

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0019a.4
(310 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
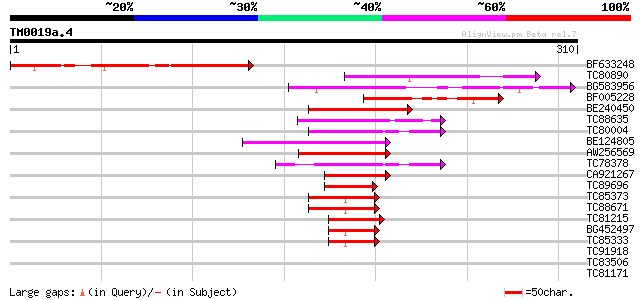
BF633248 similar to SP|P45598|ARAE_ Arabinose-proton symporter (... 105 2e-23
TC80890 similar to PIR|T04270|T04270 hypothetical protein F20B18... 84 5e-17
BG583956 homologue to GP|10177159|dbj gb|AAF63824.1~gene_id:K21P... 80 8e-16
BF005228 similar to GP|17473547|gb| unknown protein {Arabidopsis... 74 5e-14
BE240450 similar to GP|21555178|gb| transcription factor-like pr... 72 2e-13
TC88635 similar to GP|12711287|emb|CAC28528. GATA-1 zinc finger ... 62 4e-10
TC80004 similar to GP|8778844|gb|AAF79843.1| T6D22.9 {Arabidopsi... 58 4e-09
BE124805 similar to GP|10177426|db GATA-binding transcription fa... 58 5e-09
AW256569 similar to GP|10177426|dbj GATA-binding transcription f... 57 7e-09
TC78378 similar to GP|10177426|dbj|BAB10711. GATA-binding transc... 54 1e-07
CA921267 homologue to GP|10177426|dbj GATA-binding transcription... 53 2e-07
TC89696 similar to GP|10177426|dbj|BAB10711. GATA-binding transc... 52 4e-07
TC85373 similar to GP|15028099|gb|AAK76580.1 putative flowering ... 48 4e-06
TC88671 similar to PIR|JC7336|JC7336 zinc-finger protein - Arabi... 47 1e-05
TC81215 similar to GP|17064972|gb|AAL32640.1 Unknown protein {Ar... 44 8e-05
BG452497 similar to GP|15028099|gb| putative flowering protein C... 43 2e-04
TC85333 similar to GP|15028099|gb|AAK76580.1 putative flowering ... 43 2e-04
TC91918 homologue to GP|9369375|gb|AAF87124.1| F10A5.29 {Arabido... 35 0.028
TC83506 similar to PIR|T08179|T08179 LRG5 protein - Chlamydomona... 34 0.082
TC81171 similar to GP|9795609|gb|AAF98427.1| Unknown protein {Ar... 32 0.41
>BF633248 similar to SP|P45598|ARAE_ Arabinose-proton symporter (Arabinose
transporter). {Klebsiella oxytoca}, partial (5%)
Length = 514
Score = 105 bits (263), Expect = 2e-23
Identities = 73/141 (51%), Positives = 88/141 (61%), Gaps = 8/141 (5%)
Frame = +1
Query: 1 MTPVSLNPPGPS--IQGQNHLFN-SLNNQDYHASLFNILDRRQGIGIGELREN----DHQ 53
MTPVSLNPPGP+ +QGQN FN S NQD + FN+L G+ EN HQ
Sbjct: 94 MTPVSLNPPGPNSLLQGQNQFFNISPVNQDT-PTFFNLL--------GDFGENYDHHHHQ 246
Query: 54 DDKLVVWHDGSSSSSSSNHLYNSTFISPQPVMDNPSSSTCDPNLSFS-KMEEEDIKNVHG 112
D KL HDGSSSS+ LYNS S + VM + SSS D NLS S ++E+ KN HG
Sbjct: 247 DHKLAFHHDGSSSSNHQQQLYNS---SSESVMVD-SSSARDTNLSSSLELEDSSKKNSHG 414
Query: 113 SAKWMSSKMRLMKKMMSTPAT 133
S KW+SSKMR+M KM++T AT
Sbjct: 415 SEKWISSKMRVMNKMINTTAT 477
>TC80890 similar to PIR|T04270|T04270 hypothetical protein F20B18.260 -
Arabidopsis thaliana, partial (14%)
Length = 710
Score = 84.3 bits (207), Expect = 5e-17
Identities = 48/111 (43%), Positives = 60/111 (53%), Gaps = 4/111 (3%)
Frame = +1
Query: 184 PLWRSGPNGPKSLCNACGIRQRKARRAMAEAANG----LATPINTASTKTRVHHNKEKKP 239
PLWRSGP GPKSLCNACGIRQRKARRA+A AANG +A K ++ + K
Sbjct: 7 PLWRSGPTGPKSLCNACGIRQRKARRALAAAANGETLVVAEKPYVKGKKLQIKRKRSKTD 186
Query: 240 RANHFAQFKNKSKSTSSNAGSSQETKKLECFKDFAISLRNNSTFQQVFPRD 290
+ + K KS++ +N F+D S NN QVFP+D
Sbjct: 187 QCAQLLKRKGKSENKCNN------------FEDLITSWSNNLASHQVFPQD 303
>BG583956 homologue to GP|10177159|dbj gb|AAF63824.1~gene_id:K21P3.18~similar
to unknown protein {Arabidopsis thaliana}, partial (43%)
Length = 818
Score = 80.5 bits (197), Expect = 8e-16
Identities = 57/167 (34%), Positives = 81/167 (48%), Gaps = 10/167 (5%)
Frame = +2
Query: 153 HENRYSQRSPRNNN-----NSSNTTRVCSDCNTSSTPLWRSGPNGPKSLCNACGIRQRKA 207
H ++ S+ N N ++SN + C+DC TS TPLWR GP GPKSLCNACGIR RK
Sbjct: 188 HSDKVSEGEDSNPNAAVSSDNSNPKKTCADCGTSKTPLWRGGPAGPKSLCNACGIRSRKK 367
Query: 208 RRAMAEAANGLATPINTASTKTRVHHNKEKKPRANHFAQFKNKSKSTSSNAGSSQETKKL 267
+RA+ + G N E+ R K K ++ G S+ L
Sbjct: 368 KRAILGISKG----------------NNEEGTR-------KGKKSNSGGGGGGSKVGDNL 478
Query: 268 ECFKDFAISL-----RNNSTFQQVFPRDEVAEAALLLMDLSCGYVHS 309
K ++L N S ++++ E +AA+LLM LS G V++
Sbjct: 479 N-MKQRLLNLGKEVFMNRSHWKKL---GEDEQAAVLLMSLSYGSVYA 607
>BF005228 similar to GP|17473547|gb| unknown protein {Arabidopsis thaliana},
partial (5%)
Length = 742
Score = 74.3 bits (181), Expect = 5e-14
Identities = 50/85 (58%), Positives = 55/85 (63%), Gaps = 8/85 (9%)
Frame = +2
Query: 194 KSLCNACGIRQRKARRAMAEAANGLATPINTASTKTRVHHNKEKKPRANHFAQFKNKSK- 252
+SLCNACGIRQRKARRAMAEAANGLAT S KT+V K KKP QFK K+K
Sbjct: 503 QSLCNACGIRQRKARRAMAEAANGLAT-----SPKTKV--LKIKKP-----TQFKTKNKA 646
Query: 253 -------STSSNAGSSQETKKLECF 270
ST+S SSQ+ KKLE F
Sbjct: 647 STSTSSTSTTSAGSSSQDVKKLESF 721
>BE240450 similar to GP|21555178|gb| transcription factor-like protein
{Arabidopsis thaliana}, partial (18%)
Length = 529
Score = 72.4 bits (176), Expect = 2e-13
Identities = 34/58 (58%), Positives = 42/58 (71%), Gaps = 1/58 (1%)
Frame = +1
Query: 164 NNNNSSNTTRVCSDCNTSSTPLWRSGPNGPKSLCNACGIRQRK-ARRAMAEAANGLAT 220
N+NN S R C+ C+++STPLWR+GP GP+SLCNACGIR +K RRA A A AT
Sbjct: 1 NSNNDSLLARRCASCDSTSTPLWRNGPRGPESLCNACGIRYKKEERRANAAVATTAAT 174
>TC88635 similar to GP|12711287|emb|CAC28528. GATA-1 zinc finger protein
{Nicotiana tabacum}, partial (21%)
Length = 1327
Score = 61.6 bits (148), Expect = 4e-10
Identities = 33/81 (40%), Positives = 43/81 (52%)
Frame = +1
Query: 158 SQRSPRNNNNSSNTTRVCSDCNTSSTPLWRSGPNGPKSLCNACGIRQRKARRAMAEAANG 217
S + R++ + S R C+ C + TP WR GP GPK+LCNACG+R R R +
Sbjct: 775 SIETKRSSLHESIAPRKCTHCEVTETPQWREGPKGPKTLCNACGVRYRSGR--LFPEYRP 948
Query: 218 LATPINTASTKTRVHHNKEKK 238
A+P AS VH N KK
Sbjct: 949 AASPTFEAS----VHSNSHKK 999
>TC80004 similar to GP|8778844|gb|AAF79843.1| T6D22.9 {Arabidopsis
thaliana}, partial (8%)
Length = 1058
Score = 58.2 bits (139), Expect = 4e-09
Identities = 30/75 (40%), Positives = 42/75 (56%)
Frame = +3
Query: 164 NNNNSSNTTRVCSDCNTSSTPLWRSGPNGPKSLCNACGIRQRKARRAMAEAANGLATPIN 223
NN + TR C+ C + TP WR+GP GPK+LCNACG+R K+ R + E P
Sbjct: 636 NNGQNPIPTRRCTHCLSQRTPQWRAGPLGPKTLCNACGVRY-KSGRLLPE-----YRPAK 797
Query: 224 TASTKTRVHHNKEKK 238
+ + + +H N KK
Sbjct: 798 SPTFVSFLHSNSHKK 842
>BE124805 similar to GP|10177426|db GATA-binding transcription factor-like
protein {Arabidopsis thaliana}, partial (36%)
Length = 715
Score = 57.8 bits (138), Expect = 5e-09
Identities = 26/81 (32%), Positives = 41/81 (50%)
Frame = +2
Query: 128 MSTPATDKANNSTTIPISPRIQNQGHENRYSQRSPRNNNNSSNTTRVCSDCNTSSTPLWR 187
+S + ++S T+ S + + ++R + + R CS C TP WR
Sbjct: 287 ISLANSSSTSSSATLSSSNLEECSKPAEKKAKRMVSPDGEARGVPRRCSHCGVQKTPQWR 466
Query: 188 SGPNGPKSLCNACGIRQRKAR 208
+GP GPK+LCNACG+R + R
Sbjct: 467 TGPGGPKTLCNACGVRYKSGR 529
>AW256569 similar to GP|10177426|dbj GATA-binding transcription factor-like
protein {Arabidopsis thaliana}, partial (20%)
Length = 613
Score = 57.4 bits (137), Expect = 7e-09
Identities = 22/50 (44%), Positives = 31/50 (62%)
Frame = -3
Query: 159 QRSPRNNNNSSNTTRVCSDCNTSSTPLWRSGPNGPKSLCNACGIRQRKAR 208
++ P ++ R CS C+ TP WR+GP GPK+LCNACG+R + R
Sbjct: 551 RKKPEAQTGGAHFQRRCSHCHVQKTPQWRAGPLGPKTLCNACGVRFKSGR 402
>TC78378 similar to GP|10177426|dbj|BAB10711. GATA-binding transcription
factor-like protein {Arabidopsis thaliana}, partial
(23%)
Length = 1284
Score = 53.5 bits (127), Expect = 1e-07
Identities = 31/93 (33%), Positives = 42/93 (44%)
Frame = +3
Query: 146 PRIQNQGHENRYSQRSPRNNNNSSNTTRVCSDCNTSSTPLWRSGPNGPKSLCNACGIRQR 205
P Q GHE + R CS C TP WR+GP G K+LCNACG+R
Sbjct: 762 PEAQVVGHEAQ----------EEGQLQRRCSHCQVQKTPQWRTGPMGAKTLCNACGVRY- 908
Query: 206 KARRAMAEAANGLATPINTASTKTRVHHNKEKK 238
K+ R +E P + + + +H N +K
Sbjct: 909 KSGRLFSE-----YRPACSPTFSSEIHSNSHRK 992
>CA921267 homologue to GP|10177426|dbj GATA-binding transcription factor-like
protein {Arabidopsis thaliana}, partial (20%)
Length = 694
Score = 52.8 bits (125), Expect = 2e-07
Identities = 21/36 (58%), Positives = 25/36 (69%)
Frame = -1
Query: 173 RVCSDCNTSSTPLWRSGPNGPKSLCNACGIRQRKAR 208
R CS C + TP WRSGP G K+LCNACG+R + R
Sbjct: 538 RRCSHCGVTKTPQWRSGPLGAKTLCNACGVRFKSGR 431
>TC89696 similar to GP|10177426|dbj|BAB10711. GATA-binding transcription
factor-like protein {Arabidopsis thaliana}, partial
(25%)
Length = 806
Score = 51.6 bits (122), Expect = 4e-07
Identities = 19/29 (65%), Positives = 21/29 (71%)
Frame = +2
Query: 173 RVCSDCNTSSTPLWRSGPNGPKSLCNACG 201
R C C TP WR+GPNGPK+LCNACG
Sbjct: 572 RKCHHCGVDDTPQWRAGPNGPKTLCNACG 658
>TC85373 similar to GP|15028099|gb|AAK76580.1 putative flowering protein
CONSTANS {Arabidopsis thaliana}, partial (22%)
Length = 683
Score = 48.1 bits (113), Expect = 4e-06
Identities = 20/41 (48%), Positives = 26/41 (62%), Gaps = 2/41 (4%)
Frame = +1
Query: 164 NNNNSSNTTRVCSDCNTSS--TPLWRSGPNGPKSLCNACGI 202
+N+ S VC CN S TP+ R GP GP++LCNACG+
Sbjct: 217 DNDGSQQQDNVCRQCNISEKCTPMMRRGPEGPRTLCNACGL 339
>TC88671 similar to PIR|JC7336|JC7336 zinc-finger protein - Arabidopsis
thaliana, partial (44%)
Length = 1272
Score = 46.6 bits (109), Expect = 1e-05
Identities = 20/41 (48%), Positives = 30/41 (72%), Gaps = 2/41 (4%)
Frame = +3
Query: 164 NNNNSSNTTRVCSDCNTSS--TPLWRSGPNGPKSLCNACGI 202
++ ++S + C+ C TSS TP+ R GP+GP+SLCNACG+
Sbjct: 693 SSQDASPSEISCTHCGTSSKSTPMMRRGPSGPRSLCNACGL 815
>TC81215 similar to GP|17064972|gb|AAL32640.1 Unknown protein {Arabidopsis
thaliana}, partial (14%)
Length = 938
Score = 43.9 bits (102), Expect = 8e-05
Identities = 18/31 (58%), Positives = 20/31 (64%)
Frame = +3
Query: 175 CSDCNTSSTPLWRSGPNGPKSLCNACGIRQR 205
C C +STPLWR+GP LCNACG R R
Sbjct: 447 CFHCGVTSTPLWRNGPPEKPILCNACGSRWR 539
>BG452497 similar to GP|15028099|gb| putative flowering protein CONSTANS
{Arabidopsis thaliana}, partial (12%)
Length = 648
Score = 42.7 bits (99), Expect = 2e-04
Identities = 17/30 (56%), Positives = 21/30 (69%), Gaps = 2/30 (6%)
Frame = +1
Query: 175 CSDCNTSS--TPLWRSGPNGPKSLCNACGI 202
C CN S TP+ R GP GP++LCNACG+
Sbjct: 196 CRQCNISEKCTPMMRRGPEGPRTLCNACGL 285
>TC85333 similar to GP|15028099|gb|AAK76580.1 putative flowering protein
CONSTANS {Arabidopsis thaliana}, partial (22%)
Length = 1421
Score = 42.7 bits (99), Expect = 2e-04
Identities = 17/30 (56%), Positives = 21/30 (69%), Gaps = 2/30 (6%)
Frame = +1
Query: 175 CSDCNTSS--TPLWRSGPNGPKSLCNACGI 202
C CN S TP+ R GP GP++LCNACG+
Sbjct: 907 CRQCNISEKCTPMMRRGPEGPRTLCNACGL 996
>TC91918 homologue to GP|9369375|gb|AAF87124.1| F10A5.29 {Arabidopsis
thaliana}, partial (38%)
Length = 1171
Score = 35.4 bits (80), Expect = 0.028
Identities = 22/85 (25%), Positives = 41/85 (47%)
Frame = +2
Query: 98 SFSKMEEEDIKNVHGSAKWMSSKMRLMKKMMSTPATDKANNSTTIPISPRIQNQGHENRY 157
+F E + + K S +L +STP + S+ P P++Q Q ++ +
Sbjct: 209 TFETSHEAALAYDAAARKLYGSDAKLNLPELSTPPQN--TTSSPSPTPPQMQQQ-QQHPH 379
Query: 158 SQRSPRNNNNSSNTTRVCSDCNTSS 182
Q P NNNN +N+ +C++ N ++
Sbjct: 380 IQIQPNNNNNINNSFNICNNINMNN 454
>TC83506 similar to PIR|T08179|T08179 LRG5 protein - Chlamydomonas
reinhardtii, partial (2%)
Length = 539
Score = 33.9 bits (76), Expect = 0.082
Identities = 21/93 (22%), Positives = 39/93 (41%), Gaps = 5/93 (5%)
Frame = +3
Query: 89 SSSTCDPNLSF-----SKMEEEDIKNVHGSAKWMSSKMRLMKKMMSTPATDKANNSTTIP 143
S S C P+L + ++E KN H + W S+ + T + + S+ P
Sbjct: 219 SPSICSPDLDIL*RPDKETKKEKKKNHHKTRTWPMSRSGRKQPPQQTRSPRRCRRSSARP 398
Query: 144 ISPRIQNQGHENRYSQRSPRNNNNSSNTTRVCS 176
+PR + +PR ++++ + TR S
Sbjct: 399 AAPRAATSTRRGAGTSSTPRTSSSTPSRTRPTS 497
>TC81171 similar to GP|9795609|gb|AAF98427.1| Unknown protein {Arabidopsis
thaliana}, partial (40%)
Length = 932
Score = 31.6 bits (70), Expect = 0.41
Identities = 28/117 (23%), Positives = 52/117 (43%), Gaps = 10/117 (8%)
Frame = +3
Query: 134 DKANNSTTIPISPRIQNQGHENRYSQ-RSPRNNNNSSNTTRVCSDCNT---------SST 183
++ ++ T++P SP+ Q+ GH + ++Q SPR + S T+ S+ + S
Sbjct: 42 EQKSSLTSLP-SPKTQSNGHNHSHNQIPSPRPISLPSPKTQTQSNGHNHNHNHNQIPSPR 218
Query: 184 PLWRSGPNGPKSLCNACGIRQRKARRAMAEAANGLATPINTASTKTRVHHNKEKKPR 240
P+ RS P P + + + + G + AST T+ +HN P+
Sbjct: 219 PITRSEPGNPYPTTFVQA--DTTSFKQVVQMLTGSSETAKQASTSTKANHNHNIPPK 383
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.311 0.125 0.363
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 10,956,653
Number of Sequences: 36976
Number of extensions: 174741
Number of successful extensions: 1145
Number of sequences better than 10.0: 63
Number of HSP's better than 10.0 without gapping: 1103
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1125
length of query: 310
length of database: 9,014,727
effective HSP length: 96
effective length of query: 214
effective length of database: 5,465,031
effective search space: 1169516634
effective search space used: 1169516634
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.9 bits)
S2: 58 (26.9 bits)
Lotus: description of TM0019a.4


