

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0022.23
(1569 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
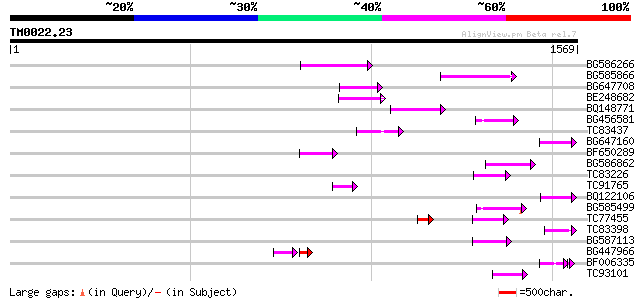
Score E
Sequences producing significant alignments: (bits) Value
BG586266 similar to GP|7267666|em RNA-directed DNA polymerase-li... 144 2e-34
BG585866 136 6e-32
BG647708 weakly similar to GP|13786450|gb| putative reverse tran... 77 5e-14
BE248682 similar to GP|18568269|gb putative gag-pol polyprotein ... 73 1e-12
BQ148771 69 1e-11
BG456581 62 2e-09
TC83437 weakly similar to PIR|D86384|D86384 unknown protein [imp... 60 7e-09
BG647160 homologue to GP|15004435|gb| maturase {Ionopsis satyrio... 60 9e-09
BF650289 weakly similar to GP|9049283|dbj| orf129a {Beta vulgari... 59 2e-08
BG586862 57 6e-08
TC83226 weakly similar to PIR|G86419|G86419 probable reverse tra... 57 7e-08
TC91765 weakly similar to GP|19881779|gb|AAM01180.1 Putative ret... 56 1e-07
BQ122106 similar to GP|9366656|emb|C probable similar to ring-h2... 55 3e-07
BG585499 52 1e-06
TC77455 similar to GP|22335695|dbj|BAC10549. nine-cis-epoxycarot... 51 3e-06
TC83398 similar to PIR|T05150|T05150 hypothetical protein F18E5.... 50 9e-06
BG587113 weakly similar to PIR|A84888|A8 hypothetical protein At... 47 4e-05
BG447966 weakly similar to PIR|G96509|G96 protein F27F5.21 [impo... 36 2e-04
BF006335 similar to GP|14334538|gb| unknown protein {Arabidopsis... 40 8e-04
TC93101 43 8e-04
>BG586266 similar to GP|7267666|em RNA-directed DNA polymerase-like protein
{Arabidopsis thaliana}, partial (18%)
Length = 789
Score = 144 bits (364), Expect = 2e-34
Identities = 80/201 (39%), Positives = 117/201 (57%), Gaps = 1/201 (0%)
Frame = -3
Query: 804 KVMVNRLRQFLQELIGPLQSSFIPGRGTIDNAIILQKIVHHMNSRRGKCQ-NVVFKLDLE 862
K++ R++ L+ +I P QS+F+PGR DN +I KI+H++ K ++ K D+
Sbjct: 724 KILSKRMQPLLRSIISPSQSAFVPGRAISDNVLITHKILHYLRQSGAKKHVSMAVKTDMT 545
Query: 863 KACGSVSWAFLRDTLCFFGFPPTSVSLIMFCVSSSSLSLLWNGVKLPPIAPSRGLR*GDP 922
KA ++W FLR+ L GF +S IM CVS+ S S L NG + PSRGLR GDP
Sbjct: 544 KAYDRIAWNFLREVLTRLGFHGIWISWIMECVSTVSYSFLINGGPQGRVLPSRGLRQGDP 365
Query: 923 LSPYLFVLCVERLTISIQDAVDQNQGELVRISRDGSPISHLFFADDVLLLAKAKVSQVRK 982
LSPYLF+LC E L+ Q A+ + V+++R+ PI+HL FADD + K+ S
Sbjct: 364 LSPYLFILCTEVLSGLCQQALRKGTLPGVKVARNCPPINHLLFADDTMFFGKSNASSCAI 185
Query: 983 VQNILADFCMASGLNVNIAKS 1003
+ +I+ + ASG +N KS
Sbjct: 184 LLSIMDKYRAASGRCIN*TKS 122
>BG585866
Length = 828
Score = 136 bits (343), Expect = 6e-32
Identities = 65/210 (30%), Positives = 112/210 (52%), Gaps = 1/210 (0%)
Frame = +3
Query: 1193 VMKTRDIIQDGFTWRIGRGDLSFWFSKCHGSKALCEKVPYVHISDTALRVRDVFQDGVCD 1252
+++ +++++ G+TWR G G+ SFW++ L + P+V I D L V+DVF G
Sbjct: 84 IIRDKNVLKSGYTWRAGSGNSSFWYTNWSSLGLLGTQAPFVDIHDLHLTVKDVFTTGGQH 263
Query: 1253 IRGLSTPITPELWALLQNLFVYVSLDDEDLVVWAPSLEGKYSVQSCYRWLLDM-EGGTAV 1311
+ L T + ++ ++ N + + D +W + G Y+ +S Y W+L E
Sbjct: 264 TQSLYTILPTDIAEVINNTHLNFNASIGDAYIWPHNSNGVYTAKSGYSWILSQTETVNYN 443
Query: 1312 SVDWA*LWKFQVPEKMKLFAWQAAHRSLPTVDTFHDRGVAIPRWCSLCHHHVET*MHCLR 1371
+ W+ +W+ ++PEK K F W A H ++PT+ + R + CS C H E+ HC+R
Sbjct: 444 NSSWSWIWRLKIPEKYKFFLWLACHNAVPTLSLLNHRNMVNSAICSRCGEHEESFFHCVR 623
Query: 1372 DCDASLQVWQNLCVSLTPGFFTASSFLDWI 1401
DC S +W + S +P FF++SS DW+
Sbjct: 624 DCRFSKIIWHKIGFS-SPDFFSSSSVQDWL 710
>BG647708 weakly similar to GP|13786450|gb| putative reverse transcriptase
{Oryza sativa}, partial (9%)
Length = 708
Score = 77.0 bits (188), Expect = 5e-14
Identities = 44/122 (36%), Positives = 67/122 (54%), Gaps = 4/122 (3%)
Frame = +1
Query: 913 PSRGLR*GDPLSPYLFVLCVERLTISIQDAVDQNQGELVRISRDGSPISHLFFADDVLLL 972
P +GLR GDPLSPYLF+LC L+ ++ ++ ++++R I+HL FADD LL
Sbjct: 4 PEKGLRQGDPLSPYLFILCANVLSGLLKREGNKQNLHGIQVARSDPKITHLLFADDSLLF 183
Query: 973 AKAKVSQVRKVQNILADFCMASGLNVNIAKSRDFSSVSVPRAKK----ADIFVVTSIPFT 1028
A+A +++ + +L + ASG VN KS S +VP +K I + T P +
Sbjct: 184 ARANLTEAATIMQVLHSYQSASGQLVNFEKSEVSYSQNVPNQEKEMICQQIAIKTGSPQS 363
Query: 1029 AN 1030
N
Sbjct: 364 VN 369
>BE248682 similar to GP|18568269|gb putative gag-pol polyprotein {Zea mays},
partial (1%)
Length = 441
Score = 72.8 bits (177), Expect = 1e-12
Identities = 50/133 (37%), Positives = 76/133 (56%), Gaps = 5/133 (3%)
Frame = +3
Query: 911 IAPSRGLR*GDPLSPYLFVLCVERLTISIQDAVDQNQGELVRISRDGSPISHLFFADDVL 970
I+ RGL+ GDPL+P+LF+L E ++ +++AV++N + + R G+ +SHL +ADD L
Sbjct: 12 ISVQRGLKQGDPLAPFLFLLVAEGISGLMKNAVNRNLFQGFDVKRGGTRVSHLQYADDTL 191
Query: 971 LLAKAKVSQVRKVQNILADFCMASGLNVNIAKSRDFSSVSVPR--AKKADIFV---VTSI 1025
+ V + ++ +L F MASGL VN KS ++VPR + A F+ SI
Sbjct: 192 CIGMPTVDNLWTLKALLQGFEMASGLKVNFHKS-SLIGINVPRDFMEAACRFLNCREESI 368
Query: 1026 PFTANLERYLGFP 1038
PF YLG P
Sbjct: 369 PFI-----YLGLP 392
>BQ148771
Length = 680
Score = 69.3 bits (168), Expect = 1e-11
Identities = 45/162 (27%), Positives = 77/162 (46%), Gaps = 10/162 (6%)
Frame = -3
Query: 1055 RVNRKFASWKGRLLNKAGKFILFKLVLNAIPVYPMQLFWFH*AVCDHLDSLAKNFIWSRG 1114
+V+ A+WK L+ A + L K V+ A+P+YPM A + + L + F+W
Sbjct: 588 QVHVMLANWKANHLSLARRVTLAKSVIEAVPLYPMMTTIIPKACIEEIQKLQRKFVWGDT 409
Query: 1115 E-RRGMNMVAWRTVTLPKCNGGLGIRKARLINTSMLGKLVWDLVNSHDKLWV*VLLDLYC 1173
E R + V W T++ PK GLG+R+ ++N + + KL W + + + L V+ Y
Sbjct: 408 EVSRRYHAVGWETMSKPKTIYGLGLRRLDVMNKACIMKLGWSIYSGSNSLCTEVMRGKYQ 229
Query: 1174 TNSD----FLHAPRRRGSCVWNAVMK-----TRDIIQDGFTW 1206
+ FL P S +W A++K R+++ W
Sbjct: 228 RSESLEEIFLEKP--TDSSLWKALVKLWPEIERNLVDSNGNW 109
>BG456581
Length = 683
Score = 61.6 bits (148), Expect = 2e-09
Identities = 38/119 (31%), Positives = 52/119 (42%), Gaps = 1/119 (0%)
Frame = +2
Query: 1290 EGKYSVQSCYRWLLDMEGGTAVSVDWA*-LWKFQVPEKMKLFAWQAAHRSLPTVDTFHDR 1348
+GK +++ Y +L WA +W +P W+ AH LPT D R
Sbjct: 5 QGKLTIKLVYSFLTSHTS----CAPWASTIWNSCIPPSHSFICWRLAHDRLPTDDNLSSR 172
Query: 1349 GVAIPRWCSLCHHHVET*MHCLRDCDASLQVWQNLCVSLTPGFFTASSFLDWIASVPRH 1407
G A+ CS C VET H C + +W LC L G SSF ++S+PRH
Sbjct: 173 GCALVSMCSFCLEQVETSDHLFLRCKFVVTLWSWLCSQLRVG-LDFSSFKALLSSLPRH 346
>TC83437 weakly similar to PIR|D86384|D86384 unknown protein [imported] -
Arabidopsis thaliana, partial (6%)
Length = 951
Score = 60.1 bits (144), Expect = 7e-09
Identities = 39/134 (29%), Positives = 64/134 (47%), Gaps = 4/134 (2%)
Frame = +2
Query: 960 ISHLFFADDVLLLAKAKVSQVRKVQNILADFCMASGLNVNIAKSR----DFSSVSVPRAK 1015
+SHL FA+D LLL + +R ++ L F SGL VN KS + + + A
Sbjct: 542 VSHLQFANDTLLLETKNWANIRALRAALVIF*AMSGLKVNFHKSGLVCVNIAPSWLSEAA 721
Query: 1016 KADIFVVTSIPFTANLERYLGFPIFQGRQTQEHFDFVLDRVNRKFASWKGRLLNKAGKFI 1075
+ V +PF YLG PI + ++ +++R+ + W R L+ G+ +
Sbjct: 722 SVLSWKVGKVPFL-----YLGMPIEGNSRRLSFWEPIVNRIKARLTGWNSRFLSFGGRLV 886
Query: 1076 LFKLVLNAIPVYPM 1089
L K VL ++ VY +
Sbjct: 887 LLKSVLTSLSVYAL 928
Score = 28.5 bits (62), Expect(2) = 1.8
Identities = 12/23 (52%), Positives = 16/23 (69%)
Frame = +1
Query: 855 VVFKLDLEKACGSVSWAFLRDTL 877
++FK+D EKA SV WA+L L
Sbjct: 304 LLFKVDFEKAYHSVDWAYLDSVL 372
Score = 21.9 bits (45), Expect(2) = 1.8
Identities = 12/26 (46%), Positives = 13/26 (49%)
Frame = +3
Query: 877 LCFFGFPPTSVSLIMFCVSSSSLSLL 902
L FGF SV L FC S S+L
Sbjct: 348 LGLFGFCSRSVCLFWFCGESGLKSVL 425
>BG647160 homologue to GP|15004435|gb| maturase {Ionopsis satyrioides}, partial
(2%)
Length = 780
Score = 59.7 bits (143), Expect = 9e-09
Identities = 33/102 (32%), Positives = 60/102 (58%), Gaps = 1/102 (0%)
Frame = +1
Query: 1467 GGVVRDAVGCWLFGYSGFIGMAE-ILKAELLAVLFGLQSCWDRGFRQLVVSSDSMVVLSL 1525
GGV+R+ + ++ G+SGFI + + I+ AEL + L+ ++V SDS++ ++L
Sbjct: 379 GGVIRNYLSTYITGFSGFISIYQNIMFAELTTLHQSLKLAISLNIEEMVCYSDSLLTVNL 558
Query: 1526 LRHGVTRFHVYAAIVGAIRELLRRDWRVELKHMLREGNVVAD 1567
++ +++ HVYA ++ I+ ++ L H LREGN AD
Sbjct: 559 IKEDISQHHVYAVLIHNIKYIM-SSRNFTLHHTLREGNQCAD 681
>BF650289 weakly similar to GP|9049283|dbj| orf129a {Beta vulgaris}, partial
(69%)
Length = 616
Score = 58.5 bits (140), Expect = 2e-08
Identities = 31/103 (30%), Positives = 57/103 (55%)
Frame = +3
Query: 803 TKVMVNRLRQFLQELIGPLQSSFIPGRGTIDNAIILQKIVHHMNSRRGKCQNVVFKLDLE 862
+K++ +R++ L ++ QS+F+ GR DN I+ ++V SR+G + K+DL
Sbjct: 294 SKILTSRMQGVLNSVVSENQSAFVKGRVIFDNIILSHELVKSY-SRKGISPRCMVKIDLX 470
Query: 863 KACGSVSWAFLRDTLCFFGFPPTSVSLIMFCVSSSSLSLLWNG 905
KA S W F++ + GFP V+ +M ++++S + NG
Sbjct: 471 KAYDSXEWPFIKHLMLELGFPYKFVNWVMAXLTTASYTFNXNG 599
>BG586862
Length = 804
Score = 57.0 bits (136), Expect = 6e-08
Identities = 34/142 (23%), Positives = 59/142 (40%), Gaps = 4/142 (2%)
Frame = -1
Query: 1318 LWKFQVPEKMKLFAWQAAHRSLPTVDTFHDRGVAIPRWCSLCHHHVET*MHCLRDCDASL 1377
+W + + K F W+ H +LP D H RG+ C C +ET H +C+ +
Sbjct: 654 VWGIKTIPRHKSFLWRLLHNALPVKDELHKRGIRCSLLCPRCESKIETVQHLFLNCEVTQ 475
Query: 1378 QVWQNLCVSLTPGFFTASSFLDWIAS-VPRHD---VAGLLVHAWFIWCRRRRFVFAGELE 1433
+ W + + F DWI + + ++D + L + IW R + VF
Sbjct: 474 KEWFGSQLGINFHSSGVLHFHDWITNFILKNDEETIIALTALLYSIWHARNQKVFENIDV 295
Query: 1434 PWRLILHRAADVLRCLRATSPS 1455
P +++ RA+ L + S
Sbjct: 294 PGDVVIQRASSSLHSFKMAQVS 229
>TC83226 weakly similar to PIR|G86419|G86419 probable reverse transcriptase
100033-105622 [imported] - Arabidopsis thaliana, partial
(2%)
Length = 885
Score = 56.6 bits (135), Expect = 7e-08
Identities = 33/113 (29%), Positives = 47/113 (41%), Gaps = 10/113 (8%)
Frame = +3
Query: 1284 VWAPSLEGKYSVQSCYR----WLLDMEGGTAVSVD----WA*LWKFQVPEKMKLFAWQAA 1335
+W + G YSV+S Y W T+ S D W +W + K+ W+
Sbjct: 3 MWMHNPTGIYSVKSGYNTLRTWQTQQINNTSTSSDETLIWKKIWSLHTIPRHKVLLWRIL 182
Query: 1336 HRSLPTVDTFHDRGVAIPRWCSLCHHHVET*MHCLRDCDASLQVW--QNLCVS 1386
+ SLP + RG+ C CH ET H C S +VW NLC++
Sbjct: 183 NDSLPVRSSLRKRGIQCYPLCPRCHSKTETITHLFMSCPLSKRVWFGSNLCIN 341
>TC91765 weakly similar to GP|19881779|gb|AAM01180.1 Putative retroelement
{Oryza sativa (japonica cultivar-group)}, partial (1%)
Length = 625
Score = 55.8 bits (133), Expect = 1e-07
Identities = 29/69 (42%), Positives = 40/69 (57%)
Frame = +2
Query: 893 CVSSSSLSLLWNGVKLPPIAPSRGLR*GDPLSPYLFVLCVERLTISIQDAVDQNQGELVR 952
CV S+ +L N + PI PSRGL+ GD LSPY+F++CVE L+ I A ++
Sbjct: 104 CVESNDYYVLVNNDAVDPIIPSRGLQQGDHLSPYIFIICVEGLSFLIPHAKERGDTHGTS 283
Query: 953 ISRDGSPIS 961
I R P+S
Sbjct: 284 I*RGAPPVS 310
Score = 33.5 bits (75), Expect = 0.67
Identities = 27/105 (25%), Positives = 44/105 (41%)
Frame = +3
Query: 955 RDGSPISHLFFADDVLLLAKAKVSQVRKVQNILADFCMASGLNVNIAKSRDFSSVSVPRA 1014
R G+ L F L + + + ++NIL + SG +++ KS + S +VP
Sbjct: 255 RRGAIPMALLFEGAPLRSLRVEEHHAQIMKNILILYEEDSGKAISLRKS*IYCSRNVPDI 434
Query: 1015 KKADIFVVTSIPFTANLERYLGFPIFQGRQTQEHFDFVLDRVNRK 1059
K I + + F +YLG P GR F + V +K
Sbjct: 435 LKTSITYILGVQFMLGTCKYLGLPSMIGRDRTTTFSSIKGGVWQK 569
>BQ122106 similar to GP|9366656|emb|C probable similar to ring-h2 finger
protein rha1a. {Trypanosoma brucei}, partial (18%)
Length = 693
Score = 54.7 bits (130), Expect = 3e-07
Identities = 35/101 (34%), Positives = 55/101 (53%), Gaps = 1/101 (0%)
Frame = -2
Query: 1468 GVVRDAVGCWLFGYSGFI-GMAEILKAELLAVLFGLQSCWDRGFRQLVVSSDSMVVLSLL 1526
G++R+ G + G+ G I ++IL AEL A+ GL+ D G V DS+ +SL+
Sbjct: 488 GLIRNIAGLFNSGFPGNITNTSDILLAELHAIFQGLRMISDMGISDFVCYFDSLHYVSLI 309
Query: 1527 RHGVTRFHVYAAIVGAIRELLRRDWRVELKHMLREGNVVAD 1567
+FHVYA ++ I++L+ + + H L EGN AD
Sbjct: 308 NGPSMKFHVYATLIQDIKDLVITS-KASVFHTLCEGNYCAD 189
>BG585499
Length = 792
Score = 52.4 bits (124), Expect = 1e-06
Identities = 37/147 (25%), Positives = 60/147 (40%), Gaps = 10/147 (6%)
Frame = +3
Query: 1292 KYSVQSCYRWLLDMEGGTAVSVDWA*LWKFQVPEKMKLFAWQAAHRSLPTVDTFHDRGVA 1351
++ ++S Y L+ + + V DW LW ++ P + + F W AH + T G
Sbjct: 162 QFKIKSSYNLLV*DQ--SIVDCDWKMLWGWRGPHRTQTFMWLVAHGCILTNYRRSRWGTR 335
Query: 1352 IPRWCSLCHHHVET*MHCLRDCDASLQVWQNLCVS-LTPGFFTASSFLDWIASVPRHDVA 1410
+ C C + ET +H L DC + QVW L S FF+ DW+
Sbjct: 336 VLATCPCCGNADETVLHVLCDCRPASQVWIRLVPSDWITNFFSFDDCRDWVFKNLSKRSN 515
Query: 1411 GL---------LVHAWFIWCRRRRFVF 1428
G+ + W +W R + +F
Sbjct: 516 GVSKFKWQPTFMTTCWHMWTWRNKAIF 596
>TC77455 similar to GP|22335695|dbj|BAC10549. nine-cis-epoxycarotenoid
dioxygenase1 {Pisum sativum}, partial (43%)
Length = 1865
Score = 51.2 bits (121), Expect = 3e-06
Identities = 22/44 (50%), Positives = 28/44 (63%)
Frame = -3
Query: 1129 LPKCNGGLGIRKARLINTSMLGKLVWDLVNSHDKLWV*VLLDLY 1172
LP+C GGLG+R RL+N S+L K W L+ LW VL D+Y
Sbjct: 966 LPRCKGGLGVRDIRLVNVSLLAKWWWRLLQDQSSLWKEVLEDIY 835
Score = 45.1 bits (105), Expect = 2e-04
Identities = 33/107 (30%), Positives = 45/107 (41%), Gaps = 8/107 (7%)
Frame = -2
Query: 1281 DLVVWAPSLEGKYSVQSCY-----RWLLDMEGGTAVSVDWA*LWKFQVPEKMKLFAWQAA 1335
D+ VW P EG +SV SCY LL+ V + LWK + P K+ F+W
Sbjct: 448 DIWVWKPDKEGVFSVNSCYFLLQNLRLLEDRLSYEEEVIFRELWKSKAPAKVLAFSWTLF 269
Query: 1336 HRSLPTVDTFHDR---GVAIPRWCSLCHHHVET*MHCLRDCDASLQV 1379
+PT+ R V + C C ET +H CD +V
Sbjct: 268 LDRIPTMVNLGKRRLLRVEDSKRCVFCGCQDETVVHLFLHCDVISKV 128
>TC83398 similar to PIR|T05150|T05150 hypothetical protein F18E5.40 -
Arabidopsis thaliana, partial (6%)
Length = 766
Score = 49.7 bits (117), Expect = 9e-06
Identities = 33/89 (37%), Positives = 49/89 (54%), Gaps = 1/89 (1%)
Frame = +2
Query: 1480 GYSGFIGMA-EILKAELLAVLFGLQSCWDRGFRQLVVSSDSMVVLSLLRHGVTRFHVYAA 1538
G+SG I + +IL AEL A+L GLQ +V SDS+ ++L+ +H YA
Sbjct: 104 GFSGHIDHSNDILFAELHAILMGLQLAQTLNIVDVVCYSDSLHYVNLINGPSVVYHAYAT 283
Query: 1539 IVGAIRELLRRDWRVELKHMLREGNVVAD 1567
++ I++L+ R+ H LREGN AD
Sbjct: 284 LIQDIKDLI----RLSKLHTLREGNRCAD 358
>BG587113 weakly similar to PIR|A84888|A8 hypothetical protein At2g45230
[imported] - Arabidopsis thaliana, partial (10%)
Length = 767
Score = 47.4 bits (111), Expect = 4e-05
Identities = 34/119 (28%), Positives = 50/119 (41%), Gaps = 9/119 (7%)
Frame = -3
Query: 1280 EDLVVWAPSLEGKYSVQSCYRWLLDM-----EGGTA--VSVD--WA*LWKFQVPEKMKLF 1330
ED W S G YSV+S Y ++ + GT S+D + +WK+ K++ F
Sbjct: 429 EDSYSWEYSKSGHYSVKSGYYVQTNIIAAANQRGTVDQPSLDDLYQRVWKYNTSPKVRHF 250
Query: 1331 AWQAAHRSLPTVDTFHDRGVAIPRWCSLCHHHVET*MHCLRDCDASLQVWQNLCVSLTP 1389
W+ SLPT R ++ CS C ET H L C + +W + P
Sbjct: 249 LWRCISNSLPTAANMRSRHISKDGSCSRCGMESETVNHILFQCPYARLIWATSPIHAPP 73
>BG447966 weakly similar to PIR|G96509|G96 protein F27F5.21 [imported] -
Arabidopsis thaliana, partial (7%)
Length = 687
Score = 35.8 bits (81), Expect(2) = 2e-04
Identities = 17/35 (48%), Positives = 25/35 (70%)
Frame = +2
Query: 803 TKVMVNRLRQFLQELIGPLQSSFIPGRGTIDNAII 837
TKV+ NR++Q L ++I QS+F+ GR DNA+I
Sbjct: 527 TKVIANRVKQTLPDVIDVEQSAFVQGRLITDNALI 631
Score = 28.5 bits (62), Expect(2) = 2e-04
Identities = 23/66 (34%), Positives = 29/66 (43%)
Frame = +3
Query: 731 RVLMAFNRFSSSSIGILWGMMFGVWFVMLLSRVLLTLSWPRL*FVLSLRLTSPLPSGSSG 790
RV MA SS SIGILWG W + V S + LSL+ + +
Sbjct: 312 RVQMASLHCSSKSIGILWGRKCNKWSSKSSTTVWKLKSLIKPSLSLSLKERIRILQRITA 491
Query: 791 QSAYAM 796
Q A+AM
Sbjct: 492 QLAFAM 509
>BF006335 similar to GP|14334538|gb| unknown protein {Arabidopsis thaliana},
partial (3%)
Length = 505
Score = 40.0 bits (92), Expect(2) = 8e-04
Identities = 26/80 (32%), Positives = 41/80 (50%)
Frame = +2
Query: 1467 GGVVRDAVGCWLFGYSGFIGMAEILKAELLAVLFGLQSCWDRGFRQLVVSSDSMVVLSLL 1526
GG+ R+ + FG+ G IG + IL AE+ +L GL+ W R+ + DS+ V+ L
Sbjct: 215 GGLDRNYDKAFQFGFYGSIGWSNILHAEIQDLLVGLKLWWKA--RKFICYYDSLHVVQLG 388
Query: 1527 RHGVTRFHVYAAIVGAIREL 1546
FH YA ++ + L
Sbjct: 389 SMNTHHFHHYANMLEIYQSL 448
Score = 22.3 bits (46), Expect(2) = 8e-04
Identities = 8/21 (38%), Positives = 11/21 (52%)
Frame = +3
Query: 1543 IRELLRRDWRVELKHMLREGN 1563
IR +++DW L EGN
Sbjct: 438 IRAYMKKDWNFSLHQTFPEGN 500
>TC93101
Length = 675
Score = 43.1 bits (100), Expect = 8e-04
Identities = 23/100 (23%), Positives = 44/100 (44%), Gaps = 4/100 (4%)
Frame = -1
Query: 1336 HRSLPTVDTFHDRGVAIPRWCSLCHHHVET*MHCLRDCDASLQVWQNLCVSLTPGFFTAS 1395
H +LP + RGV P C C+ ++ET H C+ + +VW +S+ +
Sbjct: 672 HTTLPVRXELNKRGVNCPPLCPRCYFNLETTNHIFMSCERTQRVWFGSQLSIRFPDNSTI 493
Query: 1396 SFLDWIASVPRHDVAGLLVH----AWFIWCRRRRFVFAGE 1431
+F DW+ + +++ + IW R + +F +
Sbjct: 492 NFSDWLFDAISNQTEEIIIKISAITYSIWHARNKAIFENQ 373
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.338 0.147 0.507
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 60,225,199
Number of Sequences: 36976
Number of extensions: 1042909
Number of successful extensions: 7623
Number of sequences better than 10.0: 61
Number of HSP's better than 10.0 without gapping: 4272
Number of HSP's successfully gapped in prelim test: 357
Number of HSP's that attempted gapping in prelim test: 3143
Number of HSP's gapped (non-prelim): 5022
length of query: 1569
length of database: 9,014,727
effective HSP length: 109
effective length of query: 1460
effective length of database: 4,984,343
effective search space: 7277140780
effective search space used: 7277140780
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (21.8 bits)
S2: 65 (29.6 bits)
Lotus: description of TM0022.23


