

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0011a.2
(391 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
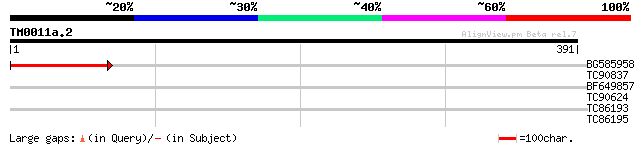
BG585958 weakly similar to GP|22830897|dbj ORF-B {Oryza sativa (... 97 8e-21
TC90837 weakly similar to GP|22202752|dbj|BAC07409. hypothetical... 31 0.72
BF649857 30 2.1
TC90624 similar to GP|6721160|gb|AAF26788.1| hypothetical protei... 29 3.6
TC86193 similar to GP|16974678|gb|AAL32439.1 SH3 domain-containi... 28 7.9
TC86195 similar to GP|16974678|gb|AAL32439.1 SH3 domain-containi... 28 7.9
>BG585958 weakly similar to GP|22830897|dbj ORF-B {Oryza sativa (japonica
cultivar-group)}, partial (13%)
Length = 769
Score = 97.4 bits (241), Expect = 8e-21
Identities = 47/71 (66%), Positives = 56/71 (78%)
Frame = +2
Query: 1 SSNCRRQRSHIERNREEGHVRLWNDYFSTNPVYTEEQFQRRYRMRKHMFLRIVEAISNTD 60
SS R QR +IERNREEG+ RL+NDYFS PVY +E F+RRYRM KH+FLRIVEA+ D
Sbjct: 458 SSRPRSQRRNIERNREEGYKRLFNDYFSEAPVYMDE*FRRRYRMHKHVFLRIVEALGQHD 637
Query: 61 DYFQMRNDATG 71
+YFQ+ DATG
Sbjct: 638 EYFQLTVDATG 670
>TC90837 weakly similar to GP|22202752|dbj|BAC07409. hypothetical
protein~similar to Oryza sativa chromosome10
OSJNBa0079H13.17, partial (7%)
Length = 596
Score = 31.2 bits (69), Expect = 0.72
Identities = 24/91 (26%), Positives = 39/91 (42%), Gaps = 11/91 (12%)
Frame = +1
Query: 242 YYLADEIYPEWATFVKTISM-----------PQGEKRKLFAQHQEGARKDVERAFGVL*S 290
YYL D+ YP+ ++ P ++ F + R +ER+FGVL
Sbjct: 10 YYLVDKGYPDKEGYMVPYPRIRYHQSQFEHEPPTNAQEAFNRAHSSLRSCIERSFGVLKK 189
Query: 291 RFAIVRGPARAWHLHILKDIIYACIILHNMI 321
R+ I+ + + + D+I A LHN I
Sbjct: 190 RWKILNKMPQ-FSVKTQIDVIIAAFALHNYI 279
>BF649857
Length = 601
Score = 29.6 bits (65), Expect = 2.1
Identities = 10/29 (34%), Positives = 16/29 (54%), Gaps = 2/29 (6%)
Frame = -3
Query: 153 GSIDCMHWI--WENGPTAWKGQYCRGDHG 179
G++ C HW W+ G + G +C D+G
Sbjct: 194 GNVICYHWFVAWQRGHRMYLGNFCNMDYG 108
>TC90624 similar to GP|6721160|gb|AAF26788.1| hypothetical protein
{Arabidopsis thaliana}, partial (7%)
Length = 455
Score = 28.9 bits (63), Expect = 3.6
Identities = 12/20 (60%), Positives = 15/20 (75%)
Frame = +2
Query: 86 RMLAYGSSTDSVDEYVCIGE 105
R L GSSTD ++ +VCIGE
Sbjct: 392 RELGAGSSTDFIETFVCIGE 451
>TC86193 similar to GP|16974678|gb|AAL32439.1 SH3 domain-containing protein 2
{Arabidopsis thaliana}, partial (97%)
Length = 1311
Score = 27.7 bits (60), Expect = 7.9
Identities = 12/36 (33%), Positives = 22/36 (60%)
Frame = +1
Query: 230 QFTLNGTTYNMGYYLADEIYPEWATFVKTISMPQGE 265
Q T NG+T +MGY+L + ++P A +++ G+
Sbjct: 991 QTTHNGSTDSMGYFLGEVLFPYSAVSEVELNLSVGD 1098
>TC86195 similar to GP|16974678|gb|AAL32439.1 SH3 domain-containing protein
2 {Arabidopsis thaliana}, partial (58%)
Length = 848
Score = 27.7 bits (60), Expect = 7.9
Identities = 12/36 (33%), Positives = 22/36 (60%)
Frame = +2
Query: 230 QFTLNGTTYNMGYYLADEIYPEWATFVKTISMPQGE 265
Q T NG+T +MGY+L + ++P A +++ G+
Sbjct: 440 QTTHNGSTDSMGYFLGEVLFPYSAVSEVELNLSVGD 547
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.323 0.138 0.434
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 12,056,332
Number of Sequences: 36976
Number of extensions: 162871
Number of successful extensions: 803
Number of sequences better than 10.0: 12
Number of HSP's better than 10.0 without gapping: 799
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 803
length of query: 391
length of database: 9,014,727
effective HSP length: 98
effective length of query: 293
effective length of database: 5,391,079
effective search space: 1579586147
effective search space used: 1579586147
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.5 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (22.0 bits)
S2: 59 (27.3 bits)
Lotus: description of TM0011a.2


