

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
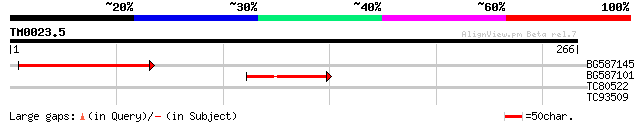
Query= TM0023.5
(266 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
BG587145 similar to PIR|H86337|H8 protein F5M15.26 [imported] - ... 53 1e-07
BG587101 similar to GP|6691191|gb F7F22.15 {Arabidopsis thaliana... 40 7e-04
TC80522 weakly similar to PIR|H96548|H96548 unknown protein [imp... 29 2.1
TC93509 weakly similar to GP|3769330|dbj|BAA33879.1 alpha-amylas... 27 8.2
>BG587145 similar to PIR|H86337|H8 protein F5M15.26 [imported] - Arabidopsis
thaliana, partial (13%)
Length = 763
Score = 53.1 bits (126), Expect = 1e-07
Identities = 25/64 (39%), Positives = 42/64 (65%)
Frame = +3
Query: 5 KPTPEVPLILYLAVTSTAVSAMLLQEEHKKYKMIYFVSHTLQGAEVRYQKIEKAALAILK 64
KP L LY+A++STAVS++L++E+ + K I++ S + E RY +EK A A++
Sbjct: 570 KPEAGDTLSLYIAISSTAVSSVLIREDRGEQKPIFYTSKRMTDPETRYPTLEKMAFAVIT 749
Query: 65 TARR 68
+AR+
Sbjct: 750 SARK 761
>BG587101 similar to GP|6691191|gb F7F22.15 {Arabidopsis thaliana}, partial
(10%)
Length = 624
Score = 40.4 bits (93), Expect = 7e-04
Identities = 19/40 (47%), Positives = 26/40 (64%)
Frame = +2
Query: 112 FAIWGADILGLFSVAKAQMKFIVVAVDYFTKWIEAEAVET 151
F +WG D +G F + K+I+VAVDY +KW+EA A T
Sbjct: 311 FDVWGIDFMGPFP-SSYNNKYILVAVDYVSKWVEAIASPT 427
>TC80522 weakly similar to PIR|H96548|H96548 unknown protein [imported] -
Arabidopsis thaliana, partial (12%)
Length = 807
Score = 28.9 bits (63), Expect = 2.1
Identities = 15/34 (44%), Positives = 22/34 (64%)
Frame = +2
Query: 186 EVENQSWRVITMNEEENNDNLIANLIMLPELQRE 219
E++ QS + TMN NN+N A + LP+LQR+
Sbjct: 482 EIDQQSQTMQTMNNNNNNNNNEA--VDLPQLQRQ 577
>TC93509 weakly similar to GP|3769330|dbj|BAA33879.1 alpha-amylase
{Phaseolus vulgaris}, partial (20%)
Length = 474
Score = 26.9 bits (58), Expect = 8.2
Identities = 25/84 (29%), Positives = 37/84 (43%), Gaps = 3/84 (3%)
Frame = -3
Query: 23 VSAMLLQEEHKKYKMIYFVSHTLQGAEVRYQKIE---KAALAILKTARRLRPYFQSFQIK 79
++A L++ ++Y++ L GA R Q +E K AL + AR +P
Sbjct: 442 LNACLMERFQQRYQL-------LLGAIGRVQVVEAPGKVALGRHRLARWRQPNVGEPSFL 284
Query: 80 VKTDILLRQVWQKPDLSGAPPEQL 103
D+LL V L APPE L
Sbjct: 283 DSRDVLLHYVVPATRLMAAPPEPL 212
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.320 0.132 0.382
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 7,290,524
Number of Sequences: 36976
Number of extensions: 82858
Number of successful extensions: 350
Number of sequences better than 10.0: 8
Number of HSP's better than 10.0 without gapping: 348
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 350
length of query: 266
length of database: 9,014,727
effective HSP length: 94
effective length of query: 172
effective length of database: 5,538,983
effective search space: 952705076
effective search space used: 952705076
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 57 (26.6 bits)
Lotus: description of TM0023.5


