C
Sbjct: 193 DYFELMMNKYEIEPSIKHYTCIVDVLGQAGLLEEAEELIKGMPLKPDAIIWGSLLSSCRK 372
Query: 486 HGNADIGKLVAKKILKVEHENAGTYSLLSNMYASVGKWKEAANVRMKMKDKGLKKQPGCS 545
H N I + A+++ ++ +A Y L+SN++A+ K++EA R+ MK+ +K+PGCS
Sbjct: 373 HRNVQIARRAAQRVYELNPSDASGYVLMSNVHAASNKFEEAIEQRLLMKENLTEKEPGCS 552
Query: 546 WIEVGNTVQVFVVGDKSHSQSEMLEYL 572
IE+ V F+ G + H +++ + +L
Sbjct: 553 SIELYGEVHEFIAGGRLHPKTQEIYHL 633
Score = 58.5 bits (140), Expect = 7e-09
Identities = 40/181 (22%), Positives = 92/181 (50%), Gaps = 1/181 (0%)
Frame = +1
Query: 262 PQKDVITWTSMMTGYAQHGLSEEALKMFTKMQANGGLKPNNGTFVTVLGACSGLASLTEG 321
P++ + W S++ G A +G EA + F+K++++ LKP++ +F+ VL AC L ++ +
Sbjct: 10 PRRGLSCWNSIIIGLAMNGHEREAFEFFSKLESSKLLKPDSVSFIGVLTACKHLGAINKA 189
Query: 322 QQIHQL-ISKTGFQENTRVVSALINMYSKCGELHIARKIFDDGLLRQRDLISWNGMIAAY 380
+ +L ++K + + + + ++++ + G L A ++ G+ + D I W ++++
Sbjct: 190 RDYFELMMNKYEIEPSIKHYTCIVDVLGQAGLLEEAEELI-KGMPLKPDAIIWGSLLSSC 366
Query: 381 AHHGYGNEAINLFNKMQELGFQANDVTYVELLTACSHAGLVDEGIQYFDKLLKNRSIQVK 440
H A ++ EL ++ YV + + + +E I+ +LL ++ K
Sbjct: 367 RKHRNVQIARRAAQRVYELN-PSDASGYVLMSNVHAASNKFEEAIE--QRLLMKENLTEK 537
Query: 441 E 441
E
Sbjct: 538 E 540
Score = 40.8 bits (94), Expect = 0.001
Identities = 32/122 (26%), Positives = 54/122 (44%), Gaps = 6/122 (4%)
Frame = +1
Query: 164 PKRNLVSWNIIIKALARCGRIEDAQWHLIRCRKGMMPLRNVVSWNAMITGYAQNRRLDEA 223
P+R L WN II LA G +A + + + VS+ ++T +++A
Sbjct: 10 PRRGLSCWNSIIIGLAMNGHEREAFEFFSKLESSKLLKPDSVSFIGVLTACKHLGAINKA 189
Query: 224 LELFERMP-----ERDMASWNAMLTGFFQNGELNRAEKLFAELPQK-DVITWTSMMTGYA 277
+ FE M E + + ++ Q G L AE+L +P K D I W S+++
Sbjct: 190 RDYFELMMNKYEIEPSIKHYTCIVDVLGQAGLLEEAEELIKGMPLKPDAIIWGSLLSSCR 369
Query: 278 QH 279
+H
Sbjct: 370 KH 375
>TC90451 similar to PIR|H84508|H84508 hypothetical protein At2g13600
[imported] - Arabidopsis thaliana, partial (12%)
Length = 1362
Score = 97.1 bits (240), Expect = 2e-20
Identities = 63/227 (27%), Positives = 110/227 (47%), Gaps = 6/227 (2%)
Frame = +2
Query: 269 WTSMMTGYAQHGLSEEALKMFTKMQANGGLKPNNGTFVTVLGACSGLASLTEGQQIHQLI 328
W +++ Y + + AL+++ M G+ P+ T VL A S ++ GQQ+H
Sbjct: 389 WNNIIRSYTRLESPQNALRIYVSM-LRAGVLPDRYTLPIVLKAVSQSFAIQLGQQVHSYG 565
Query: 329 SKTGFQENTRVVSALINMYSKCGELHIARKIFDDGLLRQRDLISWNGMIAAYAHHGYGNE 388
K G Q N S IN+Y K G+ A K+FD+ + L SWN +I+ + G +
Sbjct: 566 IKLGLQSNEYCESGFINLYCKAGDFDSAHKVFDEN--HEPKLGSWNALISGLSQGGLAMD 739
Query: 389 AINLFNKMQELGFQANDVTYVELLTACSHAGLVDEGIQYFDKLLKNRSIQVKEDHYA--- 445
AI +F M+ GF+ + +T V +++AC G + Y L Q K + +
Sbjct: 740 AIVVFVDMKRHGFEPDGITMVSVMSACGSIGDL-----YLALQLHKYVFQAKTNEWTVIL 904
Query: 446 ---CLVDLCGRAGRLKEAFYIIEGLGVKLSLSVWGPLLAGCNVHGNA 489
L+D+ G+ GR+ A+ + + + ++S W ++ G +HG+A
Sbjct: 905 MSNSLIDMYGKCGRMDLAYEVFATMEDR-NVSSWTSMIVGYAMHGHA 1042
Score = 95.5 bits (236), Expect(2) = 1e-33
Identities = 62/238 (26%), Positives = 127/238 (53%), Gaps = 7/238 (2%)
Frame = +2
Query: 170 SWNIIIKALARCGRIEDA-QWHLIRCRKGMMPLRNVVSWNAMITGYAQNRRLDEALELFE 228
+WN II++ R ++A + ++ R G++P R + ++ +Q+ + ++
Sbjct: 386 NWNNIIRSYTRLESPQNALRIYVSMLRAGVLPDRYTLP--IVLKAVSQSFAIQLGQQVHS 559
Query: 229 RMPERDMASWNAMLTGFF----QNGELNRAEKLFAELPQKDVITWTSMMTGYAQHGLSEE 284
+ + S +GF + G+ + A K+F E + + +W ++++G +Q GL+ +
Sbjct: 560 YGIKLGLQSNEYCESGFINLYCKAGDFDSAHKVFDENHEPKLGSWNALISGLSQGGLAMD 739
Query: 285 ALKMFTKMQANGGLKPNNGTFVTVLGACSGLASLTEGQQIHQLISKTGFQENTRVV--SA 342
A+ +F M+ +G +P+ T V+V+ AC + L Q+H+ + + E T ++ ++
Sbjct: 740 AIVVFVDMKRHG-FEPDGITMVSVMSACGSIGDLYLALQLHKYVFQAKTNEWTVILMSNS 916
Query: 343 LINMYSKCGELHIARKIFDDGLLRQRDLISWNGMIAAYAHHGYGNEAINLFNKMQELG 400
LI+MY KCG + +A ++F + R++ SW MI YA HG+ EA+ F+ M+ +G
Sbjct: 917 LIDMYGKCGRMDLAYEVF--ATMEDRNVSSWTSMIVGYAMHGHAKEALGCFHCMKGVG 1084
Score = 66.2 bits (160), Expect(2) = 1e-33
Identities = 34/84 (40%), Positives = 49/84 (57%)
Frame = +1
Query: 400 GFQANDVTYVELLTACSHAGLVDEGIQYFDKLLKNRSIQVKEDHYACLVDLCGRAGRLKE 459
G + N VT++ +L+AC H G V EG YFD + I + HY C+VDL GRAG +
Sbjct: 1084 GVKPNYVTFIGVLSACVHGGTVQEGRFYFDMMKNIYGITPQLQHYGCMVDLLGRAGLFDD 1263
Query: 460 AFYIIEGLGVKLSLSVWGPLLAGC 483
A ++E + +K + VWG + GC
Sbjct: 1264 ARRMVEEMPMKPNSVVWG-VFDGC 1332
Score = 60.1 bits (144), Expect(2) = 1e-12
Identities = 48/191 (25%), Positives = 80/191 (41%), Gaps = 7/191 (3%)
Frame = +2
Query: 113 VNGYVKLNQIEEAESLFYEMPERNELSWNIMIGGYGQNGQIEKALDLFRRMPKRNLVSWN 172
+N Y K + A +F E E SWN +I G Q G A+ +F M +
Sbjct: 611 INLYCKAGDFDSAHKVFDENHEPKLGSWNALISGLSQGGLAMDAIVVFVDMKRHGFEPDG 790
Query: 173 I-IIKALARCGRIEDAQWHLIRCRKGMMPLRNVVSWNAMITGYAQNRRLDEALELFERMP 231
I ++ ++ CG I D L AL+L + +
Sbjct: 791 ITMVSVMSACGSIGD---------------------------------LYLALQLHKYVF 871
Query: 232 ERDMASW------NAMLTGFFQNGELNRAEKLFAELPQKDVITWTSMMTGYAQHGLSEEA 285
+ W N+++ + + G ++ A ++FA + ++V +WTSM+ GYA HG ++EA
Sbjct: 872 QAKTNEWTVILMSNSLIDMYGKCGRMDLAYEVFATMEDRNVSSWTSMIVGYAMHGHAKEA 1051
Query: 286 LKMFTKMQANG 296
L F M+ G
Sbjct: 1052LGCFHCMKGVG 1084
Score = 48.5 bits (114), Expect = 7e-06
Identities = 40/162 (24%), Positives = 70/162 (42%), Gaps = 13/162 (8%)
Frame = +2
Query: 50 ISRLCKEGRTDHARKLFDQMPQRDKHLWTTMIDGYIKCGAIKEARKLF-------DQPDA 102
I+ CK G D A K+FD+ + W +I G + G +A +F +PD
Sbjct: 611 INLYCKAGDFDSAHKVFDENHEPKLGSWNALISGLSQGGLAMDAIVVFVDMKRHGFEPDG 790
Query: 103 KKSVVTWTTMVNGYVKLNQIEEAESLFYEMPERNELSW------NIMIGGYGQNGQIEKA 156
+T ++++ + + A L + + W N +I YG+ G+++ A
Sbjct: 791 ----ITMVSVMSACGSIGDLYLALQLHKYVFQAKTNEWTVILMSNSLIDMYGKCGRMDLA 958
Query: 157 LDLFRRMPKRNLVSWNIIIKALARCGRIEDAQWHLIRCRKGM 198
++F M RN+ SW +I A G ++A C KG+
Sbjct: 959 YEVFATMEDRNVSSWTSMIVGYAMHGHAKEA-LGCFHCMKGV 1081
Score = 31.2 bits (69), Expect(2) = 1e-12
Identities = 20/97 (20%), Positives = 46/97 (46%), Gaps = 3/97 (3%)
Frame = +1
Query: 293 QANGGLKPNNGTFVTVLGACSGLASLTEGQQIHQLISKT-GFQENTRVVSALINMYSKCG 351
+ + G+KPN TF+ VL AC ++ EG+ ++ G + ++++ + G
Sbjct: 1072 EGSRGVKPNYVTFIGVLSACVHGGTVQEGRFYFDMMKNIYGITPQLQHYGCMVDLLGRAG 1251
Query: 352 ELHIARKIFDDGLLRQRDLI--SWNGMIAAYAHHGYG 386
AR++ ++ ++ ++ ++G + GYG
Sbjct: 1252 LFDDARRMVEEMPMKPNSVVWGVFDGCM*ETWKRGYG 1362
>BF640739 weakly similar to GP|7363286|dbj Similar to Arabidopsis thaliana
DNA chromosome 4 BAC clone F4I10; vegetative storage
protein., partial (14%)
Length = 444
Score = 126 bits (317), Expect = 2e-29
Identities = 61/146 (41%), Positives = 99/146 (67%), Gaps = 1/146 (0%)
Frame = +1
Query: 284 EALKMFTKMQANGGLKPNNGTFVTVLGACSGLASLTEGQQIHQLISKTGFQENTRVVSAL 343
EAL +F MQ+ G +KP++ TFV+V+ A + L+ + + IH L +T N V +AL
Sbjct: 7 EALNLFCTMQSQG-IKPDSFTFVSVITALADLSVTRQAKWIHGLAIRTNMDTNVFVATAL 183
Query: 344 INMYSKCGELHIARKIFDDGLLRQRDLISWNGMIAAYAHHGYGNEAINLFNKMQ-ELGFQ 402
++MY+KCG + AR++FD ++++R +I+WN MI Y HG G A++LF+ MQ E +
Sbjct: 184 VDMYAKCGAIETARELFD--MMQERHVITWNAMIDGYGTHGLGKAALDLFDDMQNEASLK 357
Query: 403 ANDVTYVELLTACSHAGLVDEGIQYF 428
ND+T++ +++ACSH+G V+EG+ YF
Sbjct: 358 PNDITFLSVISACSHSGFVEEGLYYF 435
Score = 67.8 bits (164), Expect = 1e-11
Identities = 52/174 (29%), Positives = 78/174 (43%), Gaps = 5/174 (2%)
Frame = +1
Query: 153 IEKALDLFRRMPKRNL----VSWNIIIKALARCGRIEDAQW-HLIRCRKGMMPLRNVVSW 207
+ +AL+LF M + + ++ +I ALA A+W H + R M NV
Sbjct: 1 VNEALNLFCTMQSQGIKPDSFTFVSVITALADLSVTRQAKWIHGLAIRTNMDT--NVFVA 174
Query: 208 NAMITGYAQNRRLDEALELFERMPERDMASWNAMLTGFFQNGELNRAEKLFAELPQKDVI 267
A++ YA+ ++ A ELF+ M ER + +WNAM+
Sbjct: 175 TALVDMYAKCGAIETARELFDMMQERHVITWNAMI------------------------- 279
Query: 268 TWTSMMTGYAQHGLSEEALKMFTKMQANGGLKPNNGTFVTVLGACSGLASLTEG 321
GY HGL + AL +F MQ LKPN+ TF++V+ ACS + EG
Sbjct: 280 ------DGYGTHGLGKAALDLFDDMQNEASLKPNDITFLSVISACSHSGFVEEG 423
Score = 56.2 bits (134), Expect = 3e-08
Identities = 33/91 (36%), Positives = 50/91 (54%), Gaps = 5/91 (5%)
Frame = +1
Query: 105 SVVTWTTMVNGYVKLNQIEEAESLFYEMPERNELSWNIMIGGYGQNGQIEKALDLFRRMP 164
+V T +V+ Y K IE A LF M ER+ ++WN MI GYG +G + ALDLF M
Sbjct: 160 NVFVATALVDMYAKCGAIETARELFDMMQERHVITWNAMIDGYGTHGLGKAALDLFDDMQ 339
Query: 165 -----KRNLVSWNIIIKALARCGRIEDAQWH 190
K N +++ +I A + G +E+ ++
Sbjct: 340 NEASLKPNDITFLSVISACSHSGFVEEGLYY 432
Score = 52.4 bits (124), Expect = 5e-07
Identities = 30/91 (32%), Positives = 50/91 (53%), Gaps = 5/91 (5%)
Frame = +1
Query: 78 TTMIDGYIKCGAIKEARKLFDQPDAKKSVVTWTTMVNGYVKLNQIEEAESLFYEMPER-- 135
T ++D Y KCGAI+ AR+LFD ++ V+TW M++GY + A LF +M
Sbjct: 175 TALVDMYAKCGAIETARELFDMMQ-ERHVITWNAMIDGYGTHGLGKAALDLFDDMQNEAS 351
Query: 136 ---NELSWNIMIGGYGQNGQIEKALDLFRRM 163
N++++ +I +G +E+ L F+ M
Sbjct: 352 LKPNDITFLSVISACSHSGFVEEGLYYFKIM 444
Score = 40.8 bits (94), Expect = 0.001
Identities = 27/85 (31%), Positives = 41/85 (47%), Gaps = 4/85 (4%)
Frame = +1
Query: 50 ISRLCKEGRTDHARKLFDQMPQRDKHLWTTMIDGYIKCGAIKEARKLFD----QPDAKKS 105
+ K G + AR+LFD M +R W MIDGY G K A LFD + K +
Sbjct: 184 VDMYAKCGAIETARELFDMMQERHVITWNAMIDGYGTHGLGKAALDLFDDMQNEASLKPN 363
Query: 106 VVTWTTMVNGYVKLNQIEEAESLFY 130
+T+ ++++ +E E L+Y
Sbjct: 364 DITFLSVISACSHSGFVE--EGLYY 432
Score = 37.4 bits (85), Expect = 0.016
Identities = 27/105 (25%), Positives = 48/105 (45%), Gaps = 4/105 (3%)
Frame = +1
Query: 387 NEAINLFNKMQELGFQANDVTYVELLTACSHAGLVDEGIQYFDKLLKNRSIQVKEDH--- 443
NEA+NLF MQ G + + T+V ++TA L D + K + +I+ D
Sbjct: 4 NEALNLFCTMQSQGIKPDSFTFVSVITA-----LADLSVTRQAKWIHGLAIRTNMDTNVF 168
Query: 444 -YACLVDLCGRAGRLKEAFYIIEGLGVKLSLSVWGPLLAGCNVHG 487
LVD+ + G ++ A + + + + + W ++ G HG
Sbjct: 169 VATALVDMYAKCGAIETARELFDMMQER-HVITWNAMIDGYGTHG 300
>AW774626 similar to PIR|T47909|T47 hypothetical protein T20K12.70 -
Arabidopsis thaliana, partial (12%)
Length = 703
Score = 122 bits (305), Expect = 5e-28
Identities = 73/226 (32%), Positives = 120/226 (52%), Gaps = 11/226 (4%)
Frame = +3
Query: 251 LNRAEKLFA--ELPQKDVITWTSMMTGYAQHGLSEEALKMFTKMQANGGLKPNNGTFVTV 308
++ AE LF E +K+ + WT+M+TGYAQ+G +A++ F M A G ++ N TF T+
Sbjct: 33 VSEAEFLFKGLEFDRKNHVLWTAMVTGYAQNGDGYKAVEFFRYMHAQG-VECNQYTFPTI 209
Query: 309 LGACSGLASLTEGQQIHQLISKTGFQENTRVVSALINMYSKCGELHIARKIFDDGLLRQR 368
L ACS + + G+Q+H I K+GF N V SAL++MY+KCG+L A+ + + +
Sbjct: 210 LTACSSVLARCFGEQVHGFIVKSGFGSNVYVQSALVDMYAKCGDLKNAKNMLE--TMEDD 383
Query: 369 DLISWNGMIAAYAHHGYGNEAINLFNKMQELGFQANDVTYVELLTAC---------SHAG 419
D++SWN ++ + HG EA+ LF M + +D T+ +L C H
Sbjct: 384 DVVSWNSLMVGFVRHGLEEEALRLFKNMHGRNMKIDDYTFPSVLNCCVVGSINPKSVHGL 563
Query: 420 LVDEGIQYFDKLLKNRSIQVKEDHYACLVDLCGRAGRLKEAFYIIE 465
++ G + + KL+ N LVD+ + G + A+ + E
Sbjct: 564 IIKTGFENY-KLVSN-----------ALVDMYAKTGDMDCAYTVFE 665
Score = 107 bits (267), Expect = 1e-23
Identities = 82/272 (30%), Positives = 129/272 (47%), Gaps = 10/272 (3%)
Frame = +3
Query: 112 MVNGYVKLNQIEEAESLF--YEMPERNELSWNIMIGGYGQNGQIEKALDLFRRMPKR--- 166
+V+ Y K + EAE LF E +N + W M+ GY QNG KA++ FR M +
Sbjct: 3 LVDMYAKCKCVSEAEFLFKGLEFDRKNHVLWTAMVTGYAQNGDGYKAVEFFRYMHAQGVE 182
Query: 167 -NLVSWNIIIKA----LARCGRIEDAQWHLIRCRKGMMPLRNVVSWNAMITGYAQNRRLD 221
N ++ I+ A LARC E +++ G NV +A++ YA+ L
Sbjct: 183 CNQYTFPTILTACSSVLARCFG-EQVHGFIVKSGFG----SNVYVQSALVDMYAKCGDLK 347
Query: 222 EALELFERMPERDMASWNAMLTGFFQNGELNRAEKLFAELPQKDVITWTSMMTGYAQHGL 281
A + E M + D+ SWN+++ GF +HGL
Sbjct: 348 NAKNMLETMEDDDVVSWNSLMVGF-------------------------------VRHGL 434
Query: 282 SEEALKMFTKMQANGGLKPNNGTFVTVLGACSGLASLTEGQQIHQLISKTGFQENTRVVS 341
EEAL++F M +K ++ TF +VL C + + +H LI KTGF+ V +
Sbjct: 435 EEEALRLFKNMHGR-NMKIDDYTFPSVLNCC--VVGSINPKSVHGLIIKTGFENYKLVSN 605
Query: 342 ALINMYSKCGELHIARKIFDDGLLRQRDLISW 373
AL++MY+K G++ A +F+ L ++D+ISW
Sbjct: 606 ALVDMYAKTGDMDCAYTVFEKML--EKDVISW 695
Score = 61.6 bits (148), Expect = 8e-10
Identities = 38/139 (27%), Positives = 73/139 (52%), Gaps = 3/139 (2%)
Frame = +3
Query: 105 SVVTWTTMVNGYVKLNQIEEAESLFYEMPERNELSWNIMIGGYGQNGQIEKALDLFRRMP 164
+V + +V+ Y K ++ A+++ M + + +SWN ++ G+ ++G E+AL LF+ M
Sbjct: 291 NVYVQSALVDMYAKCGDLKNAKNMLETMEDDDVVSWNSLMVGFVRHGLEEEALRLFKNMH 470
Query: 165 KRNLVSWNIIIKALARC---GRIEDAQWHLIRCRKGMMPLRNVVSWNAMITGYAQNRRLD 221
RN+ + ++ C G I H + + G + V NA++ YA+ +D
Sbjct: 471 GRNMKIDDYTFPSVLNCCVVGSINPKSVHGLIIKTGFENYKLVS--NALVDMYAKTGDMD 644
Query: 222 EALELFERMPERDMASWNA 240
A +FE+M E+D+ SW +
Sbjct: 645 CAYTVFEKMLEKDVISWTS 701
Score = 59.7 bits (143), Expect = 3e-09
Identities = 42/148 (28%), Positives = 72/148 (48%), Gaps = 3/148 (2%)
Frame = +3
Query: 343 LINMYSKCGELHIARKIFDDGLLRQRDLISWNGMIAAYAHHGYGNEAINLFNKMQELGFQ 402
L++MY+KC + A +F +++ + W M+ YA +G G +A+ F M G +
Sbjct: 3 LVDMYAKCKCVSEAEFLFKGLEFDRKNHVLWTAMVTGYAQNGDGYKAVEFFRYMHAQGVE 182
Query: 403 ANDVTYVELLTACSHAGLVDEGIQYFDKLLKN---RSIQVKEDHYACLVDLCGRAGRLKE 459
N T+ +LTACS G Q ++K+ ++ V+ + LVD+ + G LK
Sbjct: 183 CNQYTFPTILTACSSVLARCFGEQVHGFIVKSGFGSNVYVQ----SALVDMYAKCGDLKN 350
Query: 460 AFYIIEGLGVKLSLSVWGPLLAGCNVHG 487
A ++E + +S W L+ G HG
Sbjct: 351 AKNMLETMEDDDVVS-WNSLMVGFVRHG 431
>BF642497 weakly similar to PIR|H96713|H96 hypothetical protein T6L1.11
[imported] - Arabidopsis thaliana, partial (20%)
Length = 461
Score = 117 bits (293), Expect = 1e-26
Identities = 54/151 (35%), Positives = 96/151 (62%)
Frame = +3
Query: 267 ITWTSMMTGYAQHGLSEEALKMFTKMQANGGLKPNNGTFVTVLGACSGLASLTEGQQIHQ 326
I+WTSM+TG+ Q+GL +A+ +F +M+ L+ + TF +VL AC G+ +L EG+Q+H
Sbjct: 6 ISWTSMITGFTQNGLDRDAIDIFREMKLEN-LQMDQYTFGSVLTACGGVMALQEGKQVHA 182
Query: 327 LISKTGFQENTRVVSALINMYSKCGELHIARKIFDDGLLRQRDLISWNGMIAAYAHHGYG 386
I +T +++N V SAL++MY KC + A +F + ++++SW M+ Y +GY
Sbjct: 183 YIIRTDYKDNIFVASALVDMYCKCKNIKSAEAVFKK--MTCKNVVSWTAMLVGYGQNGYS 356
Query: 387 NEAINLFNKMQELGFQANDVTYVELLTACSH 417
EA+ F+ MQ+ G + +D T ++++C++
Sbjct: 357 EEAVKTFSDMQKYGIEPDDFTLGSVISSCAN 449
Score = 77.0 bits (188), Expect = 2e-14
Identities = 56/182 (30%), Positives = 90/182 (48%), Gaps = 1/182 (0%)
Frame = +3
Query: 138 LSWNIMIGGYGQNGQIEKALDLFRRMPKRNL-VSWNIIIKALARCGRIEDAQWHLIRCRK 196
+SW MI G+ QNG A+D+FR M NL + L CG
Sbjct: 6 ISWTSMITGFTQNGLDRDAIDIFREMKLENLQMDQYTFGSVLTACG-------------- 143
Query: 197 GMMPLRNVVSWNAMITGYAQNRRLDEALELFERMPERDMASWNAMLTGFFQNGELNRAEK 256
G+M L+ +A I R D +F +A++ + + + AE
Sbjct: 144 GVMALQEGKQVHAYII------RTDYKDNIFVA---------SALVDMYCKCKNIKSAEA 278
Query: 257 LFAELPQKDVITWTSMMTGYAQHGLSEEALKMFTKMQANGGLKPNNGTFVTVLGACSGLA 316
+F ++ K+V++WT+M+ GY Q+G SEEA+K F+ MQ G++P++ T +V+ +C+ LA
Sbjct: 279 VFKKMTCKNVVSWTAMLVGYGQNGYSEEAVKTFSDMQ-KYGIEPDDFTLGSVISSCANLA 455
Query: 317 SL 318
SL
Sbjct: 456 SL 461
Score = 53.5 bits (127), Expect = 2e-07
Identities = 36/128 (28%), Positives = 63/128 (49%), Gaps = 10/128 (7%)
Frame = +3
Query: 48 SDISRLCKEGRTDHARKLFDQMP----QRDKHLWTTMIDGYIKCGAI------KEARKLF 97
S I+ + G A +F +M Q D++ + +++ CG + K+
Sbjct: 18 SMITGFTQNGLDRDAIDIFREMKLENLQMDQYTFGSVLTA---CGGVMALQEGKQVHAYI 188
Query: 98 DQPDAKKSVVTWTTMVNGYVKLNQIEEAESLFYEMPERNELSWNIMIGGYGQNGQIEKAL 157
+ D K ++ + +V+ Y K I+ AE++F +M +N +SW M+ GYGQNG E+A+
Sbjct: 189 IRTDYKDNIFVASALVDMYCKCKNIKSAEAVFKKMTCKNVVSWTAMLVGYGQNGYSEEAV 368
Query: 158 DLFRRMPK 165
F M K
Sbjct: 369 KTFSDMQK 392
>TC80535 weakly similar to GP|14335136|gb|AAK59848.1 At2g15690/F9O13.24
{Arabidopsis thaliana}, partial (55%)
Length = 1727
Score = 117 bits (292), Expect = 2e-26
Identities = 66/235 (28%), Positives = 124/235 (52%)
Frame = +2
Query: 273 MTGYAQHGLSEEALKMFTKMQANGGLKPNNGTFVTVLGACSGLASLTEGQQIHQLISKTG 332
+T + Q G +EAL++ K G+K + F + C S+ + +++H ++
Sbjct: 209 LTRFCQEGKVKEALELMEK-----GIKADANCFEILFDLCGKSKSVEDAKKVHDYFLQST 373
Query: 333 FQENTRVVSALINMYSKCGELHIARKIFDDGLLRQRDLISWNGMIAAYAHHGYGNEAINL 392
F+ + ++ + +I MY C + AR++FD + R++ SW+ MI YA+ G+E + L
Sbjct: 374 FRSDFKMHNKVIEMYGNCKSMTDARRVFDH--MPNRNMDSWHMMIRGYANSTMGDEGLQL 547
Query: 393 FNKMQELGFQANDVTYVELLTACSHAGLVDEGIQYFDKLLKNRSIQVKEDHYACLVDLCG 452
F +M ELG + T + +L+AC A V++ Y + + I+ +HY L+D+ G
Sbjct: 548 FEQMNELGLEITSETMLAVLSACGSAEAVEDAYIYLESMKSKYGIEPGVEHYMGLLDVLG 727
Query: 453 RAGRLKEAFYIIEGLGVKLSLSVWGPLLAGCNVHGNADIGKLVAKKILKVEHENA 507
++G LKEA IE L + +++V+ L +HG+ D+ V + I+ ++ A
Sbjct: 728 QSGYLKEAEEFIEQLPFEPTVTVFETLKNYARIHGDVDLEDHVEELIVSLDPSKA 892
Score = 70.1 bits (170), Expect = 2e-12
Identities = 53/214 (24%), Positives = 103/214 (47%), Gaps = 20/214 (9%)
Frame = +2
Query: 38 NPNSEMKRCNS-----------DISRLCKEGRTDHARKLFDQMPQRDKHLWTTMIDGYIK 86
NPN++ + N D++R C+EG+ A +L ++ + D + + + D K
Sbjct: 140 NPNNQFQTPNVQEQAPPPPSIVDLTRFCQEGKVKEALELMEKGIKADANCFEILFDLCGK 319
Query: 87 CGAIKEARKLFD---QPDAKKSVVTWTTMVNGYVKLNQIEEAESLFYEMPERNELSWNIM 143
++++A+K+ D Q + ++ Y + +A +F MP RN SW++M
Sbjct: 320 SKSVEDAKKVHDYFLQSTFRSDFKMHNKVIEMYGNCKSMTDARRVFDHMPNRNMDSWHMM 499
Query: 144 IGGYGQNGQIEKALDLFRRMPKRNL-VSWNIIIKALARCG---RIEDAQWHL--IRCRKG 197
I GY + ++ L LF +M + L ++ ++ L+ CG +EDA +L ++ + G
Sbjct: 500 IRGYANSTMGDEGLQLFEQMNELGLEITSETMLAVLSACGSAEAVEDAYIYLESMKSKYG 679
Query: 198 MMPLRNVVSWNAMITGYAQNRRLDEALELFERMP 231
+ P V + ++ Q+ L EA E E++P
Sbjct: 680 IEP--GVEHYMGLLDVLGQSGYLKEAEEFIEQLP 775
Score = 41.6 bits (96), Expect = 9e-04
Identities = 25/61 (40%), Positives = 33/61 (53%)
Frame = +2
Query: 172 NIIIKALARCGRIEDAQWHLIRCRKGMMPLRNVVSWNAMITGYAQNRRLDEALELFERMP 231
N +I+ C + DA R MP RN+ SW+ MI GYA + DE L+LFE+M
Sbjct: 398 NKVIEMYGNCKSMTDA-----RRVFDHMPNRNMDSWHMMIRGYANSTMGDEGLQLFEQMN 562
Query: 232 E 232
E
Sbjct: 563 E 565
>BF635667 weakly similar to PIR|C85359|C853 hypothetical protein AT4g30700
[imported] - Arabidopsis thaliana, partial (6%)
Length = 603
Score = 114 bits (285), Expect = 1e-25
Identities = 56/152 (36%), Positives = 86/152 (55%), Gaps = 1/152 (0%)
Frame = +3
Query: 419 GLVDEGIQYFDKLLKNRSIQVKEDHYACLVDLCGRAGRLKEAFYIIEGL-GVKLSLSVWG 477
GL+D+G + F ++ + ++ +HYAC+VDL R GRL+EA ++E + ++ S+WG
Sbjct: 30 GLLDQGKKIFGSMISDYGLEPTAEHYACMVDLFSRCGRLEEALQLLERMKSSSVTGSMWG 209
Query: 478 PLLAGCNVHGNADIGKLVAKKILKVEHENAGTYSLLSNMYASVGKWKEAANVRMKMKDKG 537
LLAGC +H N IG++ A + ++E N Y L +Y S G + + KM+D G
Sbjct: 210 ALLAGCVMHQNVKIGEVAAHHLFQLEPNNTSNYVALWGIYQSRGMVLGVSTIXGKMRDLG 389
Query: 538 LKKQPGCSWIEVGNTVQVFVVGDKSHSQSEML 569
L K PGC WI + F GD SH S ++
Sbjct: 390 LVKTPGCXWINIAGRAHKFYQGDLSHPLSHII 485
Score = 31.2 bits (69), Expect = 1.2
Identities = 27/107 (25%), Positives = 50/107 (46%), Gaps = 6/107 (5%)
Frame = +3
Query: 210 MITGYAQNRRLDEALELFERMPERDM--ASWNAMLTG--FFQNGELNR--AEKLFAELPQ 263
M+ +++ RL+EAL+L ERM + + W A+L G QN ++ A LF +L
Sbjct: 114 MVDLFSRCGRLEEALQLLERMKSSSVTGSMWGALLAGCVMHQNVKIGEVAAHHLF-QLEP 290
Query: 264 KDVITWTSMMTGYAQHGLSEEALKMFTKMQANGGLKPNNGTFVTVLG 310
+ + ++ Y G+ + KM+ G +K ++ + G
Sbjct: 291 NNTSNYVALWGIYQSRGMVLGVSTIXGKMRDLGLVKTPGCXWINIAG 431
Score = 30.0 bits (66), Expect = 2.6
Identities = 28/124 (22%), Positives = 52/124 (41%), Gaps = 11/124 (8%)
Frame = +3
Query: 57 GRTDHARKLFDQMP-----QRDKHLWTTMIDGYIKCGAIKEARKLFDQPDAKKSVVT--- 108
G D +K+F M + + M+D + +CG ++EA +L ++ K S VT
Sbjct: 30 GLLDQGKKIFGSMISDYGLEPTAEHYACMVDLFSRCGRLEEALQLLER--MKSSSVTGSM 203
Query: 109 WTTMVNGYVKLNQI---EEAESLFYEMPERNELSWNIMIGGYGQNGQIEKALDLFRRMPK 165
W ++ G V + E A +++ N ++ + G Y G + + +M
Sbjct: 204 WGALLAGCVMHQNVKIGEVAAHHLFQLEPNNTSNYVALWGIYQSRGMVLGVSTIXGKMRD 383
Query: 166 RNLV 169
LV
Sbjct: 384 LGLV 395
>BG645370 weakly similar to PIR|T00405|T004 hypothetical protein At2g44880
[imported] - Arabidopsis thaliana, partial (11%)
Length = 784
Score = 109 bits (273), Expect = 3e-24
Identities = 51/143 (35%), Positives = 90/143 (62%)
Frame = -2
Query: 408 YVELLTACSHAGLVDEGIQYFDKLLKNRSIQVKEDHYACLVDLCGRAGRLKEAFYIIEGL 467
+ +LTAC+HAG +D+G +YF+K++ N + D YACL+DL R G L++A ++E +
Sbjct: 783 FTAVLTACNHAGFIDKGEEYFNKMITNYGLSPDIDIYACLIDLYARNGNLRKARDLMEEM 604
Query: 468 GVKLSLSVWGPLLAGCNVHGNADIGKLVAKKILKVEHENAGTYSLLSNMYASVGKWKEAA 527
+ +W L+ C ++G+ ++G+ A +++K+E NA Y L+++Y + G W EA+
Sbjct: 603 PYDPNCIIWSSFLSACKIYGDVELGREAAIQLIKMEPCNAAPYLTLAHIYTTKGLWNEAS 424
Query: 528 NVRMKMKDKGLKKQPGCSWIEVG 550
VR M+ + +K PG S +E+G
Sbjct: 423 EVRSLMQQRVKRKPPGWS-MELG 358
Score = 29.3 bits (64), Expect = 4.5
Identities = 19/61 (31%), Positives = 33/61 (53%), Gaps = 3/61 (4%)
Frame = -3
Query: 535 DKGLKK--QPGC-SWIEVGNTVQVFVVGDKSHSQSEMLEYLLLGLHTKMKKFGDILDDDL 591
+KGLK+ Q G SW+EV VF V D +H QS + ++ ++ +F + +D+
Sbjct: 404 NKGLKENLQGGAWSWVEVDKHNHVFAVDDVTHQQSNEIYAE*ESIYFEILEFSPYVVEDI 225
Query: 592 S 592
+
Sbjct: 224 N 222
Score = 29.3 bits (64), Expect = 4.5
Identities = 23/96 (23%), Positives = 42/96 (42%), Gaps = 9/96 (9%)
Frame = -2
Query: 207 WNAMITGYAQNRRLDEALELFERMPER-----DMASWNAMLTGFFQNGELNRAEKLFAEL 261
+ A++T +D+ E F +M D+ + ++ + +NG L +A L E+
Sbjct: 783 FTAVLTACNHAGFIDKGEEYFNKMITNYGLSPDIDIYACLIDLYARNGNLRKARDLMEEM 604
Query: 262 P-QKDVITWTSMMTG---YAQHGLSEEALKMFTKMQ 293
P + I W+S ++ Y L EA KM+
Sbjct: 603 PYDPNCIIWSSFLSACKIYGDVELGREAAIQLIKME 496
>BG646586 weakly similar to GP|10177683|dbj selenium-binding protein-like
{Arabidopsis thaliana}, partial (9%)
Length = 687
Score = 102 bits (253), Expect = 5e-22
Identities = 45/112 (40%), Positives = 72/112 (64%)
Frame = -3
Query: 437 IQVKEDHYACLVDLCGRAGRLKEAFYIIEGLGVKLSLSVWGPLLAGCNVHGNADIGKLVA 496
I+ +HY +VDL GRAG+L++A+ +I+ + WG LLA C V+GN ++G++ A
Sbjct: 664 IEPLPEHYTGMVDLLGRAGQLEKAYSLIKENSTSADATTWGSLLAACRVYGNVELGEIAA 485
Query: 497 KKILKVEHENAGTYSLLSNMYASVGKWKEAANVRMKMKDKGLKKQPGCSWIE 548
+ + +++ ++G Y LL+N YAS KW+ A V+ M KG+KK G SWI+
Sbjct: 484 RHLFEIDPTDSGNYVLLANTYASNDKWERAEEVKKLMSKKGMKKPSGYSWIQ 329
Score = 30.8 bits (68), Expect = 1.5
Identities = 21/97 (21%), Positives = 46/97 (46%), Gaps = 5/97 (5%)
Frame = -3
Query: 77 WTTMIDGYIKCGAIKEARKLFDQPDAKKSVVTWTTM-----VNGYVKLNQIEEAESLFYE 131
+T M+D + G +++A L + TW ++ V G V+L +I A +E
Sbjct: 643 YTGMVDLLGRAGQLEKAYSLIKENSTSADATTWGSLLAACRVYGNVELGEI--AARHLFE 470
Query: 132 MPERNELSWNIMIGGYGQNGQIEKALDLFRRMPKRNL 168
+ + ++ ++ Y N + E+A ++ + M K+ +
Sbjct: 469 IDPTDSGNYVLLANTYASNDKWERAEEVKKLMSKKGM 359
>AW257410 weakly similar to GP|10177533|db selenium-binding protein-like
{Arabidopsis thaliana}, partial (21%)
Length = 724
Score = 95.9 bits (237), Expect = 4e-20
Identities = 59/217 (27%), Positives = 115/217 (52%), Gaps = 10/217 (4%)
Frame = +1
Query: 86 KCGAIKEARKLFDQPDAKKSV---VTWTTMVNGYVKL--NQIEEAESLFYEMPERNELSW 140
+C I E ++++ Q K ++ +T + ++ Y + + + A +F + N + W
Sbjct: 4 RCSNIGELKQIYVQFLKKGTIRHKLTVSRLLTTYASMEFSNLTYARMVFDRISSPNTVMW 183
Query: 141 NIMIGGYGQNGQIEKALDLFRRMPKR----NLVSWNIIIKALARCGRI-EDAQWHLIRCR 195
N MI Y + E+AL L+ +M N ++ ++KA + + E Q H+ +
Sbjct: 184 NTMIRAYSNSNDPEEALLLYHQMLHHSIPHNAYTFPFLLKACSALSALAETHQIHVQIIK 363
Query: 196 KGMMPLRNVVSWNAMITGYAQNRRLDEALELFERMPERDMASWNAMLTGFFQNGELNRAE 255
+G V + N+++ YA + + A LF+ +P RD+ SWN M+ G+ + G + A
Sbjct: 364 RGFGS--EVYATNSLLRVYAISGSIKSAHVLFDLLPSRDIVSWNTMIDGYIKCGNVEMAY 537
Query: 256 KLFAELPQKDVITWTSMMTGYAQHGLSEEALKMFTKM 292
K+F +P+K+VI+WTSM+ G+ + G+ ++AL + +M
Sbjct: 538 KIFQAMPEKNVISWTSMIVGFVRTGMHKKALSLLQQM 648
Score = 84.0 bits (206), Expect = 2e-16
Identities = 53/185 (28%), Positives = 88/185 (46%)
Frame = +1
Query: 251 LNRAEKLFAELPQKDVITWTSMMTGYAQHGLSEEALKMFTKMQANGGLKPNNGTFVTVLG 310
L A +F + + + W +M+ Y+ EEAL ++ +M + + N TF +L
Sbjct: 127 LTYARMVFDRISSPNTVMWNTMIRAYSNSNDPEEALLLYHQM-LHHSIPHNAYTFPFLLK 303
Query: 311 ACSGLASLTEGQQIHQLISKTGFQENTRVVSALINMYSKCGELHIARKIFDDGLLRQRDL 370
ACS L++L E QIH I K GF ++L+ +Y+ G + A +FD LL RD+
Sbjct: 304 ACSALSALAETHQIHVQIIKRGFGSEVYATNSLLRVYAISGSIKSAHVLFD--LLPSRDI 477
Query: 371 ISWNGMIAAYAHHGYGNEAINLFNKMQELGFQANDVTYVELLTACSHAGLVDEGIQYFDK 430
+SWN MI Y G A +F M E N +++ ++ G+ + + +
Sbjct: 478 VSWNTMIDGYIKCGNVEMAYKIFQAMPE----KNVISWTSMIVGFVRTGMHKKALSLLQQ 645
Query: 431 LLKNR 435
+L R
Sbjct: 646 MLVAR 660
Score = 74.3 bits (181), Expect = 1e-13
Identities = 48/177 (27%), Positives = 88/177 (49%), Gaps = 7/177 (3%)
Frame = +1
Query: 61 HARKLFDQMPQRDKHLWTTMIDGYIKCGAIKEARKLFDQ---PDAKKSVVTWTTMVNGYV 117
+AR +FD++ + +W TMI Y +EA L+ Q + T+ ++
Sbjct: 133 YARMVFDRISSPNTVMWNTMIRAYSNSNDPEEALLLYHQMLHHSIPHNAYTFPFLLKACS 312
Query: 118 KLNQIEEAESLFYEMPER----NELSWNIMIGGYGQNGQIEKALDLFRRMPKRNLVSWNI 173
L+ + E + ++ +R + N ++ Y +G I+ A LF +P R++VSWN
Sbjct: 313 ALSALAETHQIHVQIIKRGFGSEVYATNSLLRVYAISGSIKSAHVLFDLLPSRDIVSWNT 492
Query: 174 IIKALARCGRIEDAQWHLIRCRKGMMPLRNVVSWNAMITGYAQNRRLDEALELFERM 230
+I +CG +E A + + + MP +NV+SW +MI G+ + +AL L ++M
Sbjct: 493 MIDGYIKCGNVEMA-YKIFQA----MPEKNVISWTSMIVGFVRTGMHKKALSLLQQM 648
Score = 70.9 bits (172), Expect = 1e-12
Identities = 38/92 (41%), Positives = 55/92 (59%)
Frame = +1
Query: 41 SEMKRCNSDISRLCKEGRTDHARKLFDQMPQRDKHLWTTMIDGYIKCGAIKEARKLFDQP 100
SE+ NS + G A LFD +P RD W TMIDGYIKCG ++ A K+F Q
Sbjct: 376 SEVYATNSLLRVYAISGSIKSAHVLFDLLPSRDIVSWNTMIDGYIKCGNVEMAYKIF-QA 552
Query: 101 DAKKSVVTWTTMVNGYVKLNQIEEAESLFYEM 132
+K+V++WT+M+ G+V+ ++A SL +M
Sbjct: 553 MPEKNVISWTSMIVGFVRTGMHKKALSLLQQM 648
Score = 41.2 bits (95), Expect = 0.001
Identities = 20/61 (32%), Positives = 34/61 (54%)
Frame = +1
Query: 39 PNSEMKRCNSDISRLCKEGRTDHARKLFDQMPQRDKHLWTTMIDGYIKCGAIKEARKLFD 98
P+ ++ N+ I K G + A K+F MP+++ WT+MI G+++ G K+A L
Sbjct: 463 PSRDIVSWNTMIDGYIKCGNVEMAYKIFQAMPEKNVISWTSMIVGFVRTGMHKKALSLLQ 642
Query: 99 Q 99
Q
Sbjct: 643 Q 645
>BE202961 weakly similar to GP|6016735|gb|A hypothetical protein {Arabidopsis
thaliana}, partial (14%)
Length = 613
Score = 95.5 bits (236), Expect = 5e-20
Identities = 46/138 (33%), Positives = 80/138 (57%)
Frame = +2
Query: 253 RAEKLFAELPQKDVITWTSMMTGYAQHGLSEEALKMFTKMQANGGLKPNNGTFVTVLGAC 312
+A +F +P+KDVI W + +GYA +G+ E++ +F M ++G +P+ V +L
Sbjct: 206 KAVDIFNRMPKKDVIAWAVLFSGYADNGMVHESMWVFRNMLSSG-TRPDAIALVKILTTV 382
Query: 313 SGLASLTEGQQIHQLISKTGFQENTRVVSALINMYSKCGELHIARKIFDDGLLRQRDLIS 372
S L L + H + K GF+ N + ++LI +Y+KC + A K+F + +D+++
Sbjct: 383 SELGILQQAVCFHAFVIKNGFENNQFIGASLIEVYAKCSSIEDANKVFKG--MTYKDVVT 556
Query: 373 WNGMIAAYAHHGYGNEAI 390
W+ +IAAY HG G EA+
Sbjct: 557 WSSIIAAYGFHGQGEEAL 610
Score = 94.7 bits (234), Expect = 9e-20
Identities = 54/193 (27%), Positives = 107/193 (54%)
Frame = +2
Query: 298 LKPNNGTFVTVLGACSGLASLTEGQQIHQLISKTGFQENTRVVSALINMYSKCGELHIAR 357
+KPN T V+VL AC+ +++L EG +IH+L GF+ T V +AL++MY KC A
Sbjct: 35 IKPNWVTVVSVLRACACISNLEEGMKIHELAVNYGFEMETTVSTALMDMYMKCFSPEKAV 214
Query: 358 KIFDDGLLRQRDLISWNGMIAAYAHHGYGNEAINLFNKMQELGFQANDVTYVELLTACSH 417
IF+ + ++D+I+W + + YA +G +E++ +F M G + + + V++LT S
Sbjct: 215 DIFN--RMPKKDVIAWAVLFSGYADNGMVHESMWVFRNMLSSGTRPDAIALVKILTTVSE 388
Query: 418 AGLVDEGIQYFDKLLKNRSIQVKEDHYACLVDLCGRAGRLKEAFYIIEGLGVKLSLSVWG 477
G++ + + + ++KN + + A L+++ + +++A + +G+ K + W
Sbjct: 389 LGILQQAVCFHAFVIKN-GFENNQFIGASLIEVYAKCSSIEDANKVFKGMTYK-DVVTWS 562
Query: 478 PLLAGCNVHGNAD 490
++A HG +
Sbjct: 563 SIIAAYGFHGQGE 601
Score = 62.4 bits (150), Expect = 5e-10
Identities = 35/138 (25%), Positives = 72/138 (51%), Gaps = 4/138 (2%)
Frame = +2
Query: 154 EKALDLFRRMPKRNLVSWNIIIKALARCGRIEDAQWHLIRCRKGMMPLRNVVSWNAMITG 213
EKA+D+F RMPK+++++W ++ A G + ++ W + R + ++ ++T
Sbjct: 203 EKAVDIFNRMPKKDVIAWAVLFSGYADNGMVHESMW-VFRNMLSSGTRPDAIALVKILTT 379
Query: 214 YAQNRRLDEAL----ELFERMPERDMASWNAMLTGFFQNGELNRAEKLFAELPQKDVITW 269
++ L +A+ + + E + +++ + + + A K+F + KDV+TW
Sbjct: 380 VSELGILQQAVCFHAFVIKNGFENNQFIGASLIEVYAKCSSIEDANKVFKGMTYKDVVTW 559
Query: 270 TSMMTGYAQHGLSEEALK 287
+S++ Y HG EEALK
Sbjct: 560 SSIIAAYGFHGQGEEALK 613
Score = 60.8 bits (146), Expect = 1e-09
Identities = 41/201 (20%), Positives = 98/201 (48%), Gaps = 11/201 (5%)
Frame = +2
Query: 67 DQMPQRDKHLWTTMIDGYIKCGAI---KEARKLFDQP---DAKKSVVTWTTMVNGYVKLN 120
+ + +R K W T++ C I +E K+ + + T +++ Y+K
Sbjct: 17 EMLDKRIKPNWVTVVSVLRACACISNLEEGMKIHELAVNYGFEMETTVSTALMDMYMKCF 196
Query: 121 QIEEAESLFYEMPERNELSWNIMIGGYGQNGQIEKALDLFRRM----PKRNLVSWNIIIK 176
E+A +F MP+++ ++W ++ GY NG + +++ +FR M + + ++ I+
Sbjct: 197 SPEKAVDIFNRMPKKDVIAWAVLFSGYADNGMVHESMWVFRNMLSSGTRPDAIALVKILT 376
Query: 177 ALARCGRIEDAQ-WHLIRCRKGMMPLRNVVSWNAMITGYAQNRRLDEALELFERMPERDM 235
++ G ++ A +H + G N ++I YA+ +++A ++F+ M +D+
Sbjct: 377 TVSELGILQQAVCFHAFVIKNGFE--NNQFIGASLIEVYAKCSSIEDANKVFKGMTYKDV 550
Query: 236 ASWNAMLTGFFQNGELNRAEK 256
+W++++ + +G+ A K
Sbjct: 551 VTWSSIIAAYGFHGQGEEALK 613
Score = 45.8 bits (107), Expect = 5e-05
Identities = 33/136 (24%), Positives = 57/136 (41%), Gaps = 38/136 (27%)
Frame = +2
Query: 60 DHARKLFDQMPQRDKHLWTTMIDGYIKCGAIKEARKLF-------DQPDAKKSVVTWTT- 111
+ A +F++MP++D W + GY G + E+ +F +PDA V TT
Sbjct: 203 EKAVDIFNRMPKKDVIAWAVLFSGYADNGMVHESMWVFRNMLSSGTRPDAIALVKILTTV 382
Query: 112 ------------------------------MVNGYVKLNQIEEAESLFYEMPERNELSWN 141
++ Y K + IE+A +F M ++ ++W+
Sbjct: 383 SELGILQQAVCFHAFVIKNGFENNQFIGASLIEVYAKCSSIEDANKVFKGMTYKDVVTWS 562
Query: 142 IMIGGYGQNGQIEKAL 157
+I YG +GQ E+AL
Sbjct: 563 SIIAAYGFHGQGEEAL 610
>BQ122819 similar to PIR|C84748|C84 hypothetical protein At2g33680 [imported]
- Arabidopsis thaliana, partial (15%)
Length = 771
Score = 94.4 bits (233), Expect = 1e-19
Identities = 51/128 (39%), Positives = 73/128 (56%)
Frame = +2
Query: 453 RAGRLKEAFYIIEGLGVKLSLSVWGPLLAGCNVHGNADIGKLVAKKILKVEHENAGTYSL 512
RAG+L EA IE V L +W LL C H N ++G +K++++ + Y L
Sbjct: 26 RAGKLNEAKEFIESATVDHGLCLWRILLGACKNHRNYELGVYAGEKLVELGSPESSAYVL 205
Query: 513 LSNMYASVGKWKEAANVRMKMKDKGLKKQPGCSWIEVGNTVQVFVVGDKSHSQSEMLEYL 572
LS++Y ++G + VR MK +G+ K+PGCSWIE+ V VFVVGD H Q + + L
Sbjct: 206 LSSIYTALGDRENVERVRRIMKARGVNKEPGCSWIELKGLVHVFVVGDNQHPQVDEIR-L 382
Query: 573 LLGLHTKM 580
L L TK+
Sbjct: 383 ELELLTKL 406
>TC83657 weakly similar to GP|11994382|dbj|BAB02341.
gb|AAB82628.1~gene_id:MIL23.2~similar to unknown protein
{Arabidopsis thaliana}, partial (10%)
Length = 1299
Score = 94.0 bits (232), Expect = 1e-19
Identities = 51/143 (35%), Positives = 87/143 (60%), Gaps = 1/143 (0%)
Frame = +3
Query: 411 LLTACSHAGLVDEGIQYFDKLLKNRSIQVKEDHYACLVDLCGRAGRLKEAFYIIEGLGVK 470
+L+AC+H GL+ E ++ K+ + I++ HY C+VDL GRAG+LKEA+ +I+ + +K
Sbjct: 84 VLSACAHGGLMSEALEVISKM-EEYGIEMGIRHYGCMVDLLGRAGKLKEAYELIKRMPMK 260
Query: 471 LSLSVWGPLLAGCNVHGNADIGKLVAKKILKVEHENAGTYS-LLSNMYASVGKWKEAANV 529
+ +V G ++ C +H + + + V K I +++ LLSN+YA+ KW++A +
Sbjct: 261 PNETVLGAMIGACWIHSDMKMAEQVMKMIGADSAACVNSHNVLLSNIYAASEKWEKAEMI 440
Query: 530 RMKMKDKGLKKQPGCSWIEVGNT 552
R M D G +K PG S I + N+
Sbjct: 441 RSSMVDGGSEKIPGYSSIILSNS 509
>BG455368 weakly similar to GP|9759524|dbj selenium-binding protein-like
{Arabidopsis thaliana}, partial (33%)
Length = 687
Score = 93.2 bits (230), Expect = 3e-19
Identities = 42/129 (32%), Positives = 75/129 (57%), Gaps = 1/129 (0%)
Frame = +1
Query: 457 LKEAFYIIEGLGVKLSLSVWGPLLAGCNVHGNADIGKLVAKKILKVEHENAGTYSLLSNM 516
+++A +++ + ++ + VWG LL ++HGN D+ ++ ++ + ++E +N G Y LLS
Sbjct: 7 VEKALQLVQTMPMEPNGGVWGALLGASHIHGNPDVAEIASRSLFELEPDNLGNYLLLSKT 186
Query: 517 YASVGKWKEAANVRMKMKDKGLKKQPGCSWIEVGN-TVQVFVVGDKSHSQSEMLEYLLLG 575
YA KW + + VR M++K L+K PGCSW+E N + F GD H + ++ L
Sbjct: 187 YALAAKWDDVSRVRKLMREKQLRKNPGCSWVEAKNGIIHEFFAGDVKHPEINEIKKALDD 366
Query: 576 LHTKMKKFG 584
L ++K G
Sbjct: 367 LLQRLKCTG 393
>BG584228 weakly similar to PIR|F71411|F71 hypothetical protein - Arabidopsis
thaliana, partial (19%)
Length = 827
Score = 81.3 bits (199), Expect = 1e-15
Identities = 54/196 (27%), Positives = 95/196 (47%), Gaps = 2/196 (1%)
Frame = -3
Query: 377 IAAYAHHGYGNEAINLFNKMQ--ELGFQANDVTYVELLTACSHAGLVDEGIQYFDKLLKN 434
+ YAH G A+ F +M G + + VT V +L+AC AG+V G+Q F+ + N
Sbjct: 684 VRGYAHQGNVYMALRQFEEMTFGSRGIRLSYVTLVSILSACGRAGVVLRGMQIFESMRLN 505
Query: 435 RSIQVKEDHYACLVDLCGRAGRLKEAFYIIEGLGVKLSLSVWGPLLAGCNVHGNADIGKL 494
+I+ HYAC+V L G I+ + ++ ++S WG LL +HG +GK+
Sbjct: 504 YAIEPGTGHYACVVGLLG------SVC*FIQNMPIQPTISEWGVLLGAFRMHGKTKMGKI 343
Query: 495 VAKKILKVEHENAGTYSLLSNMYASVGKWKEAANVRMKMKDKGLKKQPGCSWIEVGNTVQ 554
A+K+ +++ E G + + W+ + + +++ + C WI V N +
Sbjct: 342 TAEKLFELDDEYFGNHVV**ACIC----WQMGRSHC*QEENERQR**EIC-WIAVKNRIH 178
Query: 555 VFVVGDKSHSQSEMLE 570
VF D SH + ++
Sbjct: 177 VFQANDSSHEMNSEIQ 130
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.320 0.136 0.409
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 18,680,339
Number of Sequences: 36976
Number of extensions: 261355
Number of successful extensions: 1605
Number of sequences better than 10.0: 134
Number of HSP's better than 10.0 without gapping: 1230
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1466
length of query: 597
length of database: 9,014,727
effective HSP length: 102
effective length of query: 495
effective length of database: 5,243,175
effective search space: 2595371625
effective search space used: 2595371625
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 61 (28.1 bits)
Lotus: description of TM0026.14


