

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0014.15
(766 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
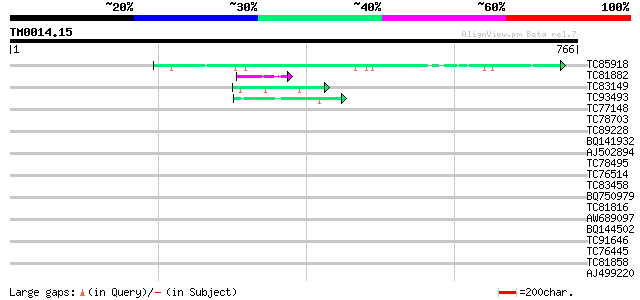
TC85918 similar to GP|13562014|gb|AAK30610.1 fibroin 1 {Plectreu... 45 8e-05
TC81882 similar to GP|9759349|dbj|BAB10004.1 contains similarity... 45 1e-04
TC83149 similar to GP|13561982|gb|AAK30594.1 flagelliform silk p... 42 7e-04
TC93493 42 9e-04
TC77148 ENOD20 40 0.003
TC78703 homologue to GP|13129456|gb|AAK13114.1 Putative retroele... 40 0.003
TC89228 similar to GP|5081555|gb|AAD39439.1| PHAP2A protein {Pet... 40 0.004
BQ141932 weakly similar to GP|6523547|emb hydroxyproline-rich gl... 40 0.004
AJ502894 weakly similar to SP|P38665|RL24 60S ribosomal protein ... 39 0.006
TC78495 homologue to PIR|S53504|S53504 extensin-like protein S3 ... 39 0.006
TC76514 homologue to PIR|T09648|T09648 nucleolin homolog nuM1 - ... 39 0.007
TC83458 homologue to PIR|T07796|T07796 DNA-directed RNA polymera... 39 0.010
BQ750979 similar to GP|13883467|gb PE_PGRS family protein {Mycob... 38 0.013
TC81816 similar to GP|8096269|dbj|BAA95789.1 KED {Nicotiana taba... 38 0.013
AW689097 weakly similar to PIR|T47565|T47 hypothetical protein F... 38 0.017
BQ144502 weakly similar to GP|6523547|emb hydroxyproline-rich gl... 38 0.017
TC91646 weakly similar to GP|6523547|emb|CAB62280.1 hydroxyproli... 37 0.022
TC76445 similar to PIR|T07623|T07623 extensin homolog HRGP2 - so... 37 0.022
TC81858 similar to GP|18252179|gb|AAL61922.1 unknown protein {Ar... 37 0.022
AJ499220 similar to SP|Q09863|YAF Hypothetical protein C29E6.10c... 37 0.028
>TC85918 similar to GP|13562014|gb|AAK30610.1 fibroin 1 {Plectreurys tristis},
partial (8%)
Length = 4117
Score = 45.4 bits (106), Expect = 8e-05
Identities = 108/630 (17%), Positives = 218/630 (34%), Gaps = 73/630 (11%)
Frame = -1
Query: 195 KKKTKKKMVLEESSEESDVPLVK-------KTKSKPDDGDDDDSEDGPPKKKQKKVRIVV 247
++K K+ + E S+E+ + P ++ K +P+ + SE+ K V V
Sbjct: 2692 EEKVKQAEISETSAEKEEKPELEPLATEEPKVTKEPEKESQEKSEEAEQPKAIAIVESTV 2513
Query: 248 KPTRVEPAAAVVRRTEAPARVTRSSAHSSKPALASDDDLNLFDALPISALLQHSSN---- 303
+ + AAA+ T+ V + +P + + P+ ++ +
Sbjct: 2512 EANEEKTAAAISEETKI-IEVEPTETVKEEPMVTEVATAEIVKEEPVVTEVEATETLKEE 2336
Query: 304 ------PLTPIPESQPAAQ----TTSPPHSPRSPFFQPSPT--EAPLWNLLKNPTSRSED 351
T + +PAA T + P P+ T E PL + + E+
Sbjct: 2335 LVATEVEATETVKEEPAATEVKVTETVKDEPVVTEVDPTETVKEEPLATEAEATETVKEE 2156
Query: 352 PTSLLTIPYDPVSSEPIIHEQPEPNQTEPQPRTSD-HSVPRASARPAARTTDTDSSTAFT 410
P + P + V EP+ E + +P ++ + P A T+ +
Sbjct: 2155 PVATEVEPTETVKEEPVATEVEATETVKDEPAATEVEATETVKEEPVATETEATETVKEE 1976
Query: 411 PVSFPINVIDSPPSNTSESIRKFMEVRKEK--VSALEEYYLTCPSPRRYPGPRPERL--- 465
PV+ + ++ + + + E KE+ + +E + P E +
Sbjct: 1975 PVATEVEATETVKEESVATEVEATETVKEEPVTTVVEPTEMVKDEPAATEFEATETVKDE 1796
Query: 466 -----VDPDEPILANPIQEA--------DPLVQQAH--------PVPDQQEPIQPDPEPE 504
VDP E + + +PLV + PV + +P + E
Sbjct: 1795 TIVTEVDPTETVKEEAVATEVEITKTVKEPLVTEVDLTESVKVKPVVTEVDPREMVKEEP 1616
Query: 505 QSVSNQSSVRSPHPLVETSDHHRGTSEPNAPMINIGSPQGASEAHSSNHQASPEPNLSII 564
+ Q++ V T T + + + Q E HS+ + +PN S I
Sbjct: 1615 VATEVQATETVKEEPVATEVEPTETVKGEPVVTEVEENQKEPEQHSTERREEEQPNASSI 1436
Query: 565 PYTRLQPTSLSECINIFYHEASLRLRNVHGQTDLSENAENVADEWNNLSTWLVAQVPIML 624
P Q T ++ I + + R + + + + +N DE T V P++
Sbjct: 1435 PE---QSTETNDVIAV-----EEKSRELEFEAAILKETKN--DEAGPAETEKVE--PVVT 1292
Query: 625 QLLHAEGSQSIEAAKQ--------------RLARRVALHE---------LEQRNKLLEAI 661
++ + I++ KQ + RV +H E N EAI
Sbjct: 1291 EVDENQSEPGIQSFKQEEEVKPKEETENQEQTGDRVEVHPPKDSDIEAVKETNNSGSEAI 1112
Query: 662 KEAKRKKDQAEEAARQAAAQAEQARLEAERLEAEAEAERLAPVVFTPAASASIPDPQAAQ 721
+ D + ++ Q E + E E E+ T ++ +I + +
Sbjct: 1111 LVKEENIDPLSNGVEEKLSEQLQVGEEVGEIIKEVEPEQATEK--TEGSNGTIKEEETEP 938
Query: 722 NVPSSSTQTSSSRLDIMEQRLDTHESMLIE 751
N ++ Q ++ +++E + E +++E
Sbjct: 937 NTTVNAAQVQTTPANLVEPSPEVEEKIVVE 848
>TC81882 similar to GP|9759349|dbj|BAB10004.1 contains similarity to unknown
protein~gb|AAF72944.1~gene_id:MAH20.11 {Arabidopsis
thaliana}, partial (5%)
Length = 875
Score = 44.7 bits (104), Expect = 1e-04
Identities = 26/76 (34%), Positives = 35/76 (45%)
Frame = +3
Query: 307 PIPESQPAAQTTSPPHSPRSPFFQPSPTEAPLWNLLKNPTSRSEDPTSLLTIPYDPVSSE 366
P P PA P +P + PS P NL KNP + S PT+ T P++ +
Sbjct: 111 PCPPPNPAISAAEPTQTPTTTLLPPSLPNPPPLNL-KNPPNFSVSPTTKST----PIT-K 272
Query: 367 PIIHEQPEPNQTEPQP 382
P++H P P T P P
Sbjct: 273 PLVHAPPNPTTTAPNP 320
>TC83149 similar to GP|13561982|gb|AAK30594.1 flagelliform silk protein
{Argiope trifasciata}, partial (3%)
Length = 1177
Score = 42.4 bits (98), Expect = 7e-04
Identities = 43/152 (28%), Positives = 51/152 (33%), Gaps = 21/152 (13%)
Frame = +3
Query: 301 SSNPLTPIPE---SQPAAQTTSPPHSPRSPFFQPSPTEAPLWNLLK-------------- 343
S PL P P P SPP RS SP PLW
Sbjct: 153 SKTPLPPPPPPLPQHPKPPQKSPPRMRRSKPSPSSPAATPLWPQRTTPPPSPSTPRPSP* 332
Query: 344 NPTSRSEDPTSLLTIPYDPVSSEPIIHEQPEPNQTEPQPRTSDHSV----PRASARPAAR 399
P + S PT+ L P + P + +P T P P+ SV P A+ R R
Sbjct: 333 TPATPSTSPTAPLPTPPPRTTLPPALTPRPPSPSTPPTPKPGPVSVSPASPSATQRVPWR 512
Query: 400 TTDTDSSTAFTPVSFPINVIDSPPSNTSESIR 431
T SST + P PS S S R
Sbjct: 513 PTARASSTKVPAAAKP*RRATRRPSAVSRSSR 608
>TC93493
Length = 456
Score = 42.0 bits (97), Expect = 9e-04
Identities = 42/161 (26%), Positives = 59/161 (36%), Gaps = 9/161 (5%)
Frame = +1
Query: 303 NPLTPIPESQPAAQTTSPPHSPRSPFFQPSPTE-APLWNLLKNP--TSRSEDPTSLLTIP 359
NP P P + P TT+P S P PSPT+ L + P RS P P
Sbjct: 4 NPAVPRPTASP---TTAPSFSAMHPSANPSPTKTTAAAELSRGPELPRRSRKPAKGTKRP 174
Query: 360 YDPVSSEPIIHEQPEPNQTEPQPRTSDHSVPRASARPAARTTDTDSSTAFTPVSFPI--- 416
P+I P P + P +TS HS A A T T +TA S
Sbjct: 175 ------APLITPPPGPPEATPS-KTSRHSPASAPATLFTSVTSTSHNTAANMASHTYRKE 333
Query: 417 ---NVIDSPPSNTSESIRKFMEVRKEKVSALEEYYLTCPSP 454
+D+ P N + + + +++ + PSP
Sbjct: 334 KYGEAVDAEPGNEKDGVNPLPTLAQKRPTKPSAQAAPVPSP 456
>TC77148 ENOD20
Length = 1108
Score = 40.0 bits (92), Expect = 0.003
Identities = 33/108 (30%), Positives = 47/108 (42%)
Frame = +3
Query: 306 TPIPESQPAAQTTSPPHSPRSPFFQPSPTEAPLWNLLKNPTSRSEDPTSLLTIPYDPVSS 365
TPIP P SPP SP PSP+ +P + +P RS P P S
Sbjct: 465 TPIPHP-PRRSLPSPPSPSPSPSPSPSPSPSPRSTPIPHPRKRS---------PASP-SP 611
Query: 366 EPIIHEQPEPNQTEPQPRTSDHSVPRASARPAARTTDTDSSTAFTPVS 413
P + + P P+++ + SV AS P++ +D S A +P S
Sbjct: 612 SPSLSKSPSPSESPSLAPSPSDSV--ASLAPSSSPSDESPSPAPSPSS 749
>TC78703 homologue to GP|13129456|gb|AAK13114.1 Putative retroelement pol
polyprotein {Oryza sativa} [Oryza sativa (japonica
cultivar-group)], partial (2%)
Length = 1235
Score = 40.0 bits (92), Expect = 0.003
Identities = 26/80 (32%), Positives = 36/80 (44%), Gaps = 7/80 (8%)
Frame = +3
Query: 295 SALLQHSSNPLTPIPESQPAAQTTSPP-------HSPRSPFFQPSPTEAPLWNLLKNPTS 347
S L HSS+ LTPIP P +++PP P SP + T P+ + +P
Sbjct: 534 SPLSSHSSDALTPIPPPSPLNGSSTPPSPVLDGSFPPPSPLDGSTLTPPPVQQVGSSPPP 713
Query: 348 RSEDPTSLLTIPYDPVSSEP 367
D T+ +T PVS P
Sbjct: 714 LGTDVTNPITPTQSPVSEPP 773
Score = 31.6 bits (70), Expect = 1.2
Identities = 35/121 (28%), Positives = 47/121 (37%), Gaps = 6/121 (4%)
Frame = +3
Query: 316 QTTSPPHSPRSPFFQPSPTEAPLWNLLKNPTSRSEDPTSLLTIPYDPVSSEPIIHEQPEP 375
QTTSPP PSP+ AP +P+ S + LT P P S P P
Sbjct: 486 QTTSPP--------SPSPSVAP------SPSPLSSHSSDALT-PIPPPSPLNGSSTPPSP 620
Query: 376 --NQTEPQPRTSDHSV----PRASARPAARTTDTDSSTAFTPVSFPINVIDSPPSNTSES 429
+ + P P D S P + TD + TP P++ + PP N + S
Sbjct: 621 VLDGSFPPPSPLDGSTLTPPPVQQVGSSPPPLGTDVTNPITPTQSPVS--EPPPPNAASS 794
Query: 430 I 430
I
Sbjct: 795 I 797
>TC89228 similar to GP|5081555|gb|AAD39439.1| PHAP2A protein {Petunia x
hybrida}, partial (25%)
Length = 774
Score = 39.7 bits (91), Expect = 0.004
Identities = 32/104 (30%), Positives = 45/104 (42%), Gaps = 1/104 (0%)
Frame = +2
Query: 138 EIRGRRMPLNDYPLWTQADHPDAIRCYV-EDLRAQGFEIDLDDFVSRLPPAPEFPSPPKK 196
E+R RRM W D PD + Y E +G E+D+D+ +
Sbjct: 233 ELR*RRM-------WDLNDSPDQRKNYESEGCSMEGDEVDVDEKGKGVGSV-------SN 370
Query: 197 KTKKKMVLEESSEESDVPLVKKTKSKPDDGDDDDSEDGPPKKKQ 240
+ +V+E+ SEE D + KK SK DDS D PP +Q
Sbjct: 371 SSSSAIVIEDGSEEEDGNMTKKKSSKIFGFSMDDSSDCPPVTRQ 502
>BQ141932 weakly similar to GP|6523547|emb hydroxyproline-rich glycoprotein
DZ-HRGP {Volvox carteri f. nagariensis}, partial (30%)
Length = 1338
Score = 39.7 bits (91), Expect = 0.004
Identities = 84/356 (23%), Positives = 125/356 (34%), Gaps = 42/356 (11%)
Frame = -3
Query: 224 DDGDDDDSEDGPPKKKQKKVRIVVKPT----RVEPAA---AVVRRTEA---PARVTRSSA 273
D D + P VR+V T R P++ +++RR A P + +
Sbjct: 1306 DSSPDQTTLSARPHAPATYVRLVPPATTSSARSTPSSIYSSMLRRLVASRRPLNPAHARS 1127
Query: 274 HSSK-PALASDDDLNLFDALPISALLQHSSNPLT-----PIPESQPAAQTTSPPHSPRSP 327
H + P L S + P ALL S PLT P P + P + P SP
Sbjct: 1126 HIIRVPPLPSPPTIPSSHLAPHRALLTLSL-PLTLSRASPRPPASPRRCLDTTPPSPHRH 950
Query: 328 FFQPSPTEAPLWNLLKNPTSRSE----------DPTSLLTIPYDPVSSEPIIHEQPEPNQ 377
+ + + P L P+ R++ P +L DP S P +
Sbjct: 949 YTHTTHSHPPQPPLRSPPSPRADFYDQHSTSHRHPPALYRYYTDPPRSTP-------HSA 791
Query: 378 TEPQPRTSDHSVPRASARPAARTTDTDSSTAFTPVSFPINVIDS--PPSN----TSESIR 431
T P T S R SA T T + FTP + S PP++ T +I
Sbjct: 790 TLPPASTHTRSCRRVSAIARRCTRRTREGSPFTPPPPRRTISGSSRPPASGAGSTGHAIP 611
Query: 432 KFMEVRKEKVSALEEYYLTCPSPRRYPGPRPERLVDPDEPILANPIQEADP------LVQ 485
F+ +++ + + P P P PR P L PI A P L
Sbjct: 610 YFVFGNRQRPTR----HPPPP*PPSIPIPRHPEPTHQPHPCLPPPIPSAPPPTRAFALPP 443
Query: 486 QAHPV----PDQQEPIQPDPEPEQSVSNQSSVRSPHPLVETSDHHRGTSEPNAPMI 537
A P P + P DP P + + +R +PL ++ P++P I
Sbjct: 442 SAPPPSDLPPPPRSPTVIDPRPPAASAVPLPLRPLYPLRDSPPRRPPPPPPHSPSI 275
Score = 35.0 bits (79), Expect = 0.11
Identities = 28/107 (26%), Positives = 40/107 (37%)
Frame = -3
Query: 293 PISALLQHSSNPLTPIPESQPAAQTTSPPHSPRSPFFQPSPTEAPLWNLLKNPTSRSEDP 352
P ++ + PL P+ +S P PPHSP + P+SR P
Sbjct: 376 PAASAVPLPLRPLYPLRDSPPRRPPPPPPHSPS----------------ILPPSSRLSPP 245
Query: 353 TSLLTIPYDPVSSEPIIHEQPEPNQTEPQPRTSDHSVPRASARPAAR 399
T L + P + P+ P P P P T+ P A P +R
Sbjct: 244 TP-LPLSSSP*PALPLPLPPPTPPPPPPAPTTTPI*APAAGYTPESR 107
Score = 32.0 bits (71), Expect = 0.91
Identities = 26/95 (27%), Positives = 31/95 (32%), Gaps = 21/95 (22%)
Frame = -1
Query: 309 PESQPAAQTTSPPHSPRSPFFQPSPTEAPLWNLLKNPTSRSEDPTSLLT----------- 357
P P A + SPP P P P P L P + P++L T
Sbjct: 480 PPPHPRAPSLSPPPPPHPPTSPPRPGRPRSSTLAPPPHRQFHSPSALCTHCGIPPPAVPP 301
Query: 358 --IPYDPVSSEPIIHE--------QPEPNQTEPQP 382
P P SS P + P PNQ P P
Sbjct: 300 PPPPTPPPSSPPALVSPPPRPFLFPPPPNQPSPSP 196
Score = 30.8 bits (68), Expect = 2.0
Identities = 42/184 (22%), Positives = 65/184 (34%), Gaps = 20/184 (10%)
Frame = -1
Query: 236 PKKKQKKVRIVVKPTRVEPAAAVV---RRTEAPARVTRSSAHSSKPA-----LASDDDLN 287
P+ + + + P + A AV+ R+ P V S AH ++P L +
Sbjct: 783 PQPLRTRAHVGAYPRLRDAAHAVLVKAHRSPRPPHVGPSLAHPARPRPGRVLLVMQYPTS 604
Query: 288 LFDALPISALLQHSSNP---LTPIPESQPAAQTTS------PPHSPRSPFFQPSPTEAPL 338
+ H NP +P ++P + T + PP PR+P P P P
Sbjct: 603 FLGTGSAPPAIHHHPNPPASQSPATLNRPTSHTPACPPPSRPPPHPRAPSLSPPPPPHPP 424
Query: 339 WNLLKNPTSRSEDPTSLLTIPYDPVSSEPIIHEQ---PEPNQTEPQPRTSDHSVPRASAR 395
+ + RS ++L P+ S + P P P P T S P A
Sbjct: 423 TSPPRPGRPRS---STLAPPPHRQFHSPSALCTHCGIPPPAVPPPPPPTPPPSSPPALVS 253
Query: 396 PAAR 399
P R
Sbjct: 252 PPPR 241
>AJ502894 weakly similar to SP|P38665|RL24 60S ribosomal protein L24 (L30).
[Yeast] {Kluyveromyces lactis}, partial (84%)
Length = 501
Score = 39.3 bits (90), Expect = 0.006
Identities = 27/78 (34%), Positives = 41/78 (51%)
Frame = +2
Query: 627 LHAEGSQSIEAAKQRLARRVALHELEQRNKLLEAIKEAKRKKDQAEEAARQAAAQAEQAR 686
+H +G E AK+R RR H+ EAIK + +K + EAAR AA+ +
Sbjct: 239 MHKKGITE-EVAKKR-TRRAVKHQRAIVGASWEAIKAKRNQKPEVREAARAEAARESKEN 412
Query: 687 LEAERLEAEAEAERLAPV 704
+AE+ + +AE +LA V
Sbjct: 413 KKAEQSKRKAEKAKLAQV 466
>TC78495 homologue to PIR|S53504|S53504 extensin-like protein S3 - alfalfa,
partial (97%)
Length = 1049
Score = 39.3 bits (90), Expect = 0.006
Identities = 29/125 (23%), Positives = 44/125 (35%), Gaps = 7/125 (5%)
Frame = +3
Query: 309 PESQPAAQTTSPPHSPRSPFFQPSPTEAPLWNLLKNPTSRSEDPTSLLTIPYDPVSSEPI 368
P S PA T P +P P P ++ + +P P + P SS P
Sbjct: 255 PNSSPAPPTPPANTPPTTPQASPPPVQSSPPPVQSSPPPLQSSPPPAQSTPPPVQSSPPP 434
Query: 369 IHEQPEPN-------QTEPQPRTSDHSVPRASARPAARTTDTDSSTAFTPVSFPINVIDS 421
+ + P P Q+ P P + + P + P A A P + + S
Sbjct: 435 VQQSPPPTPLTPPPVQSTPPPASPPPASPPPFSPPPATPPPATPPPATPPPALTPTPLSS 614
Query: 422 PPSNT 426
PP+ T
Sbjct: 615 PPATT 629
Score = 32.7 bits (73), Expect = 0.53
Identities = 41/131 (31%), Positives = 52/131 (39%), Gaps = 8/131 (6%)
Frame = +3
Query: 453 SPRRYPGP-RPERLVDPDEPILANP-IQEADPLVQQAHPVPDQQEPIQPDPEPEQSVSN- 509
SP P P P P P + P +Q + P VQ + P P+Q P P QS
Sbjct: 252 SPNSSPAPPTPPANTPPTTPQASPPPVQSSPPPVQSSPP------PLQSSPPPAQSTPPP 413
Query: 510 -QSS----VRSPHPLVETSDHHRGTSEPNAPMINIGSPQGASEAHSSNHQASPEPNLSII 564
QSS +SP P T + T P +P SP S ++ A+P P
Sbjct: 414 VQSSPPPVQQSPPPTPLTPPPVQSTPPPASP--PPASPPPFSPPPATPPPATPPP---AT 578
Query: 565 PYTRLQPTSLS 575
P L PT LS
Sbjct: 579 PPPALTPTPLS 611
Score = 31.2 bits (69), Expect = 1.6
Identities = 40/167 (23%), Positives = 53/167 (30%)
Frame = +3
Query: 315 AQTTSPPHSPRSPFFQPSPTEAPLWNLLKNPTSRSEDPTSLLTIPYDPVSSEPIIHEQPE 374
A +TSP SP P P P +P P + + P SS P P
Sbjct: 240 APSTSPNSSPAPP---TPPANTPPTTPQASPPPVQSSPPPVQSSPPPLQSSPPPAQSTPP 410
Query: 375 PNQTEPQPRTSDHSVPRASARPAARTTDTDSSTAFTPVSFPINVIDSPPSNTSESIRKFM 434
P Q+ P P + P T S +P F PP+ +
Sbjct: 411 PVQSSPPPVQQSPPPTPLTPPPVQSTPPPASPPPASPPPFSPPPATPPPATPPPA----- 575
Query: 435 EVRKEKVSALEEYYLTCPSPRRYPGPRPERLVDPDEPILANPIQEAD 481
AL L+ P P P P P +L P LA + +D
Sbjct: 576 ----TPPPALTPTPLSSP-PATTPAPAPAKL-KSKAPALAPVLSPSD 698
>TC76514 homologue to PIR|T09648|T09648 nucleolin homolog nuM1 - alfalfa,
partial (61%)
Length = 1288
Score = 38.9 bits (89), Expect = 0.007
Identities = 54/250 (21%), Positives = 85/250 (33%), Gaps = 38/250 (15%)
Frame = +3
Query: 186 PAPEFPSPPKKKTKKKMVLEESSEESD------------VPLVKKTKS--------KPDD 225
PAP +P KK KK E +SEESD P+ K S K D
Sbjct: 345 PAPSKKTPAKKGNVKKAQPETTSEESDSDSSSSDEEEVKKPVSKAVPSKNGSAPAKKVDT 524
Query: 226 GDDDDSEDGPPKKKQKKVRIVVKPTRVEPAAAVV-------------RRTEAPARVTRSS 272
+++DSE+ + K+ + V PA E PA S
Sbjct: 525 SEEEDSEESSDEDKKPAAKAVPSKNGSAPAKKAASDEEDTDESSDEDEEDEKPAAKAVPS 704
Query: 273 AHSSKPAL-----ASDDDLNLFDALPISALLQHSSNPLTPIPESQPAAQTTSPPHSPRSP 327
+ S PA +SD+D D + S N P ++ ++ S S
Sbjct: 705 KNGSVPAKKADTESSDEDSESSDEEDKKPAAKASKNVSAPTKKAASSSDEESDEESDEDE 884
Query: 328 FFQPSPTEAPLWNLLKNPTSRSEDPTSLLTIPYDPVSSEPIIHEQPEPNQTEPQPRTSDH 387
+P A + K +S S+D + D +P+ ++ + + SD
Sbjct: 885 DAKPVSKPAAVAKKSKKDSSDSDDEDDDSSSDED---KKPVAAKKEDKMNVDKDGSDSDQ 1055
Query: 388 SVPRASARPA 397
S + P+
Sbjct: 1056SEEESEDEPS 1085
Score = 30.4 bits (67), Expect = 2.7
Identities = 19/50 (38%), Positives = 26/50 (52%), Gaps = 8/50 (16%)
Frame = +3
Query: 205 EESSEESDVPLVKKTKSKPD---DGDDDD-----SEDGPPKKKQKKVRIV 246
+ SS+E P+ K + K + DG D D SED P K QKK++ V
Sbjct: 966 DSSSDEDKKPVAAKKEDKMNVDKDGSDSDQSEEESEDEPSKTPQKKIKDV 1115
Score = 29.3 bits (64), Expect = 5.9
Identities = 23/81 (28%), Positives = 36/81 (44%), Gaps = 6/81 (7%)
Frame = +3
Query: 659 EAIKEAKRKKDQAEE-AARQAAAQAEQARLEAERLEAEAEAERL-----APVVFTPAASA 712
E +K KK + EE AA+Q A + + + +E+E E+ AP TPA
Sbjct: 201 EEVKAVSAKKQKVEEVAAKQKALKVVKKEESSSEESSESEDEQPVVKAPAPSKKTPAKKG 380
Query: 713 SIPDPQAAQNVPSSSTQTSSS 733
++ Q S + +SSS
Sbjct: 381 NVKKAQPETTSEESDSDSSSS 443
>TC83458 homologue to PIR|T07796|T07796 DNA-directed RNA polymerase (EC
2.7.7.6) largest chain - soybean (fragment), partial
(63%)
Length = 1220
Score = 38.5 bits (88), Expect = 0.010
Identities = 30/103 (29%), Positives = 42/103 (40%)
Frame = +2
Query: 311 SQPAAQTTSPPHSPRSPFFQPSPTEAPLWNLLKNPTSRSEDPTSLLTIPYDPVSSEPIIH 370
+ P TSP +SP SP + PS + +PTS S PTS P P S
Sbjct: 902 TSPGYSPTSPGYSPTSPTYSPSSPGYSPTSPAYSPTSPSYSPTSPSYSPTSPSYSPTSPS 1081
Query: 371 EQPEPNQTEPQPRTSDHSVPRASARPAARTTDTDSSTAFTPVS 413
P + P TS P +SA + +S +++P S
Sbjct: 1082 YSPTSSSYSP---TSPAYSPTSSAYSPTSPAYSPTSPSYSPTS 1201
Score = 38.1 bits (87), Expect = 0.013
Identities = 34/106 (32%), Positives = 50/106 (47%), Gaps = 3/106 (2%)
Frame = +2
Query: 311 SQPAAQTTSPPHSPRSPFFQP-SPTEAPLWNLLKNPTSRSEDPTSLLTIPYDPVSSEPII 369
+ P TSP +SP SP + P SPT +P + +PTS + PTS P P S
Sbjct: 881 TSPGYSPTSPGYSPTSPGYSPTSPTYSPS-SPGYSPTSPAYSPTSPSYSPTSPSYSP--- 1048
Query: 370 HEQPEPNQTEP--QPRTSDHSVPRASARPAARTTDTDSSTAFTPVS 413
P + T P P +S +S P + A + + +S A++P S
Sbjct: 1049-TSPSYSPTSPSYSPTSSSYS-PTSPAYSPTSSAYSPTSPAYSPTS 1180
Score = 37.0 bits (84), Expect = 0.028
Identities = 36/126 (28%), Positives = 52/126 (40%), Gaps = 5/126 (3%)
Frame = +2
Query: 309 PESQPAAQTTSPPHSPRSPFFQP-SPTEAPLWNLLKNPTSRSEDPTSLLTIPYDP-VSSE 366
P S P +SP +SP SP + P SP +P + +PTS PTS P P S
Sbjct: 812 PTSSPGYSPSSPGYSPSSPGYSPTSPGYSPT-SPGYSPTSPGYSPTSPTYSPSSPGYSPT 988
Query: 367 PIIHEQPEPNQTEPQPRTSDHSVPRASARPAARTTDTD---SSTAFTPVSFPINVIDSPP 423
+ P+ + P S S + P+ T + +S A++P S +
Sbjct: 989 SPAYSPTSPSYSPTSPSYSPTSPSYSPTSPSYSPTSSSYSPTSPAYSPTSSAYSPTSPAY 1168
Query: 424 SNTSES 429
S TS S
Sbjct: 1169SPTSPS 1186
Score = 36.2 bits (82), Expect = 0.048
Identities = 29/74 (39%), Positives = 37/74 (49%), Gaps = 1/74 (1%)
Frame = +2
Query: 293 PISALLQHSSNPLTPIPESQPAAQTTSPPHSPRSPFFQP-SPTEAPLWNLLKNPTSRSED 351
P S SS +P + PA TSP +SP SP + P SP+ +P + +PTS S
Sbjct: 941 PTSPTYSPSSPGYSP---TSPAYSPTSPSYSPTSPSYSPTSPSYSPT-SPSYSPTSSSYS 1108
Query: 352 PTSLLTIPYDPVSS 365
PTS Y P SS
Sbjct: 1109 PTS---PAYSPTSS 1141
Score = 30.8 bits (68), Expect = 2.0
Identities = 23/67 (34%), Positives = 33/67 (48%), Gaps = 2/67 (2%)
Frame = +2
Query: 301 SSNPLTP-IPESQPAAQTTSPPHSPRSPFFQP-SPTEAPLWNLLKNPTSRSEDPTSLLTI 358
S +P +P + P+ TSP +SP S + P SP +P + +PTS + PTS
Sbjct: 1016 SYSPTSPSYSPTSPSYSPTSPSYSPTSSSYSPTSPAYSPT-SSAYSPTSPAYSPTSPSYS 1192
Query: 359 PYDPVSS 365
P P S
Sbjct: 1193 PTSPFYS 1213
Score = 30.4 bits (67), Expect = 2.7
Identities = 15/40 (37%), Positives = 22/40 (54%)
Frame = +2
Query: 293 PISALLQHSSNPLTPIPESQPAAQTTSPPHSPRSPFFQPS 332
P S +S+ +P + PA TSP +SP SPF+ P+
Sbjct: 1109 PTSPAYSPTSSAYSP---TSPAYSPTSPSYSPTSPFYSPT 1219
>BQ750979 similar to GP|13883467|gb PE_PGRS family protein {Mycobacterium
tuberculosis CDC1551}, partial (1%)
Length = 728
Score = 38.1 bits (87), Expect = 0.013
Identities = 22/78 (28%), Positives = 35/78 (44%)
Frame = +1
Query: 164 YVEDLRAQGFEIDLDDFVSRLPPAPEFPSPPKKKTKKKMVLEESSEESDVPLVKKTKSKP 223
+ ++ AQG ++ DDFV P+ P P+K +++ S K +SK
Sbjct: 391 WAQECEAQGSKVAEDDFVVSGLSKPQPPPKPQKAPASAQASKKNGAASQTSAAGKKRSKD 570
Query: 224 DDGDDDDSEDGPPKKKQK 241
DD+ + P KKQK
Sbjct: 571 VAQSDDEHDVKPQPKKQK 624
>TC81816 similar to GP|8096269|dbj|BAA95789.1 KED {Nicotiana tabacum},
partial (13%)
Length = 663
Score = 38.1 bits (87), Expect = 0.013
Identities = 21/49 (42%), Positives = 27/49 (54%), Gaps = 2/49 (4%)
Frame = +3
Query: 195 KKKTKKKMVLEESSEESDVPLVKKTKSKPDD--GDDDDSEDGPPKKKQK 241
KKK KK+ E+ ++ + KK K K D DDD+ EDG KKK K
Sbjct: 336 KKKEKKEKGKEDKDKDGEEKKSKKDKEKKKDKNEDDDEGEDGSKKKKNK 482
>AW689097 weakly similar to PIR|T47565|T47 hypothetical protein F24B22.20 -
Arabidopsis thaliana, partial (24%)
Length = 582
Score = 37.7 bits (86), Expect = 0.017
Identities = 18/39 (46%), Positives = 25/39 (63%), Gaps = 1/39 (2%)
Frame = -2
Query: 549 HSSNHQASPEPNLSIIPYTR-LQPTSLSECINIFYHEAS 586
H ++H + EP S++PY L PTSLS NI+ H+AS
Sbjct: 317 HXAHHSRTSEPGSSLVPYKPFLDPTSLSAPSNIYTHQAS 201
>BQ144502 weakly similar to GP|6523547|emb hydroxyproline-rich glycoprotein
DZ-HRGP {Volvox carteri f. nagariensis}, partial (39%)
Length = 1358
Score = 37.7 bits (86), Expect = 0.017
Identities = 58/240 (24%), Positives = 72/240 (29%), Gaps = 15/240 (6%)
Frame = -3
Query: 299 QHSSNPLTPIPESQPAAQTTSPPHSPRSPFFQPSPTEAPLWNLLKNPTSRSEDPTSLLTI 358
QH S P P +SPP P Q + AP P +R P+ +
Sbjct: 726 QHPSRAPRRHPPPPPPPPPSSPPGMGTRPSCQENWWAAP-------PGARPSRPSRYIPP 568
Query: 359 PYDPVSSEPIIHEQPEPNQT--------EPQPRTSDHSVPRASARPAARTTDTDSSTAFT 410
P P + P P+ T P PR P A RP R
Sbjct: 567 PTSPPPASPPAPVPPQSPSTIRGRAPPPAPPPRRPPARPPPAPQRPRPRRPPA------- 409
Query: 411 PVSFPINVIDSPPSNTSESIRKFMEVRKEKVSALEEYYLTCPSP--RRYPGPRPERLVDP 468
PPS R +VRK P+P +P PR P
Sbjct: 408 ----------RPPSGRRRGAR--AKVRKLPKGGTPGSRAPSPAPPSPTHPAPRRPWTGPP 265
Query: 469 DEPILANPIQEA-----DPLVQQAHPVPDQQEPIQPDPEPEQSVSNQSSVRSPHPLVETS 523
P +P Q A P + P P P+ P P P +S PHP TS
Sbjct: 264 PRPRPHSP*QAACGRGHPPRHRPPPPGPPPPPPLGPRPPPHPP---RSGPPRPHPPTRTS 94
Score = 33.5 bits (75), Expect = 0.31
Identities = 54/231 (23%), Positives = 70/231 (29%), Gaps = 11/231 (4%)
Frame = -1
Query: 292 LPISALLQHSSNPLTPIPESQPAAQTTSPPHSPRSPFFQPSPTEAPLWNLLKNPTSRSED 351
+P+ + + P P P P + +PP S +P P P A L+ P+ + D
Sbjct: 599 VPLVRVATYPRPPRPPPPVPPPPSPPRAPPQSEAAPPPPPPPRGARPRAPLRPPS--APD 426
Query: 352 PTSLLTIPYDPVSSEP---------IIHEQPEPNQTEPQPRTSDHSVPRASARPAARTTD 402
P P V+ P H PEP P PR P + PAA
Sbjct: 425 PAGHPRAPRQGVAVVPGPRSGNSPRAAHRGPEP---RPLPRP-----PPPTRPPAAPGPG 270
Query: 403 TDSSTAFTPVSFPINVIDSPPSNTSESIRKFMEVRKEKVSALEEYYLTCPSPRRYPGPRP 462
+ TP+S P PP+ T P PR P
Sbjct: 269 PPPAPGPTPLSKPPAAEGIPPA-------------------------TAPPPRARRLRPP 165
Query: 463 ERLVDPDEPILANPIQEADPLVQ--QAHPVPDQQEPIQPDPEPEQSVSNQS 511
P P A P P H P P P P P + N S
Sbjct: 164 SAPAPPPTPPAAAPPAPTPPHAPA*APHLPPPPPAPRPPRPPPFWYIINSS 12
Score = 28.9 bits (63), Expect = 7.7
Identities = 53/221 (23%), Positives = 65/221 (28%), Gaps = 6/221 (2%)
Frame = -2
Query: 304 PLTPIPESQPAAQTTS----PPHSPRSPFFQPSPTEAPLWNLLKNPTSRSEDPTSLLTIP 359
P P P S P + + P SP P P P P +R SL P
Sbjct: 850 PGPPDPASTPLDRARACCARRPPSPHRP--APPPQAHP---------ARERRTASLARTP 704
Query: 360 YDPVSSEPIIHEQPEPNQTEPQPRTSDHSV-PRASARPAARTTDTDSSTAFTPVSFPINV 418
P SS P P PR + S+ PR AR + S + TP
Sbjct: 703 PPPPSSPPPAPLLP--------PRNGNASLLPRKLVGSPARRSSLSSESLHTPAHL---- 560
Query: 419 IDSPPSNTSESIRKFMEVRKEKVSALEEYYLTCPSPRRYPGPRPERLVDPDEPILANPI- 477
+PP + P P P PRP P P A P
Sbjct: 559 --APPRQSPRPRPP-------------------PEPLHNPRPRPPPRPPPAAPARAPPSG 443
Query: 478 QEADPLVQQAHPVPDQQEPIQPDPEPEQSVSNQSSVRSPHP 518
A P P + P P E + V+SP P
Sbjct: 442 PPAPPTPPATRAPPVRASPWCPGQGQETPQGRHTGVQSPVP 320
>TC91646 weakly similar to GP|6523547|emb|CAB62280.1 hydroxyproline-rich
glycoprotein DZ-HRGP {Volvox carteri f. nagariensis},
partial (15%)
Length = 984
Score = 37.4 bits (85), Expect = 0.022
Identities = 33/105 (31%), Positives = 45/105 (42%), Gaps = 1/105 (0%)
Frame = +1
Query: 295 SALLQHSSNPLTPIPESQPAAQTTSPPHSPRSPFFQPSPTEAPLWNLLKNPTSRSEDPTS 354
S L++ S +P P S + P +P S PSPT +P P RS P
Sbjct: 232 SPLMEAPSPRPSPSPLSPLRFPSRLPLTTPPSLTTSPSPTASPPPRSPPRPPPRSP-PRQ 408
Query: 355 LLTIPYD-PVSSEPIIHEQPEPNQTEPQPRTSDHSVPRASARPAA 398
L++ P P S P PE ++T P PR P ++ PAA
Sbjct: 409 LMSPPRPLPRSPSPPPRNPPELSRTTPTPRPPMSPSPSPTSPPAA 543
Score = 32.3 bits (72), Expect = 0.70
Identities = 25/85 (29%), Positives = 40/85 (46%), Gaps = 5/85 (5%)
Frame = +3
Query: 334 TEAPLWNLLKNPTSRSEDPTSLLTIPYDPVSS--EPIIHEQPEPNQTEPQPRTSDHSVPR 391
TE P + P + +++PT T +P++S EP +PN T S VP
Sbjct: 363 TEEPTETATEEPATTTDEPTE--TSTEEPITSTEEPTGTVTDDPNTTASDESFSFTDVPT 536
Query: 392 AS---ARPAARTTDTDSSTAFTPVS 413
S + P+A TT T+++T V+
Sbjct: 537 GSDSTSVPSATTTGTETATPSATVA 611
Score = 28.9 bits (63), Expect = 7.7
Identities = 27/105 (25%), Positives = 37/105 (34%), Gaps = 1/105 (0%)
Frame = +1
Query: 323 SPRSPFFQPSPTEAPLWNLLKNPTSRSEDPTSLLTIPYDPVSSEPIIHEQPEPNQTEPQP 382
SP P P+ +PL L P SL T P S P +P P Q
Sbjct: 232 SPLMEAPSPRPSPSPLSPLRFPSRLPLTTPPSLTTSPSPTASPPPRSPPRPPPRSPPRQL 411
Query: 383 RTSDHSVPRASARPAARTTDTDSSTAF-TPVSFPINVIDSPPSNT 426
+ +PR+ + P + +T P P SPP+ T
Sbjct: 412 MSPPRPLPRSPSPPPRNPPELSRTTPTPRPPMSPSPSPTSPPAAT 546
>TC76445 similar to PIR|T07623|T07623 extensin homolog HRGP2 - soybean
(fragment), partial (76%)
Length = 821
Score = 37.4 bits (85), Expect = 0.022
Identities = 45/181 (24%), Positives = 56/181 (30%), Gaps = 5/181 (2%)
Frame = +2
Query: 309 PESQPAAQTTSPPH---SPRSPFFQPSPTEAPLWNLLKNPTSRSEDPTSLLTIPYDPVSS 365
P P SPP SP SP++ SP P S S P P P S
Sbjct: 2 PSPPPPYYYKSPPPPSPSPPSPYYYKSPP----------PPSPSPPPPYYYKSPPPPSPS 151
Query: 366 EP--IIHEQPEPNQTEPQPRTSDHSVPRASARPAARTTDTDSSTAFTPVSFPINVIDSPP 423
P ++ P P P P S P S P+S P N SPP
Sbjct: 152 PPPPYYYKSPPPPSPSPPPPYHYQSPPPPS-----------------PISHPPNYYKSPP 280
Query: 424 SNTSESIRKFMEVRKEKVSALEEYYLTCPSPRRYPGPRPERLVDPDEPILANPIQEADPL 483
+ + +Y++ P P + P P P PI P PL
Sbjct: 281 PPSPSPPPPY-------------HYVSPPPPVKSPPPPAYIYASPPPPIYN*PHNTKSPL 421
Query: 484 V 484
V
Sbjct: 422 V 424
>TC81858 similar to GP|18252179|gb|AAL61922.1 unknown protein {Arabidopsis
thaliana}, partial (9%)
Length = 748
Score = 37.4 bits (85), Expect = 0.022
Identities = 23/73 (31%), Positives = 32/73 (43%)
Frame = +1
Query: 371 EQPEPNQTEPQPRTSDHSVPRASARPAARTTDTDSSTAFTPVSFPINVIDSPPSNTSESI 430
E P + T + P+A+A P + TT S+A +P S P + SPPS T S
Sbjct: 136 EAPSTSPTAAPKASHAAPAPKATATPPSSTTTPPKSSATSPTSSPAPKVSSPPSPTPTSA 315
Query: 431 RKFMEVRKEKVSA 443
+E E A
Sbjct: 316 EAPVESPTESPPA 354
>AJ499220 similar to SP|Q09863|YAF Hypothetical protein C29E6.10c in
chromosome I. [Fission yeast] {Schizosaccharomyces
pombe}, partial (12%)
Length = 518
Score = 37.0 bits (84), Expect = 0.028
Identities = 30/81 (37%), Positives = 48/81 (59%), Gaps = 5/81 (6%)
Frame = -3
Query: 625 QLLHAEGSQSIEAAKQRLARRVALHELEQRNKL-----LEAIKEAKRKKDQAEEAARQAA 679
++L+A + + +Q+L L ELE+ N+L L+ +KE +RKK A+ A +
Sbjct: 315 RVLNAYREKVAQERQQKL-----LEELEEENRLKEERELKKLKEKERKK--AKNRALKQQ 157
Query: 680 AQAEQARLEAERLEAEAEAER 700
+ E+AR EAERL AE +A+R
Sbjct: 156 KEEERARREAERL-AEEKAQR 97
Score = 33.5 bits (75), Expect = 0.31
Identities = 19/66 (28%), Positives = 39/66 (58%), Gaps = 1/66 (1%)
Frame = -3
Query: 633 QSIEAAKQRLARRVALHELEQRNKLLEAIKEAKR-KKDQAEEAARQAAAQAEQARLEAER 691
+ ++ K++ ++ L+Q+ + A +EA+R +++A+ A R+ + E+ R E ER
Sbjct: 222 RELKKLKEKERKKAKNRALKQQKEEERARREAERLAEEKAQRAEREKKLEEERKRREEER 43
Query: 692 LEAEAE 697
L+ EAE
Sbjct: 42 LKKEAE 25
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.312 0.128 0.366
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 22,602,607
Number of Sequences: 36976
Number of extensions: 371920
Number of successful extensions: 3702
Number of sequences better than 10.0: 226
Number of HSP's better than 10.0 without gapping: 2956
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 3523
length of query: 766
length of database: 9,014,727
effective HSP length: 104
effective length of query: 662
effective length of database: 5,169,223
effective search space: 3422025626
effective search space used: 3422025626
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.8 bits)
S2: 62 (28.5 bits)
Lotus: description of TM0014.15


