

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0016.15
(1469 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
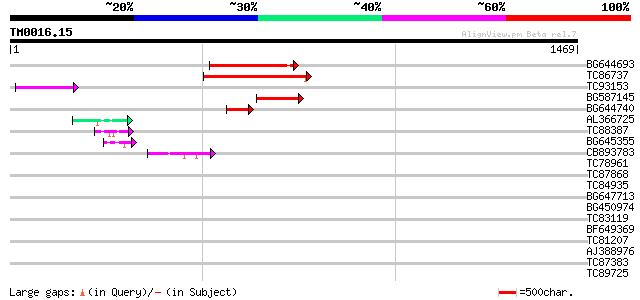
Score E
Sequences producing significant alignments: (bits) Value
BG644693 weakly similar to GP|18767374|g Putative 22 kDa kafirin... 228 1e-59
TC86737 weakly similar to GP|6683624|dbj|BAA89272.1 Pol {Alterna... 225 1e-58
TC93153 similar to GP|14715220|emb|CAC44106. gag polyprotein {Ci... 100 3e-21
BG587145 similar to PIR|H86337|H8 protein F5M15.26 [imported] - ... 96 1e-19
BG644740 similar to PIR|A84460|A84 probable retroelement pol pol... 87 6e-17
AL366725 57 7e-08
TC88387 similar to GP|13357253|gb|AAK20050.1 putative zinc finge... 48 3e-05
BG645355 similar to PIR|G96590|G965 hypothetical protein T24C10.... 47 4e-05
CB893783 weakly similar to GP|22830935|dbj hypothetical protein~... 47 5e-05
TC78961 similar to GP|18252179|gb|AAL61922.1 unknown protein {Ar... 42 0.001
TC87868 similar to PIR|T05112|T05112 splicing factor 9G8-like SR... 42 0.002
TC84935 similar to PIR|G96631|G96631 probable RNA-binding protei... 39 0.011
BG647713 homologue to GP|15042313|gb| 232R {Chilo iridescent vir... 39 0.011
BG450974 similar to PIR|T05112|T05 splicing factor 9G8-like SR p... 39 0.019
TC83119 similar to GP|13357253|gb|AAK20050.1 putative zinc finge... 38 0.025
BF649369 38 0.025
TC81207 similar to GP|21322752|dbj|BAB78536. cold shock protein-... 38 0.025
AJ388976 similar to PIR|E84638|E84 probable RSZp22 splicing fact... 38 0.033
TC87383 similar to GP|19168656|emb|CAD26175. DNA-DIRECTED RNA PO... 37 0.057
TC89725 similar to PIR|T05494|T05494 glycine-rich protein T19K4.... 37 0.057
>BG644693 weakly similar to GP|18767374|g Putative 22 kDa kafirin cluster;
Ty3-Gypsy type {Oryza sativa}, partial (15%)
Length = 716
Score = 228 bits (581), Expect = 1e-59
Identities = 127/235 (54%), Positives = 155/235 (65%), Gaps = 5/235 (2%)
Frame = +2
Query: 518 LPPVRDIEFAIDIVPGTGPISIAPYRMAPAELVELKSQLEDLLSKGFIRPSVSPWGAPVL 577
+PP I+F ID++P PI I YR+ P +L LK QL+DLL KGFI+PS+ P G VL
Sbjct: 17 VPPEWKIDFGIDLLPNMNPI*IPSYRINPLKLKVLKLQLKDLLEKGFIQPSIYP*GVVVL 196
Query: 578 LVKKKDGKSRLCVDYRQLNKVTVKNRYPLPRIDDLMDQLRGAAIFSKIDLKSGYHQIRVK 637
+KKKDG R+ +DY QLN V +K +YPLP ID+L D L+G+ F KIDL+ G HQ RV
Sbjct: 197 FLKKKDGFLRMSIDYPQLNNVNIKIKYPLPLIDELFDNLQGSKWFFKIDLRLG*HQHRVI 376
Query: 638 TEDIQKTAFRTRYGHYEYLVMPFGVTNAPAIFMDYMNRTFHTFLDRFVVVFIDDILIYSK 697
ED+ KTAFR RYGHYE LVM FG TN P FM+ MNR F +LD V+VF +DILIYSK
Sbjct: 377 GEDVPKTAFRIRYGHYEILVMSFG*TNPPMAFMELMNRVFQDYLDSLVIVFSNDILIYSK 556
Query: 698 NVVEHEGHLRQVLQVLREKELYANPSKCEF----WLEEVKFLG-HVISKEGIAVD 747
N EHE HLR L+VL++ L C+ L EV F HVIS EG+ VD
Sbjct: 557 NENEHENHLRLALKVLKDIGL------CQISYV*ILVEVGFFSLHVISGEGLKVD 703
>TC86737 weakly similar to GP|6683624|dbj|BAA89272.1 Pol {Alternaria
alternata}, partial (21%)
Length = 1540
Score = 225 bits (573), Expect = 1e-58
Identities = 126/294 (42%), Positives = 185/294 (62%), Gaps = 15/294 (5%)
Frame = +1
Query: 503 VVKEFTDVF-PGDVPGLPPVRDI-EFAIDIVPGTG----PISIAP-YRMAPAELVELKSQ 555
V++EF D+F P +P R + + AI ++P P+ P Y M+ EL+ LK
Sbjct: 343 VLEEFPDLFNPEKAYQVPASRGLLDHAIPLIPDKDGNDPPLPWGPLYGMSRQELLVLKKT 522
Query: 556 LEDLLSKGFIRPSVSPWGAPVLLVKKKDGKSRLCVDYRQLNKVTVKNRYPLPRIDDLMDQ 615
LEDLL KGFI+ S S GAPVL V+K G R CVDYR LN +T K+RYPLP I + + +
Sbjct: 523 LEDLLDKGFIKASGSAAGAPVLFVRKPGGGIRFCVDYRALNAITKKDRYPLPLISETLRR 702
Query: 616 LRGAAIFSKIDLKSGYHQIRVKTEDIQKTAFRTRYGHYEYLVMPFGVTNAPAIFMDYMNR 675
+ GA F+K+D+ + +H++R+K ED +KTAFRTRYG +E++V PFG+T APA F Y+N+
Sbjct: 703 VAGARWFTKLDVVAAFHKMRIKDEDQEKTAFRTRYGLFEWIVCPFGLTGAPATFQRYINK 882
Query: 676 TFHTFLDRFVVVFIDDILIYSK-NVVEHEGHLRQVLQVLREKELYANPSKCEFWLEEVKF 734
T H FLD FV +IDD+LIY+ + +HE +R+VL+ L + L +P KCEF + VK+
Sbjct: 883 TLHEFLDDFVTAYIDDVLIYTTGSKKDHEAQVRRVLRRLADAGLSLDPKKCEFSVTTVKY 1062
Query: 735 LGHVISK-EGIAVDPAKVETVLSWERPKTV------LGVSLVWRDIIVALSRIS 781
+G +++ +G++ DP K+ + W P +V LG ++D I S I+
Sbjct: 1063VGFILTAGKGVSCDPLKLAAIRDWLPPGSVKGARSFLGFCNYYKDFIPGYSEIT 1224
>TC93153 similar to GP|14715220|emb|CAC44106. gag polyprotein {Cicer
arietinum}, partial (8%)
Length = 516
Score = 100 bits (250), Expect = 3e-21
Identities = 57/167 (34%), Positives = 88/167 (52%), Gaps = 2/167 (1%)
Frame = +2
Query: 14 ANDSAMRRAAEEAREQHQRQREVTL--DQNKGLNDFRRHDPPKFSGGTDPDKADLWIQEI 71
++D+A+ A E + Q+ +V D + L F R+ PP F G PD A W++EI
Sbjct: 11 SSDAALVAALEAVAQAVQQLPKVDTGSDGTRMLETFLRNHPPTFKGRYAPDGA*KWLKEI 190
Query: 72 EKIFGVLQTAEGAKLGMATYLLLGDAEYWWKGTRGIMEDNHEEIN*NSFRMAFLEKYFPT 131
E+IF V+Q E K+ T++L +A+ WW ++E + + FR FL +YFP
Sbjct: 191 ERIFRVMQCFETQKVQFGTHMLAEEADDWWISLLPVLEQDDAVVTWAMFRKEFLGRYFPE 370
Query: 132 SARDERESQFLTLRQGGMSVPEFASKLESLAKHFQFFHDHVNERYMC 178
R ++E +FL L+QG MSV E+A+K LA + + E C
Sbjct: 371 DVRGKKEIEFLELKQGDMSVTEYAAKFVELATFYPHYSAETAEFSKC 511
>BG587145 similar to PIR|H86337|H8 protein F5M15.26 [imported] - Arabidopsis
thaliana, partial (13%)
Length = 763
Score = 95.5 bits (236), Expect = 1e-19
Identities = 46/123 (37%), Positives = 75/123 (60%)
Frame = +2
Query: 639 EDIQKTAFRTRYGHYEYLVMPFGVTNAPAIFMDYMNRTFHTFLDRFVVVFIDDILIYSKN 698
+D++KTAF T G Y Y VMPFG+ NA + + +NR F L + V+IDD+L+ S
Sbjct: 11 DDLEKTAFITDRGTYCYKVMPFGLKNAGSTYQRLVNRMFADKLGNTMEVYIDDMLVKSLR 190
Query: 699 VVEHEGHLRQVLQVLREKELYANPSKCEFWLEEVKFLGHVISKEGIAVDPAKVETVLSWE 758
+H HL++ + L E + NP+KC F + +FLG++++++GI V+P ++ +L
Sbjct: 191 ATDHLNHLKE*FKTLDEYIMKLNPAKCTFGVTSGEFLGYIVTQQGIEVNPKQITAILDLP 370
Query: 759 RPK 761
PK
Sbjct: 371 SPK 379
>BG644740 similar to PIR|A84460|A84 probable retroelement pol polyprotein
[imported] - Arabidopsis thaliana, partial (4%)
Length = 754
Score = 86.7 bits (213), Expect = 6e-17
Identities = 40/69 (57%), Positives = 52/69 (74%)
Frame = -1
Query: 562 KGFIRPSVSPWGAPVLLVKKKDGKSRLCVDYRQLNKVTVKNRYPLPRIDDLMDQLRGAAI 621
K F +PS+SP GA +L V+KKDG R+C+DYRQ NKVT KN+YPLPRID+L D+++
Sbjct: 274 KRFQQPSISP*GAALLFVRKKDGYFRMCIDYRQFNKVTTKNKYPLPRIDNLFDKIQEDCY 95
Query: 622 FSKIDLKSG 630
F IDL+ G
Sbjct: 94 F*NIDLRLG 68
>AL366725
Length = 485
Score = 56.6 bits (135), Expect = 7e-08
Identities = 44/169 (26%), Positives = 65/169 (38%), Gaps = 13/169 (7%)
Frame = +2
Query: 163 KHFQFFHDHVNERYMCKRFVNGLRPDIEDSVRPLGIMRFQSLVEKATEVE---LMKNRRL 219
K + + E C +F NGLRPDI+ ++ + F LV E ++ +
Sbjct: 2 KFYPHYAAETAEFSKCIKFENGLRPDIKRAIGYQQLRVFPDLVNTCRIYEEDTKAHDKVV 181
Query: 220 NRAGTGG----------PMRSGSQNFQNRGKFQNRRPYQRPAGRGFASGSYRPMVGAAGG 269
N T G P G Q + +RRP ++ A +Y G G
Sbjct: 182 NERKTKGQ*SRPKPYSAPADKGKQRMVD-----DRRPKKKDAPAEIVCFNY----GEKGH 334
Query: 270 SGDQTQNRELTCFKCGKPGHFARACPDTRPQCYNCNKLGHTAAQCRVPK 318
+ C +C K GH C C+NCN+ GH +QC+ PK
Sbjct: 335 KSNVCPKEIKKCVRCDKKGHIVADCKRNDIVCFNCNEEGHIGSQCKQPK 481
>TC88387 similar to GP|13357253|gb|AAK20050.1 putative zinc finger protein
{Oryza sativa (japonica cultivar-group)}, partial (96%)
Length = 1286
Score = 47.8 bits (112), Expect = 3e-05
Identities = 36/124 (29%), Positives = 55/124 (44%), Gaps = 21/124 (16%)
Frame = +1
Query: 219 LNRAGTGGPM--RSGSQNFQNRGKFQNRRPYQRPAGRGFAS----------GSYR---PM 263
++R+ + P+ R S+ F +R PY+R + RGF+ G Y P
Sbjct: 307 MDRSRSRSPVDRRIRSERFSHR-----EAPYRRDSRRGFSQDNLCKNCKRPGHYVRECPN 471
Query: 264 VGAA------GGSGDQTQNRELTCFKCGKPGHFARACPDTRPQCYNCNKLGHTAAQCRVP 317
V G + + L C+ C +PGH A +CP+ C+ C K GH A +C VP
Sbjct: 472 VAVCHNCSLPGHIASECSTKSL-CWNCKEPGHMASSCPN-EGICHTCGKAGHRARECTVP 645
Query: 318 KAEP 321
+ P
Sbjct: 646 QKPP 657
Score = 40.8 bits (94), Expect = 0.004
Identities = 19/37 (51%), Positives = 20/37 (53%)
Frame = +1
Query: 278 ELTCFKCGKPGHFARACPDTRPQCYNCNKLGHTAAQC 314
E C C K GH AR CP+ P C CN GH A QC
Sbjct: 721 EKACNNCRKTGHLARDCPND-PICNLCNISGHVARQC 828
Score = 39.7 bits (91), Expect = 0.009
Identities = 22/47 (46%), Positives = 23/47 (48%), Gaps = 8/47 (17%)
Frame = +1
Query: 281 CFKCGKPGHFARACPDTRPQ--------CYNCNKLGHTAAQCRVPKA 319
C CGK GH AR C T PQ C NC K GH A +C KA
Sbjct: 595 CHTCGKAGHRAREC--TVPQKPPGDLRLCNNCYKQGHIAVECTNEKA 729
Score = 32.0 bits (71), Expect = 1.8
Identities = 18/59 (30%), Positives = 25/59 (41%), Gaps = 7/59 (11%)
Frame = +1
Query: 263 MVGAAGGSGDQTQN------RELTCFKCGKPGHFARACPDTRPQ-CYNCNKLGHTAAQC 314
++G GG G R++ C C + GH +R C C NC GH A +C
Sbjct: 841 VIGDRGGGGSLRGGYRDGGFRDVVCRSCQQFGHMSRDCMGGPLMICQNCGGRGHQAYEC 1017
>BG645355 similar to PIR|G96590|G965 hypothetical protein T24C10.5 [imported]
- Arabidopsis thaliana, partial (5%)
Length = 627
Score = 47.4 bits (111), Expect = 4e-05
Identities = 30/103 (29%), Positives = 44/103 (42%), Gaps = 17/103 (16%)
Frame = -1
Query: 244 RRPYQRPAGRGFASGSYRPMVGAAGGSGDQTQNRELTCFKCGKPGHFARACPD------- 296
R P ++ AG G+Y V +GG+ + C+KC +PGH+A CP
Sbjct: 576 RNPPEQTAG-----GAYVNTVSGSGGASGK-------CYKCQQPGHWASNCPSMSAANRV 433
Query: 297 ------TRPQCYNCNKLGHTAAQC----RVPKAEPTVNTARGK 329
CY CN+ GH A C P++ NT +G+
Sbjct: 432 SGGSGGASGNCYKCNQPGHWANNCPNMSAAPQSHGNSNTGQGR 304
>CB893783 weakly similar to GP|22830935|dbj hypothetical protein~similar to
gag-pol polyprotein {Oryza sativa (japonica
cultivar-group)}, partial (8%)
Length = 853
Score = 47.0 bits (110), Expect = 5e-05
Identities = 54/207 (26%), Positives = 88/207 (42%), Gaps = 31/207 (14%)
Frame = +2
Query: 358 IDGNLLSVLFDSGATHSFISRECAIRLRLPIT--ALSFDLIVTTPAKTLLANTACMYCSV 415
+ G + V+ DSG+ + +S +L LP + L + + C+ V
Sbjct: 245 LQGIVCDVIIDSGSCENVVSNYMVEKLELPTKDHPHRYKLQWLKKGNEVRVSKCCL---V 415
Query: 416 TYN-NRVYHANLVC--LTLKNLDVILGMDWLSHYHCLLD------------CN*KRVVFP 460
+++ + Y N+ C +++ ++LG W H L D K V P
Sbjct: 416 SFSIGQKYKDNVWCDVISMDACHMLLGRPWQYDRHALYDGHANTYTFVKYGVKIKLVPLP 595
Query: 461 DSVLSEYLFANHISVSLNGGS---------QEYSVLLSL----ESKGNPEVDSIPVVKEF 507
+ E VSL Q+ S++L + ES EV+ + V +F
Sbjct: 596 PNAFDEGKKDFKPIVSLVSKEPFKVTTKDIQDMSLILLVKSNEESTIQKEVEHLLV--DF 769
Query: 508 TDVFPGDVP-GLPPVRDIEFAIDIVPG 533
TDV P ++P GLPP+RDI+ AID +PG
Sbjct: 770 TDVVPSEIPSGLPPMRDIQHAIDFIPG 850
>TC78961 similar to GP|18252179|gb|AAL61922.1 unknown protein {Arabidopsis
thaliana}, partial (71%)
Length = 974
Score = 42.4 bits (98), Expect = 0.001
Identities = 22/63 (34%), Positives = 28/63 (43%), Gaps = 12/63 (19%)
Frame = +2
Query: 265 GAAGGSGDQTQNRELT----------CFKCGKPGHFARACP--DTRPQCYNCNKLGHTAA 312
G GS D +RE CF CG GH+AR C D + +CY C + GH
Sbjct: 281 GVPRGSRDSRDSREYLGRGPPPGSGRCFNCGIDGHWARDCKAGDWKNKCYRCGERGHIEK 460
Query: 313 QCR 315
C+
Sbjct: 461 NCK 469
>TC87868 similar to PIR|T05112|T05112 splicing factor 9G8-like SR protein
RSZp22 [validated] - Arabidopsis thaliana, partial (91%)
Length = 860
Score = 41.6 bits (96), Expect = 0.002
Identities = 20/46 (43%), Positives = 23/46 (49%)
Frame = +3
Query: 249 RPAGRGFASGSYRPMVGAAGGSGDQTQNRELTCFKCGKPGHFARAC 294
R G G G R G GG D L C++CG+PGHFAR C
Sbjct: 258 RSGGGGGGGGGGRGRGGGGGGGSD------LKCYECGEPGHFAREC 377
>TC84935 similar to PIR|G96631|G96631 probable RNA-binding protein F8A5.17
[imported] - Arabidopsis thaliana, partial (41%)
Length = 552
Score = 39.3 bits (90), Expect = 0.011
Identities = 19/42 (45%), Positives = 26/42 (61%)
Frame = +2
Query: 254 GFASGSYRPMVGAAGGSGDQTQNRELTCFKCGKPGHFARACP 295
GF+SG + G+GD+ + CFKCG+PGH+AR CP
Sbjct: 413 GFSSGGR-----GSYGAGDRVGQDD--CFKCGRPGHWARDCP 517
>BG647713 homologue to GP|15042313|gb| 232R {Chilo iridescent virus}, partial
(1%)
Length = 726
Score = 39.3 bits (90), Expect = 0.011
Identities = 19/55 (34%), Positives = 27/55 (48%)
Frame = +2
Query: 51 DPPKFSGGTDPDKADLWIQEIEKIFGVLQTAEGAKLGMATYLLLGDAEYWWKGTR 105
D P F G PD+ W+Q IE++F + AE K+ + L A WWK +
Sbjct: 341 DIPDF*GKLQPDEFVDWLQTIERVFKYKEVAEEQKVKIVAAKLKKHASIWWKNLK 505
>BG450974 similar to PIR|T05112|T05 splicing factor 9G8-like SR protein
RSZp22 [validated] - Arabidopsis thaliana, partial (54%)
Length = 364
Score = 38.5 bits (88), Expect = 0.019
Identities = 17/39 (43%), Positives = 22/39 (55%), Gaps = 3/39 (7%)
Frame = +1
Query: 259 SYRPMVGAAGGSGDQTQNR---ELTCFKCGKPGHFARAC 294
S+ G GG G R +L C++CG+PGHFAR C
Sbjct: 244 SHNSKSGGGGGRGGGRGGRGGDDLKCYECGEPGHFAREC 360
>TC83119 similar to GP|13357253|gb|AAK20050.1 putative zinc finger protein
{Oryza sativa (japonica cultivar-group)}, partial (16%)
Length = 421
Score = 38.1 bits (87), Expect = 0.025
Identities = 30/109 (27%), Positives = 50/109 (45%), Gaps = 7/109 (6%)
Frame = +3
Query: 203 SLVEKATEVELMKNRRLNRAGTGGPMRSGSQNFQNR-GKFQNRR------PYQRPAGRGF 255
SLV++ + + +K +R+ + RS S++ R K ++ R PY+R + RGF
Sbjct: 153 SLVKEVKKEKNLKMSSDSRSRSRSRSRSRSRSRSPRIRKIRSDRHSYRDAPYRRDSSRGF 332
Query: 256 ASGSYRPMVGAAGGSGDQTQNRELTCFKCGKPGHFARACPDTRPQCYNC 304
+ R+ C C +PGH+AR CP+ C+NC
Sbjct: 333 S--------------------RDNLCKNCKRPGHYARECPNV-AVCHNC 416
>BF649369
Length = 631
Score = 38.1 bits (87), Expect = 0.025
Identities = 41/181 (22%), Positives = 73/181 (39%)
Frame = +3
Query: 32 RQREVTLDQNKGLNDFRRHDPPKFSGGTDPDKADLWIQEIEKIFGVLQTAEGAKLGMATY 91
R+ V QN+ ++ P F G D WI E F V T + ++ ++
Sbjct: 72 REEYVESSQNESRLAGKKVKLPLFEG----DDPVAWITRAEIYFDVQNTPDDMRVKLSRL 239
Query: 92 LLLGDAEYWWKGTRGIMEDNHEEIN*NSFRMAFLEKYFPTSARDERESQFLTLRQGGMSV 151
+ G +W+ ++ + ++++ + A + +Y + E + TLRQ G SV
Sbjct: 240 SMEGPTIHWF----NLLMETEDDLSREKLKKALIARYDGRRLENPFE-ELSTLRQIG-SV 401
Query: 152 PEFASKLESLAKHFQFFHDHVNERYMCKRFVNGLRPDIEDSVRPLGIMRFQSLVEKATEV 211
EF E L+ + E F++GL+ I VR L ++ A +V
Sbjct: 402 EEFVEAFELLSSQV----GRLPEEQYLGYFMSGLKAHIRRRVRTLNPTTRMQMMRIAKDV 569
Query: 212 E 212
E
Sbjct: 570 E 572
>TC81207 similar to GP|21322752|dbj|BAB78536. cold shock protein-1 {Triticum
aestivum}, partial (39%)
Length = 630
Score = 38.1 bits (87), Expect = 0.025
Identities = 21/66 (31%), Positives = 25/66 (37%), Gaps = 14/66 (21%)
Frame = +3
Query: 268 GGSGDQTQNRELTCFKCGKPGHFARACP--------------DTRPQCYNCNKLGHTAAQ 313
GG GD R+ C+ CG HFAR C CY C +GH A
Sbjct: 435 GGGGD----RDRACYTCGSFEHFARDCMRGGGNNNNGGGGYGGGGTSCYRCGGVGHIARD 602
Query: 314 CRVPKA 319
C P +
Sbjct: 603 CATPSS 620
Score = 34.3 bits (77), Expect = 0.37
Identities = 24/80 (30%), Positives = 25/80 (31%), Gaps = 17/80 (21%)
Frame = +3
Query: 252 GRGFASGSYRPMVGAAGGSGDQTQNRELTCFKCGKPGHFARACP---------------- 295
GRGF G R GG G C+ CG GH AR C
Sbjct: 306 GRGFRGGERRN-----GGGG---------CYTCGDTGHIARDCDRSDRNDRNDRSGGGGG 443
Query: 296 -DTRPQCYNCNKLGHTAAQC 314
D CY C H A C
Sbjct: 444 GDRDRACYTCGSFEHFARDC 503
>AJ388976 similar to PIR|E84638|E84 probable RSZp22 splicing factor
[imported] - Arabidopsis thaliana, partial (62%)
Length = 508
Score = 37.7 bits (86), Expect = 0.033
Identities = 19/43 (44%), Positives = 23/43 (53%)
Frame = +2
Query: 252 GRGFASGSYRPMVGAAGGSGDQTQNRELTCFKCGKPGHFARAC 294
G G G R G +GG G +L C+ CG+PGHFAR C
Sbjct: 281 GGGGGGGGGR---GRSGGGGS-----DLKCYXCGEPGHFARXC 385
>TC87383 similar to GP|19168656|emb|CAD26175. DNA-DIRECTED RNA POLYMERASE II
{Encephalitozoon cuniculi}, partial (0%)
Length = 1247
Score = 37.0 bits (84), Expect = 0.057
Identities = 24/94 (25%), Positives = 40/94 (42%)
Frame = -2
Query: 12 VQANDSAMRRAAEEAREQHQRQREVTLDQNKGLNDFRRHDPPKFSGGTDPDKADLWIQEI 71
V ++DS R++ R + Q +ND + D P F G PD W+Q +
Sbjct: 808 VSSSDSLSSRSSRSQRREFQ------------MNDIK*-DIPDFEGNLQPDDLLDWLQIM 668
Query: 72 EKIFGVLQTAEGAKLGMATYLLLGDAEYWWKGTR 105
E++F + E K+ + L A WW+ +
Sbjct: 667 ERLFKYKEVLEEQKVKIVAAKLKKLASIWWENVK 566
>TC89725 similar to PIR|T05494|T05494 glycine-rich protein T19K4.150 -
Arabidopsis thaliana, partial (17%)
Length = 378
Score = 37.0 bits (84), Expect = 0.057
Identities = 18/45 (40%), Positives = 22/45 (48%)
Frame = +1
Query: 251 AGRGFASGSYRPMVGAAGGSGDQTQNRELTCFKCGKPGHFARACP 295
+G G G Y G GG G +C+ CG+ GHFAR CP
Sbjct: 61 SGGGGGGGRYGGGGGGGGGGGGGG-----SCYSCGESGHFARDCP 180
Score = 31.2 bits (69), Expect = 3.1
Identities = 15/54 (27%), Positives = 19/54 (34%), Gaps = 20/54 (37%)
Frame = +1
Query: 281 CFKCGKPGHFARACPD--------------------TRPQCYNCNKLGHTAAQC 314
C+ CG+ GH AR C CY+C + GH A C
Sbjct: 16 CYNCGESGHMARECTSGGGGGGGRYGGGGGGGGGGGGGGSCYSCGESGHFARDC 177
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.345 0.152 0.510
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 46,926,357
Number of Sequences: 36976
Number of extensions: 711755
Number of successful extensions: 5919
Number of sequences better than 10.0: 54
Number of HSP's better than 10.0 without gapping: 2365
Number of HSP's successfully gapped in prelim test: 306
Number of HSP's that attempted gapping in prelim test: 3405
Number of HSP's gapped (non-prelim): 2931
length of query: 1469
length of database: 9,014,727
effective HSP length: 108
effective length of query: 1361
effective length of database: 5,021,319
effective search space: 6834015159
effective search space used: 6834015159
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.5 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 38 (21.6 bits)
S2: 65 (29.6 bits)
Lotus: description of TM0016.15


