

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0024.11
(148 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
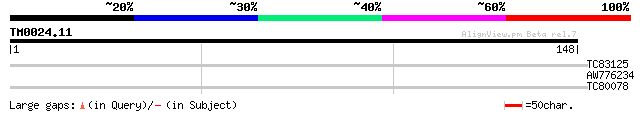
Score E
Sequences producing significant alignments: (bits) Value
TC83125 similar to PIR|T05841|T05841 spliceosome-associated prot... 28 1.0
AW776234 similar to GP|21555241|gb| putative SET protein phospa... 27 3.0
TC80078 similar to PIR|T02105|T02105 calcium-dependent protein k... 25 8.8
>TC83125 similar to PIR|T05841|T05841 spliceosome-associated protein homolog
F17L22.120 - Arabidopsis thaliana, partial (24%)
Length = 722
Score = 28.5 bits (62), Expect = 1.0
Identities = 21/79 (26%), Positives = 37/79 (46%), Gaps = 1/79 (1%)
Frame = +2
Query: 67 KEISVARLPES-EYYRAYSPEFRNETLVEGKIKRIFKFSKDSKSHLKRSSKSEIAYSGHA 125
+++ + +PE + Y EFR KI F F++ + S + + K ++A A
Sbjct: 224 EQVEIEYVPEKVDLYEGMDEEFR-------KIFEKFSFTEVAAS--EETDKKDVAEETAA 376
Query: 126 RRKAANSCHAYTDSTRESE 144
+K ANS Y D ++E
Sbjct: 377 TKKKANSDSDYEDEENDNE 433
>AW776234 similar to GP|21555241|gb| putative SET protein phospatase 2A
inhibitor {Arabidopsis thaliana}, partial (32%)
Length = 378
Score = 26.9 bits (58), Expect = 3.0
Identities = 22/86 (25%), Positives = 44/86 (50%), Gaps = 7/86 (8%)
Frame = +2
Query: 42 FQNSKQISKIWEKRYLKSQKRWFRSKEIS--VARLPESEYYRAYSPEFRN-----ETLVE 94
F +S + I+ +R ++++RW ++ S ++P+ + SP RN T
Sbjct: 35 FSSSSTSAAIFTQRRSEAKRRWLETEPRSRNSPKIPKKLTLNSSSPS-RNCKRSKMTSKR 211
Query: 95 GKIKRIFKFSKDSKSHLKRSSKSEIA 120
+KR+ +FSK S+S ++ S+ I+
Sbjct: 212 LMMKRVIRFSKLSRSIMRLGSRFTIS 289
>TC80078 similar to PIR|T02105|T02105 calcium-dependent protein kinase (EC
2.7.1.-) T3K9.9 - Arabidopsis thaliana, partial (47%)
Length = 1326
Score = 25.4 bits (54), Expect = 8.8
Identities = 15/34 (44%), Positives = 21/34 (61%)
Frame = -3
Query: 89 NETLVEGKIKRIFKFSKDSKSHLKRSSKSEIAYS 122
N+T VE + +F+F K SK HL R ++AYS
Sbjct: 1111 NKTRVE---ESVFRFVKHSKGHLTR----DVAYS 1031
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.316 0.128 0.360
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 4,088,861
Number of Sequences: 36976
Number of extensions: 46425
Number of successful extensions: 283
Number of sequences better than 10.0: 6
Number of HSP's better than 10.0 without gapping: 283
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 283
length of query: 148
length of database: 9,014,727
effective HSP length: 87
effective length of query: 61
effective length of database: 5,797,815
effective search space: 353666715
effective search space used: 353666715
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 54 (25.4 bits)
Lotus: description of TM0024.11


