

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
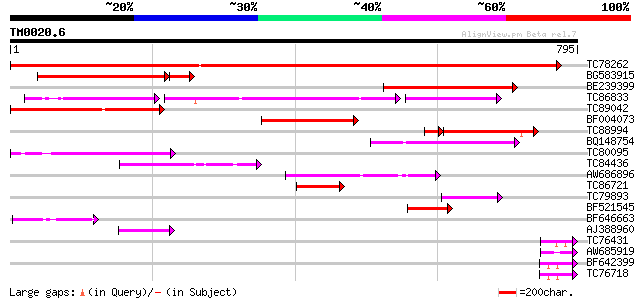
Query= TM0020.6
(795 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC78262 similar to GP|20260620|gb|AAM13208.1 unknown protein {Ar... 917 0.0
BG583915 similar to PIR|C86445|C86 hypothetical protein AAG23449... 318 3e-96
BE239399 similar to PIR|C86445|C86 hypothetical protein AAG23449... 343 1e-94
TC86833 weakly similar to PIR|H86427|H86427 unknown protein [imp... 197 2e-91
TC89042 similar to GP|18176281|gb|AAL60016.1 unknown protein {Ar... 265 5e-71
BF004073 similar to GP|11994398|db gb|AAD36947.1~gene_id:MIL23.1... 192 6e-49
TC88994 similar to GP|20260620|gb|AAM13208.1 unknown protein {Ar... 155 1e-44
BQ148754 weakly similar to GP|15146264|gb At1g10080/T27I1_10 {Ar... 146 3e-35
TC80095 weakly similar to GP|12597787|gb|AAG60099.1 unknown prot... 127 2e-29
TC84436 weakly similar to GP|498707|emb|CAA55187.1| HYP1 {Arabid... 124 2e-28
AW686896 similar to GP|12597787|gb unknown protein {Arabidopsis ... 112 4e-25
TC86721 similar to GP|9757997|dbj|BAB08419.1 gene_id:MFH8.10~pir... 79 5e-15
TC79893 weakly similar to GP|12597787|gb|AAG60099.1 unknown prot... 75 1e-13
BF521545 similar to GP|498707|emb|C HYP1 {Arabidopsis thaliana},... 63 4e-10
BF646663 similar to GP|6714477|gb|A hypothetical protein {Arabid... 58 2e-08
AJ388960 weakly similar to GP|14596049|gb| Unknown protein {Arab... 44 2e-04
TC76431 similar to GP|8132441|gb|AAF73291.1| extensin {Pisum sat... 44 3e-04
AW685919 homologue to PIR|S25298|S252 extensin (clone Tom J-10) ... 43 4e-04
BF642399 similar to PIR|T10863|T10 extensin precursor - kidney b... 42 7e-04
TC76718 homologue to GP|8132441|gb|AAF73291.1| extensin {Pisum s... 42 7e-04
>TC78262 similar to GP|20260620|gb|AAM13208.1 unknown protein {Arabidopsis
thaliana}, partial (93%)
Length = 3014
Score = 917 bits (2369), Expect = 0.0
Identities = 438/775 (56%), Positives = 589/775 (75%), Gaps = 2/775 (0%)
Frame = +3
Query: 1 MATLADIGVSAAINILSAFAFLLAFALLRIQPINDRVYFPKWYINGERTSPRSTGNFVGK 60
MA LADI ++A INILSAF F +AFA+LR+QP+NDRVYFPKWY+ G RT P G F+ K
Sbjct: 285 MAKLADISLAAGINILSAFIFFVAFAILRLQPLNDRVYFPKWYLKGLRTDPVHGGAFMRK 464
Query: 61 FVNLNFKTYLTFLNWMPQALRMSENEIINHAGLDSAVFLRIYTLGLKIFVPVTVVALLIL 120
VNL++++Y+ FLNWMP ALRM E E+I+HAGLDSAV+LRIY LGLKIFVP+ +A +L
Sbjct: 465 IVNLDWRSYIRFLNWMPAALRMPEPELIDHAGLDSAVYLRIYLLGLKIFVPIAFLAWAVL 644
Query: 121 IPVN-VSSGALFFLKRDLVVSDIDKLSISNVPSKSSRFFVHIGLEYMFTLWICFLLYKEY 179
+PVN SSG +++ SDIDK+SISNV S RF+ HI + Y FT W C+ L KEY
Sbjct: 645 VPVNWTSSGLENAGIKNITSSDIDKISISNVQRGSERFWSHIVVAYAFTFWTCYTLMKEY 824
Query: 180 DNIAMMRLHFLASQRRRVEQFTVVVRSVPHISGHSVSDSVESFFQTNHPNHYIGHQAVYN 239
+ MRL FLA+++RR +QFTV+VR++P + SV + VE FF NHP++Y+ HQ VYN
Sbjct: 825 GKVTAMRLQFLATEKRRPDQFTVLVRNIPPDTDESVGELVEHFFLVNHPDNYLTHQVVYN 1004
Query: 240 ANKFAKLVRRRDRLQNWLDYYQLKFQRHPDKRPTINTGVLGLWGRKVDAIEYYSHTIKEL 299
ANK K V+++ +LQNWL YYQ K +R KRP + TG LGL G+KVDAI+YY+ I +L
Sbjct: 1005 ANKLEKFVKKKSKLQNWLVYYQNKLER-TSKRPEMKTGFLGLHGKKVDAIDYYTTEIDKL 1181
Query: 300 DKVMTLERQRIIKDPKCILPVAFLSFSSRWGASVCAQTQQSKNPTLWLTDWAPEPRDIYW 359
K + LER ++ DPK +P AF+SF SRWGA+VCAQTQQ++NPT+WLT+WAPEPRD+YW
Sbjct: 1182 SKEIALERDKVTNDPKSTMPAAFVSFKSRWGAAVCAQTQQTRNPTIWLTEWAPEPRDVYW 1361
Query: 360 RNLAIPFVSLSIRKLVISLSVFALVFFYMIPIAFVQSLANLEGLERVAPFLRPVIELKFI 419
+NLAIP+VSL++R+L+I+++ F L FF+MIPIA VQ LA+L+G+++ AP+L P++ + +
Sbjct: 1362 QNLAIPYVSLTVRRLIIAVAFFFLTFFFMIPIAIVQGLASLDGIQKAAPWLNPLVRVPVV 1541
Query: 420 KSFLQGFLPGLALKIFLYILPAVLMIMSKIEGHIAKSTLERKTAAKYYYFMLVNVFLGSI 479
SF+QGFLPG+ LK+FL LP +LM+MSK EG + S+LER++A++YY F VN+FLG++
Sbjct: 1542 MSFIQGFLPGIVLKLFLIFLPTILMMMSKFEGFGSISSLERRSASRYYLFCFVNIFLGNL 1721
Query: 480 VTGTAFEQLHSFLHQSPTQIPRTIGVSIPMKATFFMTYIMVDGWAGIAGEILRLKPLVIY 539
+ G+AF+QL +F+HQ + P TIG +IP+KA+FF+TYIMVDGW+GIA E+L LKPL++Y
Sbjct: 1722 LAGSAFQQLDTFIHQPANEYPITIGTAIPLKASFFITYIMVDGWSGIAAEVLMLKPLIMY 1901
Query: 540 HLKNMFIVKTERDRGKAMDPGSVDFPETIPSLQLYFLLGIVYAVVTPILLPFILVFFALA 599
HLKN F+VKTE+DR +AM+PGS+ F P +QLYFLLG+VYA VTP +LPFI++FF LA
Sbjct: 1902 HLKNFFLVKTEKDREEAMNPGSIGFNTGEPRIQLYFLLGLVYAAVTPTVLPFIIIFFGLA 2081
Query: 600 YLVYRHQIINVYNQQYESAAAFWPHVHSRIIASLLISQLLMLGLLSTKKAAKSTPLLAIL 659
Y+V+RHQIINVYNQ+YES AAFWP VH R+I +LL+SQ++++GLL+TKKAA STP L +L
Sbjct: 2082 YVVFRHQIINVYNQEYESGAAFWPDVHFRVIIALLVSQIVLMGLLTTKKAASSTPFLIVL 2261
Query: 660 PILTFAFHKYCQHRFEPAFRKYPLEEAMAKDLLEKTTEPDLNIKAYLADAYLHPIFRSFE 719
PILT FH+YC+ RFE AF K+PL+EAM KD LE+ TEP+LN+K YL AY+HP+F++
Sbjct: 2262 PILTIWFHRYCKGRFESAFVKFPLQEAMMKDTLERATEPNLNVKGYLQHAYVHPVFKASH 2441
Query: 720 VEEELVEVRVDKQPETQVASPTLSEPSSPS-PLHDHVHQPSPPQHVNQYSHYPTS 773
++ E + + ET+ A+ S S PL S P ++ + P S
Sbjct: 2442 DDDADEEDAMSLKWETESATVATKRQSRRSTPLPSRFSGASSPSMLDSIKNDPES 2606
>BG583915 similar to PIR|C86445|C86 hypothetical protein AAG23449.1
[imported] - Arabidopsis thaliana, partial (20%)
Length = 667
Score = 318 bits (816), Expect(2) = 3e-96
Identities = 157/185 (84%), Positives = 172/185 (92%)
Frame = +2
Query: 40 PKWYINGERTSPRSTGNFVGKFVNLNFKTYLTFLNWMPQALRMSENEIINHAGLDSAVFL 99
PKWYI+G R++PRS+ NFVGKFVNLNFKTYLTFLNWMPQALRMSE EIINHAGLDSAVFL
Sbjct: 2 PKWYISGGRSNPRSSANFVGKFVNLNFKTYLTFLNWMPQALRMSETEIINHAGLDSAVFL 181
Query: 100 RIYTLGLKIFVPVTVVALLILIPVNVSSGALFFLKRDLVVSDIDKLSISNVPSKSSRFFV 159
RIYTLGLK+F+PVT+VALLILIPVNVSSG LFFL+R+LVVSDIDKLSISNVP KS RFFV
Sbjct: 182 RIYTLGLKMFIPVTIVALLILIPVNVSSGTLFFLRRELVVSDIDKLSISNVPPKSLRFFV 361
Query: 160 HIGLEYMFTLWICFLLYKEYDNIAMMRLHFLASQRRRVEQFTVVVRSVPHISGHSVSDSV 219
HIGLEYM T+ F KEYDN+A+MRLHFLASQRRRVEQFTVVVR+VPHISGHSVSDSV
Sbjct: 362 HIGLEYMLTIMDLFSALKEYDNVALMRLHFLASQRRRVEQFTVVVRNVPHISGHSVSDSV 541
Query: 220 ESFFQ 224
+SFF+
Sbjct: 542 DSFFK 556
Score = 52.8 bits (125), Expect(2) = 3e-96
Identities = 23/35 (65%), Positives = 25/35 (70%)
Frame = +1
Query: 225 TNHPNHYIGHQAVYNANKFAKLVRRRDRLQNWLDY 259
TNHP+HYIGHQ K VR+RDRLQNWLDY
Sbjct: 556 TNHPDHYIGHQLSTMQIDSLKFVRKRDRLQNWLDY 660
>BE239399 similar to PIR|C86445|C86 hypothetical protein AAG23449.1
[imported] - Arabidopsis thaliana, partial (23%)
Length = 565
Score = 343 bits (880), Expect = 1e-94
Identities = 168/188 (89%), Positives = 181/188 (95%)
Frame = +2
Query: 524 AGIAGEILRLKPLVIYHLKNMFIVKTERDRGKAMDPGSVDFPETIPSLQLYFLLGIVYAV 583
AGIAGEILRLKPLVIYHLKNMFIVKTERDRGKAMDPGSV+FPET+PSLQLYFLLGIVYAV
Sbjct: 2 AGIAGEILRLKPLVIYHLKNMFIVKTERDRGKAMDPGSVEFPETLPSLQLYFLLGIVYAV 181
Query: 584 VTPILLPFILVFFALAYLVYRHQIINVYNQQYESAAAFWPHVHSRIIASLLISQLLMLGL 643
+TPILLPFILVFFA AYLVYRHQIINVY+QQYESAAAFWP VHSRIIASL++SQ+L+ GL
Sbjct: 182 MTPILLPFILVFFAFAYLVYRHQIINVYHQQYESAAAFWPQVHSRIIASLILSQILLFGL 361
Query: 644 LSTKKAAKSTPLLAILPILTFAFHKYCQHRFEPAFRKYPLEEAMAKDLLEKTTEPDLNIK 703
LSTKKA KSTPLL +LPILTFAFHKYC+ RFEPAF KYP+EEAMAKD+LEKTTEPDLNIK
Sbjct: 362 LSTKKAVKSTPLLIMLPILTFAFHKYCKRRFEPAFXKYPVEEAMAKDILEKTTEPDLNIK 541
Query: 704 AYLADAYL 711
AYLAD+YL
Sbjct: 542 AYLADSYL 565
>TC86833 weakly similar to PIR|H86427|H86427 unknown protein [imported] -
Arabidopsis thaliana, partial (90%)
Length = 2640
Score = 197 bits (500), Expect(3) = 2e-91
Identities = 112/338 (33%), Positives = 190/338 (56%), Gaps = 6/338 (1%)
Frame = +2
Query: 217 DSVESFFQTNHPNHYIGHQAVYNANKFAKLVRRRDRLQNWLD-----YYQLKFQRHPD-K 270
+ V+S+F+ +P + + + K K+ + + L Y K P+
Sbjct: 878 EQVDSYFKAIYPETFYRSMIITDNKKVNKIWEELEGYKKKLARAEVVYAGSKTTAKPEGT 1057
Query: 271 RPTINTGVLGLWGRKVDAIEYYSHTIKELDKVMTLERQRIIKDPKCILPVAFLSFSSRWG 330
RPT TG LGL G+KVD+IEY + I EL + E++ +++ + + F FS+R
Sbjct: 1058 RPTNKTGCLGLIGKKVDSIEYCNEKINELVAKLESEQKVTLREKQQNAAIVF--FSNRVI 1231
Query: 331 ASVCAQTQQSKNPTLWLTDWAPEPRDIYWRNLAIPFVSLSIRKLVISLSVFALVFFYMIP 390
A+ AQ+ ++ W APEP + W NL I + +R+ ++ V +FFYM+P
Sbjct: 1232 AASAAQSLHAQVVDHWSVFGAPEPCQLLWPNLKIKYFQRELRQYLVYFIVTLAIFFYMVP 1411
Query: 391 IAFVQSLANLEGLERVAPFLRPVIELKFIKSFLQGFLPGLALKIFLYILPAVLMIMSKIE 450
I FV + L+ LE++ PF++P++++ +K+ L+ +LP LAL IFL +LP +LM +SK+E
Sbjct: 1412 ITFVSAFTTLKSLEKLLPFIKPIVKIITLKTVLEAYLPQLALIIFLAMLPKLLMFLSKLE 1591
Query: 451 GHIAKSTLERKTAAKYYYFMLVNVFLGSIVTGTAFEQLHSFLHQSPTQIPRTIGVSIPMK 510
G +S R + KY+YF ++NVF+G ++GT F+ + P I + S+P +
Sbjct: 1592 GIPTESHAARAASGKYFYFTVLNVFIGVTLSGTLFDTFKR-IQNKPKDIVPVLAESLPGR 1768
Query: 511 ATFFMTYIMVDGWAGIAGEILRLKPLVIYHLKNMFIVK 548
ATFF+T++ + + G E+ RL PL+IY+LK F+++
Sbjct: 1769 ATFFLTFVALKFFVGYGLELSRLVPLIIYNLKKKFLLQ 1882
Score = 95.9 bits (237), Expect(3) = 2e-91
Identities = 47/135 (34%), Positives = 80/135 (58%)
Frame = +1
Query: 555 KAMDPGSVDFPETIPSLQLYFLLGIVYAVVTPILLPFILVFFALAYLVYRHQIINVYNQQ 614
+A PG + + IP+ L + + Y+ + P+++PF ++F L +LV R+Q + VY +
Sbjct: 1903 EAWAPGDLGYATRIPADMLIVTIVLCYSCIAPLIIPFGALYFGLGWLVLRNQALKVYVPR 2082
Query: 615 YESAAAFWPHVHSRIIASLLISQLLMLGLLSTKKAAKSTPLLAILPILTFAFHKYCQHRF 674
YES WPH+++RI+AS+++ Q+ M G ++ + PLL LPILT F C +F
Sbjct: 2083 YESYGRMWPHINNRILASMVLYQVTMFGYFGVQQFVYA-PLLIPLPILTVLFGFICSKKF 2259
Query: 675 EPAFRKYPLEEAMAK 689
P+F+ LE A ++
Sbjct: 2260 YPSFQHQALEVAASE 2304
Score = 84.0 bits (206), Expect(3) = 2e-91
Identities = 58/190 (30%), Positives = 99/190 (51%), Gaps = 1/190 (0%)
Frame = +1
Query: 22 LLAFALLRIQPINDRVYFPKWYINGERTSPRSTGNFVGKFVNLNFKTYLTFLNWMPQALR 81
++ FALL+ +P N+ VY+P + G P G+ KT F +W+ +A
Sbjct: 325 MILFALLQSKPGNNVVYYPNRILKG--LDPFEGGS----------KTRNPF-SWIKEAFS 465
Query: 82 MSENEIINHAGLDSAVFLRIYTLGLKIFVPVTVVALLILIPVNVSSGALFFLKR-DLVVS 140
SE ++I +GLD+AVF + I V ++ L +L+P+ V+ GA L + +
Sbjct: 466 SSEQDVIAMSGLDTAVFFVFLSTVFSILVICGIILLPVLLPIAVTGGAGKKLTTSEGTFN 645
Query: 141 DIDKLSISNVPSKSSRFFVHIGLEYMFTLWICFLLYKEYDNIAMMRLHFLASQRRRVEQF 200
++D+LS+ N+ +KS R + Y +L FLL+K Y +++ +R S + EQF
Sbjct: 646 ELDQLSMGNITAKSVRLWAFFIACYFVSLVSLFLLWKAYKHVSWLRTKAFKSIDVKPEQF 825
Query: 201 TVVVRSVPHI 210
+VVR +P +
Sbjct: 826 AIVVRDIPPV 855
>TC89042 similar to GP|18176281|gb|AAL60016.1 unknown protein {Arabidopsis
thaliana}, partial (53%)
Length = 770
Score = 265 bits (677), Expect = 5e-71
Identities = 129/216 (59%), Positives = 171/216 (78%)
Frame = +2
Query: 1 MATLADIGVSAAINILSAFAFLLAFALLRIQPINDRVYFPKWYINGERTSPRSTGNFVGK 60
MA+L DIG++AAINIL+A FLLAFA+LRIQPINDRVYFPKWY+ G R+SP G FV K
Sbjct: 128 MASLGDIGLAAAINILTAIVFLLAFAILRIQPINDRVYFPKWYMKGLRSSPLQGGAFVSK 307
Query: 61 FVNLNFKTYLTFLNWMPQALRMSENEIINHAGLDSAVFLRIYTLGLKIFVPVTVVALLIL 120
FVN++F++Y+ FLNWMP AL+M E E+I HAGLDSAV+LRIY LGLKIFVP++++A ++
Sbjct: 308 FVNIDFRSYIRFLNWMPAALQMPEPELIEHAGLDSAVYLRIYLLGLKIFVPISLLAFSVM 487
Query: 121 IPVNVSSGALFFLKRDLVVSDIDKLSISNVPSKSSRFFVHIGLEYMFTLWICFLLYKEYD 180
+PVN ++ L + ++V + IDKLSISN+P S+RF+ H+ + Y FT W C++L +EY
Sbjct: 488 VPVNWTNDTL--KRSNVVYTSIDKLSISNIPLGSNRFWTHLVMAYAFTFWTCYILKREYQ 661
Query: 181 NIAMMRLHFLASQRRRVEQFTVVVRSVPHISGHSVS 216
+A MRL FLAS+RRR +QFTV+VR+VP + SVS
Sbjct: 662 IVAAMRLSFLASERRRPDQFTVLVRNVPPDADESVS 769
>BF004073 similar to GP|11994398|db gb|AAD36947.1~gene_id:MIL23.19~strong
similarity to unknown protein {Arabidopsis thaliana},
partial (18%)
Length = 424
Score = 192 bits (487), Expect = 6e-49
Identities = 90/136 (66%), Positives = 114/136 (83%)
Frame = +3
Query: 353 EPRDIYWRNLAIPFVSLSIRKLVISLSVFALVFFYMIPIAFVQSLANLEGLERVAPFLRP 412
EPRDIYW N+AIP+ SLSIR+LVI ++ F L FF+MIPIAFVQSLAN+EG+E+ APFL+
Sbjct: 3 EPRDIYWDNMAIPYFSLSIRRLVIGVAFFFLTFFFMIPIAFVQSLANIEGIEKAAPFLKS 182
Query: 413 VIELKFIKSFLQGFLPGLALKIFLYILPAVLMIMSKIEGHIAKSTLERKTAAKYYYFMLV 472
+IE+ IKSF+QGFLPG+ALK+FL LP + MIMSK G I +S+LER+ A++YY F +
Sbjct: 183 IIEIHVIKSFIQGFLPGIALKLFLIFLPTISMIMSKFAGFIIQSSLERRCASRYYIFQFI 362
Query: 473 NVFLGSIVTGTAFEQL 488
NVFLGSI+TGTAF+QL
Sbjct: 363 NVFLGSIITGTAFQQL 410
>TC88994 similar to GP|20260620|gb|AAM13208.1 unknown protein {Arabidopsis
thaliana}, partial (16%)
Length = 888
Score = 155 bits (391), Expect(2) = 1e-44
Identities = 75/143 (52%), Positives = 101/143 (70%), Gaps = 10/143 (6%)
Frame = +2
Query: 609 NVYNQQYESAAAFWPHVHSRIIASLLISQLLMLGLLSTKKAAKSTPLLAILPILTFAFHK 668
NVYNQ+YESA AFWP VH RI+ +L++SQLL++GLLSTK+AA STPLL LP+LT FH+
Sbjct: 89 NVYNQEYESAGAFWPDVHGRIVFALVVSQLLLMGLLSTKEAANSTPLLIALPVLTIWFHR 268
Query: 669 YCQHRFEPAFRKYPLEEAMAKDLLEKTTEPDLNIKAYLADAYLHPIFR----------SF 718
+C+ +EPAF +PL+EAM KD LE+T EP+ N+K +L DAY+HP+F S
Sbjct: 269 FCKGSYEPAFTTHPLQEAMVKDTLERTKEPNFNLKDFLHDAYIHPVFNDDGDTDSDVMSQ 448
Query: 719 EVEEELVEVRVDKQPETQVASPT 741
E +EE V V+ +Q +P+
Sbjct: 449 EWKEEPVIVQTKRQSRKNTPAPS 517
Score = 44.3 bits (103), Expect(2) = 1e-44
Identities = 20/27 (74%), Positives = 23/27 (85%)
Frame = +1
Query: 582 AVVTPILLPFILVFFALAYLVYRHQII 608
+VVT LLP+I+VF LAYLVYRHQII
Sbjct: 7 SVVTQFLLPYIIVFLGLAYLVYRHQII 87
>BQ148754 weakly similar to GP|15146264|gb At1g10080/T27I1_10 {Arabidopsis
thaliana}, partial (19%)
Length = 685
Score = 146 bits (369), Expect = 3e-35
Identities = 75/209 (35%), Positives = 123/209 (57%)
Frame = +1
Query: 506 SIPMKATFFMTYIMVDGWAGIAGEILRLKPLVIYHLKNMFIVKTERDRGKAMDPGSVDFP 565
++P + TFF TYI+ GWA + E+L++ PL +Y+L F+++ + D ++ + +
Sbjct: 10 AVPAQGTFFTTYILSSGWASLGFELLQVCPL-LYNLFQRFLLRVKDD---TLNGITFPYH 177
Query: 566 ETIPSLQLYFLLGIVYAVVTPILLPFILVFFALAYLVYRHQIINVYNQQYESAAAFWPHV 625
+P L L+ LG +++ P++LPF+L++F LAYLVYR+QIINVY +Y+S +WP
Sbjct: 178 TEVPRLLLFGFLGFTCSILAPLMLPFLLIYFFLAYLVYRNQIINVYITKYDSGGQYWPIA 357
Query: 626 HSRIIASLLISQLLMLGLLSTKKAAKSTPLLAILPILTFAFHKYCQHRFEPAFRKYPLEE 685
H+ + SLL +QL+ LG+ K++ S +A L I+T FH+YC+ RF P FR +
Sbjct: 358 HNATVCSLLFAQLIALGVFGLKRSTVSAGFIAPLLIVTILFHQYCRKRFLPVFRSNSAQI 537
Query: 686 AMAKDLLEKTTEPDLNIKAYLADAYLHPI 714
+ D ++ I L AY P+
Sbjct: 538 LIDLDKKDEQCGRMEEIYEQLRSAYKQPM 624
>TC80095 weakly similar to GP|12597787|gb|AAG60099.1 unknown protein
{Arabidopsis thaliana}, partial (18%)
Length = 826
Score = 127 bits (318), Expect = 2e-29
Identities = 74/232 (31%), Positives = 126/232 (53%), Gaps = 1/232 (0%)
Frame = +3
Query: 2 ATLADIGVSAAINILSAFAFLLAFALLRIQPINDRVYFPKWYINGERTSPRSTGNFVGKF 61
A L +G++ A+ +L F +++LR QP N +VY P+ G + +
Sbjct: 174 ALLTSVGINTALCVL----FFTLYSILRKQPSNYKVYIPRLLAEG-----------ISRK 308
Query: 62 VNLNFKTYLTFLNWMPQALRMSENEIINHAGLDSAVFLRIYTLGLKIFVPVTVVALLILI 121
K + +W+ +A +SE+E+ + +GLD+ VF+ + T LKIF V+ + +L+
Sbjct: 309 KPFKLKQLIPSPDWVAKAWNLSEDELFSSSGLDALVFMHLITFSLKIFTFAGVIGIFVLL 488
Query: 122 PVNVSSGALFFLKR-DLVVSDIDKLSISNVPSKSSRFFVHIGLEYMFTLWICFLLYKEYD 180
PVN+ L + D + +D +ISNV S S+ +VH YM + + CF L+ EY
Sbjct: 489 PVNLWGNQLEEVDMYDFAGNSLDVFTISNVNSGSNWLWVHFLAVYMVSGFTCFQLHHEYK 668
Query: 181 NIAMMRLHFLASQRRRVEQFTVVVRSVPHISGHSVSDSVESFFQTNHPNHYI 232
IA R+ + +S + QFT++V S+P S S+S+SV+SFF+ +P+ Y+
Sbjct: 669 YIASKRISYFSSSKPLPHQFTILVHSIPTSSSCSISESVDSFFRELYPSTYL 824
>TC84436 weakly similar to GP|498707|emb|CAA55187.1| HYP1 {Arabidopsis
thaliana}, partial (17%)
Length = 716
Score = 124 bits (310), Expect = 2e-28
Identities = 67/198 (33%), Positives = 108/198 (53%)
Frame = +3
Query: 155 SRFFVHIGLEYMFTLWICFLLYKEYDNIAMMRLHFLASQRRRVEQFTVVVRSVPHISGHS 214
SR ++H Y+FT +C+LLY EY I+ R+ S + FTV+VR +P G +
Sbjct: 84 SRLWIHFSAAYVFTGVVCYLLYYEYRYISSKRIACFYSSEPQPHHFTVLVRGIPIPPGST 263
Query: 215 VSDSVESFFQTNHPNHYIGHQAVYNANKFAKLVRRRDRLQNWLDYYQLKFQRHPDKRPTI 274
+D+V+ FF HP+ Y+ H V ++K L+ D+L L LK + KR T
Sbjct: 264 CTDAVQRFFSEYHPSTYLSHSVVRRSSKLHNLITDADKLYKKLT--NLKQKNDAPKRQT- 434
Query: 275 NTGVLGLWGRKVDAIEYYSHTIKELDKVMTLERQRIIKDPKCILPVAFLSFSSRWGASVC 334
G GL+G KVD +++Y + ++ + +E+ + +P AF+SF +R+GA++
Sbjct: 435 REGCCGLFGPKVDTVDHYERRLGNIEDNVRMEQSSLASKE---VPAAFVSFKTRFGAAIA 605
Query: 335 AQTQQSKNPTLWLTDWAP 352
Q+S NPT W+T+ AP
Sbjct: 606 LHIQESVNPTEWITEEAP 659
>AW686896 similar to GP|12597787|gb unknown protein {Arabidopsis thaliana},
partial (28%)
Length = 661
Score = 112 bits (281), Expect = 4e-25
Identities = 65/220 (29%), Positives = 118/220 (53%), Gaps = 2/220 (0%)
Frame = +1
Query: 387 YMIPIAFVQSLANLEGLERVAPFLRPVIELKFIKSFLQGFLPGLALKIFLYILPAVLMIM 446
++IP+ VQ L NL+ LE + PFL ++ +KF+ + G+LP L L + L ++P V+ +
Sbjct: 16 FLIPVVLVQGLTNLKQLEILFPFLASILSIKFVTQIITGYLPSLILXMSLKLVPPVMGFL 195
Query: 447 SKIEGHIAKSTLERKTAAKYYYFMLVNVFLGSIVTGTAFEQLHSFLHQSPTQIPRTIGVS 506
S I+G+I+ S +E + K +F + NVF ++ +G+ QL+ L +I + V+
Sbjct: 196 SSIQGYISHSDIEMSASKKVIWFTVWNVFFATVFSGSILHQLYVIL--DLREITSNLAVA 369
Query: 507 IPMKATFFMTYIMVDGWAGIAGEILRLKPLVIYHLKNMFIVKTERDRGKAMDPGSVDFPE 566
+P +A+FF+ Y+ GW + E+ ++ P + +K F + RG S+
Sbjct: 370 VPAQASFFIPYVATTGWTNVLSELFQILPFISSLIKRPF----TKQRG*I--*SSISCLP 531
Query: 567 TIPSLQLYFLLGIVYAVVTPI--LLPFILVFFALAYLVYR 604
S + + L +V + + + L PF+ V+ LAY +YR
Sbjct: 532 QXCSKESFSLDFLVLHISS*LR*LYPFLWVYLWLAYXIYR 651
>TC86721 similar to GP|9757997|dbj|BAB08419.1
gene_id:MFH8.10~pir||T02893~similar to unknown protein
{Arabidopsis thaliana}, partial (90%)
Length = 1362
Score = 79.3 bits (194), Expect = 5e-15
Identities = 34/67 (50%), Positives = 52/67 (76%)
Frame = +2
Query: 403 LERVAPFLRPVIELKFIKSFLQGFLPGLALKIFLYILPAVLMIMSKIEGHIAKSTLERKT 462
L++ AP+L P++ + + SF+QGFLPG+ LK+FL LP +LM+MSK EG + S+LER++
Sbjct: 1160 LQKAAPWLNPLVRVPVVMSFIQGFLPGIVLKLFLIFLPTILMMMSKFEGFGSISSLERRS 1339
Query: 463 AAKYYYF 469
A++YY F
Sbjct: 1340 ASRYYLF 1360
>TC79893 weakly similar to GP|12597787|gb|AAG60099.1 unknown protein
{Arabidopsis thaliana}, partial (15%)
Length = 867
Score = 75.1 bits (183), Expect = 1e-13
Identities = 35/85 (41%), Positives = 50/85 (58%)
Frame = +2
Query: 606 QIINVYNQQYESAAAFWPHVHSRIIASLLISQLLMLGLLSTKKAAKSTPLLAILPILTFA 665
Q INVY +YE+A FWP VH +I SL++ QL+ +G + KK + ++ LP+ T
Sbjct: 2 QFINVYTPKYETAGRFWPIVHDSMIFSLVLMQLIAVGSFALKKLSPASTWTLPLPVFTLL 181
Query: 666 FHKYCQHRFEPAFRKYPLEEAMAKD 690
F+ YC+ RF P F Y E + KD
Sbjct: 182 FNYYCRRRFLPIFTAYSAESLVKKD 256
>BF521545 similar to GP|498707|emb|C HYP1 {Arabidopsis thaliana}, partial
(13%)
Length = 213
Score = 63.2 bits (152), Expect = 4e-10
Identities = 23/64 (35%), Positives = 44/64 (67%)
Frame = +3
Query: 558 DPGSVDFPETIPSLQLYFLLGIVYAVVTPILLPFILVFFALAYLVYRHQIINVYNQQYES 617
+P S+ + IP ++L+ LLG+ Y + P++LPF+L++F L Y+++R+Q + VY ++E+
Sbjct: 21 EPPSIPYHSEIPRIRLFGLLGVTYFFLAPLILPFLLIYFCLGYIIFRNQFLKVYVPKFET 200
Query: 618 AAAF 621
F
Sbjct: 201 GGEF 212
>BF646663 similar to GP|6714477|gb|A hypothetical protein {Arabidopsis
thaliana}, partial (41%)
Length = 570
Score = 57.8 bits (138), Expect = 2e-08
Identities = 35/121 (28%), Positives = 63/121 (51%)
Frame = +3
Query: 4 LADIGVSAAINILSAFAFLLAFALLRIQPINDRVYFPKWYINGERTSPRSTGNFVGKFVN 63
L+ + S AIN+ F +++LR QP N VY P++ G+ + G F
Sbjct: 231 LSALLTSVAINLGLCLLFFTLYSILRKQPGNINVYVPRFVAEGK---VKEGGQF------ 383
Query: 64 LNFKTYLTFLNWMPQALRMSENEIINHAGLDSAVFLRIYTLGLKIFVPVTVVALLILIPV 123
N + L W+ +A + +E ++ +GLD+ VF+R++ LK+F ++ ++LIP+
Sbjct: 384 -NLERLLPTAGWVRKAWEPTXDEFLSTSGLDAFVFMRMFXFSLKVFTFGAIIG-IVLIPI 557
Query: 124 N 124
N
Sbjct: 558 N 560
>AJ388960 weakly similar to GP|14596049|gb| Unknown protein {Arabidopsis
thaliana}, partial (14%)
Length = 477
Score = 43.9 bits (102), Expect = 2e-04
Identities = 24/80 (30%), Positives = 42/80 (52%), Gaps = 1/80 (1%)
Frame = +2
Query: 153 KSSRFFVHIGLEYMFTLWICFLLYKEYDNIAMMRLHFLASQRRRVEQFTVVVRSVPHI-S 211
KS R + Y +L FLL+K Y +++ +R S + EQF +VVR +P +
Sbjct: 2 KSVRLWAFFIACYFVSLVSLFLLWKAYKHVSWLRTEAFKSIDVKPEQFAIVVRDIPPVLD 181
Query: 212 GHSVSDSVESFFQTNHPNHY 231
G + + V+S+F+ +P +
Sbjct: 182 GQTRKEQVDSYFKAIYPEEH 241
>TC76431 similar to GP|8132441|gb|AAF73291.1| extensin {Pisum sativum},
partial (30%)
Length = 890
Score = 43.5 bits (101), Expect = 3e-04
Identities = 22/60 (36%), Positives = 26/60 (42%), Gaps = 9/60 (15%)
Frame = +1
Query: 745 PSSPSPLHDHVHQPSPPQHVN---QYSHYPTSPPGY------YYHPPPPPHYAYQYQDEP 795
PS P P H H P P H +Y + PP + YYH PPPP Y+Y P
Sbjct: 136 PSPPPPTHHHYPHPHPVYHSPPPPKYKYSSPPPPVHHVSPKPYYHSPPPPKKPYKYSSPP 315
Score = 37.0 bits (84), Expect = 0.030
Identities = 24/81 (29%), Positives = 31/81 (37%), Gaps = 25/81 (30%)
Frame = +1
Query: 740 PTLSEPSSPSPLHD-----HVHQPSPPQHVNQYS------HYPTSPPGYY---------- 778
P S P P+H + H P PP+ +YS H P +P Y
Sbjct: 205 PKYKYSSPPPPVHHVSPKPYYHSPPPPKKPYKYSSPPPPVHPPYTPKPVYHSPPVHPPYT 384
Query: 779 ----YHPPPPPHYAYQYQDEP 795
YH PPPP ++Y Y P
Sbjct: 385 PHPVYHSPPPPVHSYIYASPP 447
Score = 35.0 bits (79), Expect = 0.11
Identities = 18/52 (34%), Positives = 24/52 (45%), Gaps = 6/52 (11%)
Frame = +1
Query: 746 SSPSPLHDHVH-QPSPPQHVNQYSHYPTSPPG-----YYYHPPPPPHYAYQY 791
S P P H + H P P H + Y + + PP Y PPPP H+ Y +
Sbjct: 19 SPPPPYHPYPHPHPHPHPHPHPYPVHHSPPPSPKKPYKYPSPPPPTHHHYPH 174
Score = 34.7 bits (78), Expect = 0.15
Identities = 18/56 (32%), Positives = 21/56 (37%), Gaps = 8/56 (14%)
Frame = +1
Query: 748 PSPLHDHVHQPSPPQHVNQYSHYPTSPPGYY--------YHPPPPPHYAYQYQDEP 795
P P VH PP Y + PP ++ YH PPPP Y Y P
Sbjct: 70 PHPHPYPVHHSPPPSPKKPYKYPSPPPPTHHHYPHPHPVYHSPPPPKYKYSSPPPP 237
Score = 32.7 bits (73), Expect = 0.56
Identities = 17/56 (30%), Positives = 25/56 (44%)
Frame = +1
Query: 733 PETQVASPTLSEPSSPSPLHDHVHQPSPPQHVNQYSHYPTSPPGYYYHPPPPPHYA 788
P+ SP + P +P P++ H P PP H Y Y PPPP+++
Sbjct: 340 PKPVYHSPPVHPPYTPHPVY---HSPPPPVH------------SYIYASPPPPYHS 462
Score = 30.0 bits (66), Expect = 3.6
Identities = 15/48 (31%), Positives = 21/48 (43%)
Frame = +1
Query: 740 PTLSEPSSPSPLHDHVHQPSPPQHVNQYSHYPTSPPGYYYHPPPPPHY 787
P + P +P P++ H PP + H P P Y + PPP Y
Sbjct: 316 PPVHPPYTPKPVY-HSPPVHPPYTPHPVYHSPPPPVHSYIYASPPPPY 456
>AW685919 homologue to PIR|S25298|S252 extensin (clone Tom J-10) - tomato,
partial (23%)
Length = 663
Score = 43.1 bits (100), Expect = 4e-04
Identities = 20/51 (39%), Positives = 25/51 (48%)
Frame = +1
Query: 745 PSSPSPLHDHVHQPSPPQHVNQYSHYPTSPPGYYYHPPPPPHYAYQYQDEP 795
P SPSP + ++ PP P+ PP YYY PPPP Y+Y P
Sbjct: 127 PPSPSPPPPYYYKSPPPPS-------PSPPPPYYYKSPPPPAPVYKYNSPP 258
Score = 39.7 bits (91), Expect = 0.005
Identities = 23/55 (41%), Positives = 27/55 (48%)
Frame = +1
Query: 733 PETQVASPTLSEPSSPSPLHDHVHQPSPPQHVNQYSHYPTSPPGYYYHPPPPPHY 787
P SP PS P P + P PP V +Y+ P PP +YY PPPPHY
Sbjct: 148 PPYYYKSPPPPSPSPPPPYY--YKSPPPPAPVYKYNSPP--PPVHYY-SPPPPHY 297
Score = 37.7 bits (86), Expect = 0.017
Identities = 20/55 (36%), Positives = 26/55 (46%), Gaps = 4/55 (7%)
Frame = +1
Query: 745 PSSPSPLHDHVHQPSPPQHVNQYSHYPTSPPGYYYHPPPPPHYA----YQYQDEP 795
P SPSP + ++ PP P+ PP YYY PPPP + Y Y+ P
Sbjct: 79 PPSPSPPPPYYYKSPPPPS-------PSPPPPYYYKSPPPPSPSPPPPYYYKSPP 222
Score = 37.7 bits (86), Expect = 0.017
Identities = 20/55 (36%), Positives = 26/55 (46%), Gaps = 4/55 (7%)
Frame = +1
Query: 745 PSSPSPLHDHVHQPSPPQHVNQYSHYPTSPPGYYYHPPPPPHYA----YQYQDEP 795
P SPSP + ++ PP P+ PP YYY PPPP + Y Y+ P
Sbjct: 31 PPSPSPPPPYYYKSPPPPS-------PSPPPPYYYKSPPPPSPSPPPPYYYKSPP 174
>BF642399 similar to PIR|T10863|T10 extensin precursor - kidney bean, partial
(33%)
Length = 653
Score = 42.4 bits (98), Expect = 7e-04
Identities = 23/66 (34%), Positives = 27/66 (40%), Gaps = 13/66 (19%)
Frame = +3
Query: 743 SEPSSPSPLHDH-----VHQPSPPQHVNQ--------YSHYPTSPPGYYYHPPPPPHYAY 789
S P P P + H VH P PP H + Y + PP Y Y PPPP Y Y
Sbjct: 105 SPPPPPKPYYYHSPPPPVHSPPPPYHYSSPPPPPKKPYKYASPPPPVYKYKSPPPPVYKY 284
Query: 790 QYQDEP 795
+ P
Sbjct: 285 KSPPPP 302
Score = 40.8 bits (94), Expect = 0.002
Identities = 20/56 (35%), Positives = 26/56 (45%)
Frame = +3
Query: 740 PTLSEPSSPSPLHDHVHQPSPPQHVNQYSHYPTSPPGYYYHPPPPPHYAYQYQDEP 795
P S P P++ + P PP+ +Y P PP Y Y PPPP Y Y+ P
Sbjct: 240 PVYKYKSPPPPVYKYKSPPPPPKKPYKYPSPP--PPVYKYKSPPPPVYKYKSPPPP 401
Score = 40.0 bits (92), Expect = 0.003
Identities = 21/61 (34%), Positives = 29/61 (47%), Gaps = 5/61 (8%)
Frame = +3
Query: 740 PTLSEPSSPSPLHDHVHQPSPPQHVNQYSHYPTSPPGYYYHPPPPPHYA-----YQYQDE 794
P S P P++ + P PP+ +Y P PP Y Y PPPP Y+ Y+Y+
Sbjct: 339 PVYKYKSPPPPVYKYKSPPPPPKKPYKYPSPP--PPVYKYKSPPPPVYSPPPPVYKYKSP 512
Query: 795 P 795
P
Sbjct: 513 P 515
Score = 40.0 bits (92), Expect = 0.003
Identities = 23/57 (40%), Positives = 25/57 (43%), Gaps = 1/57 (1%)
Frame = +3
Query: 740 PTLSEPSSPSPLHDHVHQPSPPQHVNQYSHYPTSPPGY-YYHPPPPPHYAYQYQDEP 795
P S P P PSPP V +Y P PP Y Y PPPPP Y+Y P
Sbjct: 270 PVYKYKSPPPPPKKPYKYPSPPPPVYKYKSPP--PPVYKYKSPPPPPKKPYKYPSPP 434
Score = 37.0 bits (84), Expect = 0.030
Identities = 23/64 (35%), Positives = 25/64 (38%), Gaps = 1/64 (1%)
Frame = +3
Query: 733 PETQVASPTLSEPSSPSPLHDHVHQPSPPQHVNQYSHYPTSPPGY-YYHPPPPPHYAYQY 791
P P S P P SPP V +Y P PP Y Y PPPPP Y+Y
Sbjct: 150 PPVHSPPPPYHYSSPPPPPKKPYKYASPPPPVYKYKSPP--PPVYKYKSPPPPPKKPYKY 323
Query: 792 QDEP 795
P
Sbjct: 324 PSPP 335
Score = 36.2 bits (82), Expect = 0.050
Identities = 20/52 (38%), Positives = 26/52 (49%), Gaps = 1/52 (1%)
Frame = +3
Query: 745 PSSPSPLHDHVHQPSPPQHVNQYSHYPTSPPGYYYHP-PPPPHYAYQYQDEP 795
PS P P++ + SPP V +Y P P Y +P PPPP Y Y+ P
Sbjct: 324 PSPPPPVYKY---KSPPPPVYKYKSPPPPPKKPYKYPSPPPPVYKYKSPPPP 470
Score = 35.0 bits (79), Expect = 0.11
Identities = 22/65 (33%), Positives = 30/65 (45%), Gaps = 2/65 (3%)
Frame = +3
Query: 733 PETQVASPTLSEPSSPSPLHDHVHQPSPPQHVNQYSH-YPTSPPGYYYHP-PPPPHYAYQ 790
P + SP S P P++ + P PP H + Y + PP Y +P PPPP Y Y+
Sbjct: 438 PVYKYKSPPPPVYSPPPPVYKY-KSPPPPVHSPPXPYKYKSPPPXPYKYPSPPPPVYKYK 614
Query: 791 YQDEP 795
P
Sbjct: 615 SPPSP 629
Score = 34.3 bits (77), Expect = 0.19
Identities = 15/33 (45%), Positives = 17/33 (51%), Gaps = 4/33 (12%)
Frame = +3
Query: 767 YSHYPTSPPGYYYHPPPPPHYA----YQYQDEP 795
YS P P YYYH PPPP ++ Y Y P
Sbjct: 99 YSSPPPPPKPYYYHSPPPPVHSPPPPYHYSSPP 197
Score = 31.6 bits (70), Expect = 1.2
Identities = 23/66 (34%), Positives = 26/66 (38%), Gaps = 6/66 (9%)
Frame = +3
Query: 731 KQPETQVASP---TLSEPSSPSPLHDH---VHQPSPPQHVNQYSHYPTSPPGYYYHPPPP 784
K P V SP S P P+H SPP +Y P PP Y Y PP
Sbjct: 453 KSPPPPVYSPPPPVYKYKSPPPPVHSPPXPYKYKSPPPXPYKYPSPP--PPVYKYKSPPS 626
Query: 785 PHYAYQ 790
P Y Y+
Sbjct: 627 PVYKYK 644
Score = 30.0 bits (66), Expect = 3.6
Identities = 19/52 (36%), Positives = 25/52 (47%), Gaps = 2/52 (3%)
Frame = +3
Query: 746 SSPSPLHDHVHQPSPPQHVNQYSHYPTSP--PGYYYHPPPPPHYAYQYQDEP 795
S P P++ + SPP V +Y P P P Y PPPP Y+Y+ P
Sbjct: 228 SPPPPVYKY---KSPPPPVYKYKSPPPPPKKPYKYPSPPPP---VYKYKSPP 365
>TC76718 homologue to GP|8132441|gb|AAF73291.1| extensin {Pisum sativum},
partial (63%)
Length = 712
Score = 42.4 bits (98), Expect = 7e-04
Identities = 23/66 (34%), Positives = 27/66 (40%), Gaps = 13/66 (19%)
Frame = +1
Query: 743 SEPSSPSPLHDH-----VHQPSPPQHVNQ--------YSHYPTSPPGYYYHPPPPPHYAY 789
S P P P + H VH P PP H + Y + PP Y Y PPPP Y Y
Sbjct: 130 SPPPPPKPYYYHSPPPPVHSPPPPYHYSSPPPPPKKPYKYASPPPPVYKYKSPPPPVYKY 309
Query: 790 QYQDEP 795
+ P
Sbjct: 310 KSPPPP 327
Score = 40.8 bits (94), Expect = 0.002
Identities = 20/56 (35%), Positives = 26/56 (45%)
Frame = +1
Query: 740 PTLSEPSSPSPLHDHVHQPSPPQHVNQYSHYPTSPPGYYYHPPPPPHYAYQYQDEP 795
P S P P++ + P PP+ +Y P PP Y Y PPPP Y Y+ P
Sbjct: 265 PVYKYKSPPPPVYKYKSPPPPPKKPYKYPSPP--PPVYKYKSPPPPVYKYKSPPPP 426
Score = 40.4 bits (93), Expect = 0.003
Identities = 22/56 (39%), Positives = 24/56 (42%)
Frame = +1
Query: 740 PTLSEPSSPSPLHDHVHQPSPPQHVNQYSHYPTSPPGYYYHPPPPPHYAYQYQDEP 795
P S P P PSPP V +Y P PP Y Y PPPP Y Y+ P
Sbjct: 295 PVYKYKSPPPPPKKPYKYPSPPPPVYKYKSPP--PPVYKYKSPPPPVYKYKSPPPP 456
Score = 38.9 bits (89), Expect = 0.008
Identities = 22/58 (37%), Positives = 27/58 (45%), Gaps = 7/58 (12%)
Frame = +1
Query: 745 PSSPSPLHDHVHQP-------SPPQHVNQYSHYPTSPPGYYYHPPPPPHYAYQYQDEP 795
PS P P++ + P SPP V +Y P PP Y Y PPPP Y Y+ P
Sbjct: 349 PSPPPPVYKYKSPPPPVYKYKSPPPPVYKYKSPP--PPVYKYKSPPPPVYKYKSPPPP 516
Score = 37.0 bits (84), Expect = 0.030
Identities = 22/57 (38%), Positives = 27/57 (46%), Gaps = 1/57 (1%)
Frame = +1
Query: 740 PTLSEPSSPSPLHDHVHQPSPPQHVNQYSHYPTSPPGYYYH-PPPPPHYAYQYQDEP 795
P S P P++ + SPP V +Y P PP Y Y PPPPP Y+Y P
Sbjct: 394 PVYKYKSPPPPVYKY---KSPPPPVYKYKSPP--PPVYKYKSPPPPPKKPYKYPSPP 549
Score = 37.0 bits (84), Expect = 0.030
Identities = 23/64 (35%), Positives = 25/64 (38%), Gaps = 1/64 (1%)
Frame = +1
Query: 733 PETQVASPTLSEPSSPSPLHDHVHQPSPPQHVNQYSHYPTSPPGY-YYHPPPPPHYAYQY 791
P P S P P SPP V +Y P PP Y Y PPPPP Y+Y
Sbjct: 175 PPVHSPPPPYHYSSPPPPPKKPYKYASPPPPVYKYKSPP--PPVYKYKSPPPPPKKPYKY 348
Query: 792 QDEP 795
P
Sbjct: 349 PSPP 360
Score = 36.6 bits (83), Expect = 0.039
Identities = 18/51 (35%), Positives = 24/51 (46%)
Frame = +1
Query: 740 PTLSEPSSPSPLHDHVHQPSPPQHVNQYSHYPTSPPGYYYHPPPPPHYAYQ 790
P S P P++ + P PP+ +Y P PP Y Y P PP Y Y+
Sbjct: 454 PVYKYKSPPPPVYKYKSPPPPPKKPYKYPSPP--PPVYKYKSPXPPVYKYK 600
Score = 34.7 bits (78), Expect = 0.15
Identities = 20/57 (35%), Positives = 26/57 (45%), Gaps = 1/57 (1%)
Frame = +1
Query: 740 PTLSEPSSPSPLHDHVHQPSPPQHVNQYSHYPTSPPGYYYHP-PPPPHYAYQYQDEP 795
P S P P++ + SPP V +Y P P Y +P PPPP Y Y+ P
Sbjct: 424 PVYKYKSPPPPVYKY---KSPPPPVYKYKSPPPPPKKPYKYPSPPPPVYKYKSPXPP 585
Score = 34.3 bits (77), Expect = 0.19
Identities = 15/33 (45%), Positives = 17/33 (51%), Gaps = 4/33 (12%)
Frame = +1
Query: 767 YSHYPTSPPGYYYHPPPPPHYA----YQYQDEP 795
YS P P YYYH PPPP ++ Y Y P
Sbjct: 124 YSSPPPPPKPYYYHSPPPPVHSPPPPYHYSSPP 222
Score = 30.0 bits (66), Expect = 3.6
Identities = 19/52 (36%), Positives = 25/52 (47%), Gaps = 2/52 (3%)
Frame = +1
Query: 746 SSPSPLHDHVHQPSPPQHVNQYSHYPTSP--PGYYYHPPPPPHYAYQYQDEP 795
S P P++ + SPP V +Y P P P Y PPPP Y+Y+ P
Sbjct: 253 SPPPPVYKY---KSPPPPVYKYKSPPPPPKKPYKYPSPPPP---VYKYKSPP 390
Score = 29.6 bits (65), Expect = 4.7
Identities = 18/65 (27%), Positives = 27/65 (40%), Gaps = 8/65 (12%)
Frame = +2
Query: 739 SPTLSEPSSPSPLHDHVHQPSPPQHVNQYSHYPTSPPGYYYH--------PPPPPHYAYQ 790
+P L + +S S LH H+ H+N + H+ + H PPPP Y+
Sbjct: 470 NPLLHQCTSTSLLHHHLRS-----HINTHLHHRQFTSTNHLHLQFTSTNLHPPPPKKPYK 634
Query: 791 YQDEP 795
Y P
Sbjct: 635 YPSPP 649
Score = 29.3 bits (64), Expect = 6.2
Identities = 19/57 (33%), Positives = 25/57 (43%), Gaps = 11/57 (19%)
Frame = +1
Query: 745 PSSPSPLHDHVHQPSPPQHVNQYSHYPTSPPG-----------YYYHPPPPPHYAYQ 790
PS P P++ + P PP V +Y PT+ Y Y PPPP Y Y+
Sbjct: 538 PSPPPPVYKY-KSPXPP--VYKYKSPPTTTQETIQVPITTTTRYKYKSPPPPVYKYK 699
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.326 0.140 0.426
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 26,663,175
Number of Sequences: 36976
Number of extensions: 465074
Number of successful extensions: 7919
Number of sequences better than 10.0: 217
Number of HSP's better than 10.0 without gapping: 3778
Number of HSP's successfully gapped in prelim test: 223
Number of HSP's that attempted gapping in prelim test: 977
Number of HSP's gapped (non-prelim): 5266
length of query: 795
length of database: 9,014,727
effective HSP length: 104
effective length of query: 691
effective length of database: 5,169,223
effective search space: 3571933093
effective search space used: 3571933093
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 62 (28.5 bits)
Lotus: description of TM0020.6


