

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
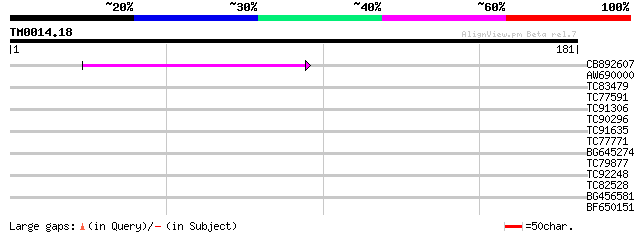
Query= TM0014.18
(181 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
CB892607 similar to GP|8927657|gb| EST gb|N38213 comes from this... 50 6e-07
AW690000 37 0.006
TC83479 similar to GP|23476992|emb|CAD48949. hypothetical protei... 36 0.010
TC77591 weakly similar to GP|21554074|gb|AAM63155.1 putative aci... 31 0.31
TC91306 weakly similar to PIR|T06371|T06371 probable UDP-glucuro... 28 1.5
TC90296 homologue to GP|22652121|gb|AAN03624.1 BEL1-related home... 28 2.0
TC91635 similar to PIR|A84854|A84854 hypothetical protein At2g42... 27 4.4
TC77771 homologue to GP|6137207|gb|AAF04377.1| P72 DEAD box prot... 27 4.4
BG645274 27 4.4
TC79877 similar to PIR|B86170|B86170 ADK1 [imported] - Arabidops... 27 5.8
TC92248 weakly similar to GP|22535591|dbj|BAC10766. putative DEI... 26 7.6
TC82528 weakly similar to PIR|C86356|C86356 hypothetical protein... 26 7.6
BG456581 26 9.9
BF650151 similar to GP|20161613|dbj endopeptidase-like protein {... 26 9.9
>CB892607 similar to GP|8927657|gb| EST gb|N38213 comes from this gene.
{Arabidopsis thaliana}, partial (1%)
Length = 782
Score = 49.7 bits (117), Expect = 6e-07
Identities = 29/73 (39%), Positives = 40/73 (54%)
Frame = +1
Query: 24 PLDAVWVKLNSDGAYISSADMAACRGIIRDRFSAFVIAYARRLGSYSTLQAELWGILCGA 83
P ++ V L D ++ AAC G+I+D FV YA +LGS S LQAELWGI G
Sbjct: 559 PPNSCEVALKCDYVVLNCDLNAACGGLIQDDQGHFVFHYANKLGSCSVLQAELWGI*HGL 738
Query: 84 RLLQERDY*RVLI 96
+ R Y ++ +
Sbjct: 739 SIDWNRGYSKIRV 777
>AW690000
Length = 652
Score = 36.6 bits (83), Expect = 0.006
Identities = 18/55 (32%), Positives = 29/55 (52%)
Frame = +3
Query: 22 WWPLDAVWVKLNSDGAYISSADMAACRGIIRDRFSAFVIAYARRLGSYSTLQAEL 76
W P W+K N+DG+ S + +AC GI R+ + ++ +A G + AEL
Sbjct: 411 WRPPIPHWIKCNTDGS--SRSHSSACGGIFRNHDTDLLLCFAENTGECNAFHAEL 569
>TC83479 similar to GP|23476992|emb|CAD48949. hypothetical protein {Plasmodium
falciparum 3D7}, partial (0%)
Length = 1222
Score = 35.8 bits (81), Expect = 0.010
Identities = 22/78 (28%), Positives = 36/78 (45%), Gaps = 1/78 (1%)
Frame = +2
Query: 6 QVTRVVSPV-RDAPAIRWWPLDAVWVKLNSDGAYISSADMAACRGIIRDRFSAFVIAYAR 64
Q+ +++PV R W + W KLN+DG+ + + A G++RD + A+
Sbjct: 926 QIANILNPVSRSIIWCEWKKPEIGWTKLNTDGSV--NKETAGFGGLLRDYRGEPICAFVS 1099
Query: 65 RLGSYSTLQAELWGILCG 82
+ T ELW I G
Sbjct: 1100 KAPQGDTFLVELWAIWRG 1153
>TC77591 weakly similar to GP|21554074|gb|AAM63155.1 putative acid
phosphatase {Arabidopsis thaliana}, partial (74%)
Length = 1067
Score = 30.8 bits (68), Expect = 0.31
Identities = 16/42 (38%), Positives = 24/42 (57%)
Frame = -1
Query: 95 LIETNLSILGVVLVLIPVLLLLTK*SRLFSRVSPRLGAVTCS 136
++++ I G + +P+L LL *SR FS SP L TC+
Sbjct: 806 ILQSTEIIFGPTMFPLPILALLII*SRTFSSSSPSLQPCTCA 681
>TC91306 weakly similar to PIR|T06371|T06371 probable
UDP-glucuronosyltransferase (EC 2.4.1.-) - garden pea,
partial (31%)
Length = 724
Score = 28.5 bits (62), Expect = 1.5
Identities = 8/16 (50%), Positives = 13/16 (81%)
Frame = +1
Query: 18 PAIRWWPLDAVWVKLN 33
P+ WW LD++W++LN
Sbjct: 169 PSFNWWILDSLWMELN 216
>TC90296 homologue to GP|22652121|gb|AAN03624.1 BEL1-related homeotic
protein 14 {Solanum tuberosum}, partial (15%)
Length = 715
Score = 28.1 bits (61), Expect = 2.0
Identities = 12/21 (57%), Positives = 14/21 (66%)
Frame = -2
Query: 49 GIIRDRFSAFVIAYARRLGSY 69
G +RDRF AY RRLG+Y
Sbjct: 63 GFLRDRFGGPKYAYQRRLGTY 1
>TC91635 similar to PIR|A84854|A84854 hypothetical protein At2g42450
[imported] - Arabidopsis thaliana, partial (71%)
Length = 1919
Score = 26.9 bits (58), Expect = 4.4
Identities = 22/71 (30%), Positives = 33/71 (45%), Gaps = 1/71 (1%)
Frame = +1
Query: 11 VSPVRDAPAIRWWPLDAVWVKLNSDGAYISSADMAACRGIIRDRFSAFVIAYARR-LGSY 69
V PV++ P R PL VK NSD I A + + F + Y +R LGS
Sbjct: 1339 VDPVKELPVAREAPLPPKEVKENSDVQKIEETKPA----VPEELFIPGTVYYLKRNLGSQ 1506
Query: 70 STLQAELWGIL 80
+ + E++ +L
Sbjct: 1507 NDVGKEVYTLL 1539
>TC77771 homologue to GP|6137207|gb|AAF04377.1| P72 DEAD box protein {Pisum
sativum}, partial (69%)
Length = 1736
Score = 26.9 bits (58), Expect = 4.4
Identities = 17/59 (28%), Positives = 31/59 (51%)
Frame = +2
Query: 77 WGILCGARLLQERDY*RVLIETNLSILGVVLVLIPVLLLLTK*SRLFSRVSPRLGAVTC 135
W + CG R+ +R VL ++ ++++L+ L LL LF+ ++PRL + C
Sbjct: 1337 WKLFCGHRIKGQR*LFSVL--PRKCVINLLVILLASLELL-----LFTGINPRLIGIMC 1492
>BG645274
Length = 641
Score = 26.9 bits (58), Expect = 4.4
Identities = 12/25 (48%), Positives = 13/25 (52%)
Frame = -3
Query: 29 WVKLNSDGAYISSADMAACRGIIRD 53
WVK N DG+ A C GI RD
Sbjct: 357 WVKCNIDGSIKGCFRPATCSGIFRD 283
>TC79877 similar to PIR|B86170|B86170 ADK1 [imported] - Arabidopsis
thaliana, partial (32%)
Length = 1283
Score = 26.6 bits (57), Expect = 5.8
Identities = 13/18 (72%), Positives = 14/18 (77%)
Frame = +3
Query: 51 IRDRFSAFVIAYARRLGS 68
IRDRFS V A+ARR GS
Sbjct: 426 IRDRFSGAVQAFARRNGS 479
>TC92248 weakly similar to GP|22535591|dbj|BAC10766. putative DEIH-box
RNA/DNA helicase {Oryza sativa (japonica
cultivar-group)}, partial (4%)
Length = 605
Score = 26.2 bits (56), Expect = 7.6
Identities = 12/47 (25%), Positives = 22/47 (46%)
Frame = -1
Query: 31 KLNSDGAYISSADMAACRGIIRDRFSAFVIAYARRLGSYSTLQAELW 77
+ +D +IS+ + C G R + + Y R GS+ ++ LW
Sbjct: 473 RFGTDPTWISNVP*SRCAGCRRSKVLVSTMIYKRCRGSFWSITIPLW 333
>TC82528 weakly similar to PIR|C86356|C86356 hypothetical protein AAF87257.1
[imported] - Arabidopsis thaliana, partial (35%)
Length = 1376
Score = 26.2 bits (56), Expect = 7.6
Identities = 7/16 (43%), Positives = 11/16 (68%)
Frame = +3
Query: 18 PAIRWWPLDAVWVKLN 33
P WW LD++W++ N
Sbjct: 927 PPFNWWILDSLWMEFN 974
>BG456581
Length = 683
Score = 25.8 bits (55), Expect = 9.9
Identities = 9/24 (37%), Positives = 17/24 (70%)
Frame = +2
Query: 19 AIRWWPLDAVWVKLNSDGAYISSA 42
++ W P A WVK N+DG+ ++++
Sbjct: 593 SVCWKPPSAPWVKGNTDGSXLNNS 664
>BF650151 similar to GP|20161613|dbj endopeptidase-like protein {Oryza sativa
(japonica cultivar-group)}, partial (3%)
Length = 652
Score = 25.8 bits (55), Expect = 9.9
Identities = 8/21 (38%), Positives = 13/21 (61%)
Frame = -2
Query: 15 RDAPAIRWWPLDAVWVKLNSD 35
R +PA+ WW +D +W+ D
Sbjct: 366 RGSPAVVWWWIDLLWLGEEKD 304
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.336 0.145 0.454
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 5,504,744
Number of Sequences: 36976
Number of extensions: 70945
Number of successful extensions: 507
Number of sequences better than 10.0: 28
Number of HSP's better than 10.0 without gapping: 503
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 505
length of query: 181
length of database: 9,014,727
effective HSP length: 90
effective length of query: 91
effective length of database: 5,686,887
effective search space: 517506717
effective search space used: 517506717
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (21.7 bits)
S2: 55 (25.8 bits)
Lotus: description of TM0014.18


