

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0010.6
(493 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
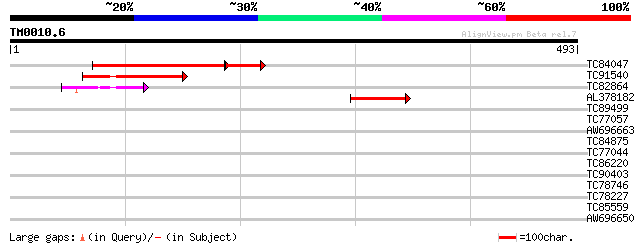
Score E
Sequences producing significant alignments: (bits) Value
TC84047 homologue to GP|3695063|gb|AAC62626.1| rac GTPase activa... 157 2e-50
TC91540 similar to GP|9757821|dbj|BAB08339.1 rac GTPase activati... 103 2e-22
TC82864 similar to GP|3695061|gb|AAC62625.1| rac GTPase activati... 64 1e-10
AL378182 similar to GP|3695063|gb|A rac GTPase activating protei... 50 2e-06
TC89499 similar to GP|20197852|gb|AAM15282.1 hypothetical protei... 32 0.56
TC77057 similar to GP|15982817|gb|AAL09756.1 At2g20760/F5H14.27 ... 31 0.95
AW696663 weakly similar to GP|17827193|dbj ORF1 {TT virus}, part... 30 2.1
TC84875 similar to GP|4508080|gb|AAD21424.1| 69873 {Arabidopsis ... 29 3.6
TC77044 homologue to GP|4103324|gb|AAD01737.1| GDP-mannose pyrop... 29 3.6
TC86220 similar to GP|7339705|dbj|BAA92910.1 ESTs D23839(R0339) ... 29 4.7
TC90403 weakly similar to GP|7339705|dbj|BAA92910.1 ESTs D23839(... 29 4.7
TC78746 similar to GP|22137234|gb|AAM91462.1 AT5g39360/MUL8_40 {... 29 4.7
TC78227 similar to GP|10176883|dbj|BAB10113. contains similarity... 29 4.7
TC85559 similar to PIR|T07012|T07012 acetyl-CoA carboxylase (EC ... 28 8.0
AW696650 similar to PIR|D96594|D965 unknown protein 71207-66119... 28 8.0
>TC84047 homologue to GP|3695063|gb|AAC62626.1| rac GTPase activating
protein 3 {Lotus japonicus}, partial (34%)
Length = 630
Score = 157 bits (396), Expect(2) = 2e-50
Identities = 77/119 (64%), Positives = 92/119 (76%)
Frame = +3
Query: 73 RKSLIGSCSTSPRDSGALSSSSMEIGWPSNVRHVAHVTFDRFHGFLGLPVEFEPEVPRRP 132
+KS++ SP D + MEIGWP+NV+HV HVTFDRF+GFLGLP+E E VP
Sbjct: 162 KKSMVACSVESPDDVISAVHHPMEIGWPTNVKHVNHVTFDRFNGFLGLPLELEVHVPAPV 341
Query: 133 PSASTSVFGVSTESMQLSFDARGNSVPTILLLMQRHLYARGGLQAEGIFRINAENSQEE 191
PSAS SVFGVS ESMQ S+D++GNSVPTILLLMQ LY++GGL+AEGIFRIN EN +EE
Sbjct: 342 PSASVSVFGVSAESMQCSYDSKGNSVPTILLLMQERLYSQGGLKAEGIFRINPENGEEE 518
Score = 60.8 bits (146), Expect(2) = 2e-50
Identities = 25/35 (71%), Positives = 31/35 (88%)
Frame = +1
Query: 188 SQEELVREQLNRGVVPNGVDVHCLAGLIKAWFREL 222
++ ++REQLN G+VPN +DVHCLAGLIKAWFREL
Sbjct: 508 AKRNILREQLNSGIVPNDIDVHCLAGLIKAWFREL 612
>TC91540 similar to GP|9757821|dbj|BAB08339.1 rac GTPase activating protein
{Arabidopsis thaliana}, partial (13%)
Length = 645
Score = 103 bits (256), Expect = 2e-22
Identities = 53/91 (58%), Positives = 67/91 (73%)
Frame = +1
Query: 64 LLTLLIATFRKSLIGSCSTSPRDSGALSSSSMEIGWPSNVRHVAHVTFDRFHGFLGLPVE 123
L LL+ FRKSL S+ +D +M+I P+NVRHVAHVTFDRF+GFLGLP E
Sbjct: 388 LFELLVTLFRKSLFPFKSSGNKDL-----CNMDISPPTNVRHVAHVTFDRFNGFLGLPDE 552
Query: 124 FEPEVPRRPPSASTSVFGVSTESMQLSFDAR 154
FEP+ PRRPPSAS +VF VST+S+Q S+D++
Sbjct: 553 FEPDFPRRPPSASATVFEVSTKSIQFSYDSK 645
>TC82864 similar to GP|3695061|gb|AAC62625.1| rac GTPase activating protein
2 {Lotus japonicus}, partial (14%)
Length = 583
Score = 63.9 bits (154), Expect = 1e-10
Identities = 38/101 (37%), Positives = 53/101 (51%), Gaps = 26/101 (25%)
Frame = +2
Query: 46 EEDEEEEKEKD--------------------------REGDQLSLLTLLIATFRKSLIGS 79
EEDEEEE E D +Q ++L +++A +KS++ +
Sbjct: 299 EEDEEEELESDDNDNDDDDEIYSSNNASRQFTKGSNNNHQNQFAILDIVMAALKKSIV-T 475
Query: 80 CSTSPRDSGALSSSSMEIGWPSNVRHVAHVTFDRFHGFLGL 120
CS D SS++I WP+ VRHV+HVTFDRF+GFLGL
Sbjct: 476 CSVEREDV-----SSLDISWPTEVRHVSHVTFDRFNGFLGL 583
>AL378182 similar to GP|3695063|gb|A rac GTPase activating protein 3 {Lotus
japonicus}, partial (12%)
Length = 218
Score = 50.4 bits (119), Expect = 2e-06
Identities = 28/53 (52%), Positives = 37/53 (68%), Gaps = 1/53 (1%)
Frame = +3
Query: 297 PLTALMYAVQVMSFLKTLVVKTLREREESIVKS-NPVPNLNSFDDDGHQSDSQ 348
P+TALM+AVQVM+ LKTL++KTLREREE+ +P+ SF + DSQ
Sbjct: 3 PVTALMHAVQVMNLLKTLIMKTLREREETATGGYSPMSFRTSFRRSEDEYDSQ 161
>TC89499 similar to GP|20197852|gb|AAM15282.1 hypothetical protein
{Arabidopsis thaliana}, partial (21%)
Length = 1002
Score = 32.0 bits (71), Expect = 0.56
Identities = 16/51 (31%), Positives = 28/51 (54%), Gaps = 2/51 (3%)
Frame = +1
Query: 64 LLTLLIATFRKSLIGSCSTSP--RDSGALSSSSMEIGWPSNVRHVAHVTFD 112
++ ++I T + G C + + MEIG+P++V+HVAHV +D
Sbjct: 331 IIGIVIFTMATTFKGICKSLKYITQMFVVKEREMEIGYPTDVKHVAHVGWD 483
>TC77057 similar to GP|15982817|gb|AAL09756.1 At2g20760/F5H14.27
{Arabidopsis thaliana}, partial (49%)
Length = 1280
Score = 31.2 bits (69), Expect = 0.95
Identities = 14/30 (46%), Positives = 19/30 (62%)
Frame = +1
Query: 349 VLPKDGSENGNDCSDEDTVFVSAEPSQPSP 378
V ++G++NGN D+D VFVS P P P
Sbjct: 313 VTVENGNDNGNGYGDDDGVFVSDGPILPPP 402
>AW696663 weakly similar to GP|17827193|dbj ORF1 {TT virus}, partial (2%)
Length = 660
Score = 30.0 bits (66), Expect = 2.1
Identities = 15/43 (34%), Positives = 26/43 (59%)
Frame = -1
Query: 17 DGSHQTALINNSSSVEEGVLQLQTHLDLVEEDEEEEKEKDREG 59
+ +H T ++ +S + Q L+ EE+EEEE+E++REG
Sbjct: 222 NSNHPTLCVHAETSYHQ-----QQQLEEEEEEEEEEEEEEREG 109
>TC84875 similar to GP|4508080|gb|AAD21424.1| 69873 {Arabidopsis thaliana},
partial (21%)
Length = 692
Score = 29.3 bits (64), Expect = 3.6
Identities = 10/18 (55%), Positives = 15/18 (82%)
Frame = +1
Query: 95 MEIGWPSNVRHVAHVTFD 112
MEIG+P++V+HV H+ D
Sbjct: 427 MEIGYPTDVKHVTHIGLD 480
>TC77044 homologue to GP|4103324|gb|AAD01737.1| GDP-mannose pyrophosphorylase
{Solanum tuberosum}, complete
Length = 1631
Score = 29.3 bits (64), Expect = 3.6
Identities = 21/97 (21%), Positives = 38/97 (38%), Gaps = 4/97 (4%)
Frame = -3
Query: 310 FLKTLVVKTLREREESIVKSNPVPNLNSFDDDGHQSDSQVLPKDGSENGNDCSDEDTVFV 369
+L+ + + L + + N + N+N F D H S LP G N + ++F+
Sbjct: 1257 WLQNICLNLLVG*NHTTIAVNLITNMNIFSKDSHILHSSPLPDSGMPPNNTAGNA-SMFL 1081
Query: 370 SAEPSQ----PSPTHHTEDGCETESGSETSPTPAENF 402
+ S P + T+S ++ T NF
Sbjct: 1080 DPDSSHYCAARKPNTRLNNAARTDSNIRSNETSLTNF 970
>TC86220 similar to GP|7339705|dbj|BAA92910.1 ESTs D23839(R0339)
AU082696(E61918) correspond to a region of the predicted
gene.~Similar to, partial (8%)
Length = 658
Score = 28.9 bits (63), Expect = 4.7
Identities = 24/89 (26%), Positives = 38/89 (41%), Gaps = 9/89 (10%)
Frame = +2
Query: 315 VVKTLREREESIVKSNPVPNLNSFDDDGHQSDSQVLPKDGSENGNDCSDEDTVFVSAEPS 374
V+ L+E EE + N D +S+S+V + NG+D D+D+V V S
Sbjct: 311 VITFLKESEEGSAEQGKEVN------DNDESESKVKDDEEKGNGDDNEDKDSVSVEDSQS 472
Query: 375 QPSPTHHT---------EDGCETESGSET 394
+ ++ T E E E G E+
Sbjct: 473 EKVNSNPTIMRAVMLVFERVAEDEGGDES 559
>TC90403 weakly similar to GP|7339705|dbj|BAA92910.1 ESTs D23839(R0339)
AU082696(E61918) correspond to a region of the predicted
gene.~Similar to, partial (18%)
Length = 1244
Score = 28.9 bits (63), Expect = 4.7
Identities = 23/91 (25%), Positives = 39/91 (42%), Gaps = 9/91 (9%)
Frame = +2
Query: 313 TLVVKTLREREESIVKSNPVPNLNSFDDDGHQSDSQVLPKDGSENGNDCSDEDTVFVSAE 372
+ ++ L+E EE + N D +S+S+V + NG+D D+D+V V
Sbjct: 71 SFIITFLKESEEGSAEQGKEVN------DNDESESKVKDDEEKGNGDDNEDKDSVSVEDS 232
Query: 373 PSQPSPTHHT---------EDGCETESGSET 394
S+ ++ T E E E G E+
Sbjct: 233 QSEKVNSNPTIMRAVMLVFERVAEDEGGDES 325
>TC78746 similar to GP|22137234|gb|AAM91462.1 AT5g39360/MUL8_40 {Arabidopsis
thaliana}, partial (97%)
Length = 1222
Score = 28.9 bits (63), Expect = 4.7
Identities = 12/37 (32%), Positives = 23/37 (61%)
Frame = -3
Query: 25 INNSSSVEEGVLQLQTHLDLVEEDEEEEKEKDREGDQ 61
++ + V+E +THL L + E E+E++REG++
Sbjct: 128 LSRGNLVQEKHRNEETHLKLRNAEREREREREREGER 18
>TC78227 similar to GP|10176883|dbj|BAB10113. contains similarity to unknown
protein~gb|AAD10691.1~gene_id:K9L2.3 {Arabidopsis
thaliana}, partial (27%)
Length = 1707
Score = 28.9 bits (63), Expect = 4.7
Identities = 13/40 (32%), Positives = 25/40 (62%)
Frame = +1
Query: 25 INNSSSVEEGVLQLQTHLDLVEEDEEEEKEKDREGDQLSL 64
INN+S + + Q H EEDE+EE+++D + + +++
Sbjct: 175 INNTSQPNDELESDQRHSSENEEDEDEEEDEDDDEEYVAI 294
>TC85559 similar to PIR|T07012|T07012 acetyl-CoA carboxylase (EC 6.4.1.2) -
potato chloroplast, partial (66%)
Length = 5011
Score = 28.1 bits (61), Expect = 8.0
Identities = 18/69 (26%), Positives = 33/69 (47%)
Frame = +3
Query: 333 PNLNSFDDDGHQSDSQVLPKDGSENGNDCSDEDTVFVSAEPSQPSPTHHTEDGCETESGS 392
P+ NS +D+ S++ L + SE G + DE + + +PS ++ +E+
Sbjct: 3537 PSENSLEDEDEPSENS-LEDEPSEKGLEDEDEPSEKGLEDEDEPSEKGLEDEDEPSENSL 3713
Query: 393 ETSPTPAEN 401
E P+EN
Sbjct: 3714 EDEDEPSEN 3740
>AW696650 similar to PIR|D96594|D965 unknown protein 71207-66119 [imported]
- Arabidopsis thaliana, partial (1%)
Length = 663
Score = 28.1 bits (61), Expect = 8.0
Identities = 26/74 (35%), Positives = 34/74 (45%), Gaps = 8/74 (10%)
Frame = +1
Query: 5 LQLPSPSRRCSNDGSHQTALINNSSSVEEGVLQLQTHLDLVEEDEE--------EEKEKD 56
L P SR N +H NS + + GVLQLQ LV++ E EEK K+
Sbjct: 49 LLTPISSRVSRNSSNHSP----NSKTADAGVLQLQNQ-QLVQQTEVQKQAIQDLEEKTKE 213
Query: 57 REGDQLSLLTLLIA 70
+ Q S +LIA
Sbjct: 214 LKERQNSYDDILIA 255
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.313 0.130 0.370
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 14,086,238
Number of Sequences: 36976
Number of extensions: 192190
Number of successful extensions: 992
Number of sequences better than 10.0: 30
Number of HSP's better than 10.0 without gapping: 955
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 983
length of query: 493
length of database: 9,014,727
effective HSP length: 100
effective length of query: 393
effective length of database: 5,317,127
effective search space: 2089630911
effective search space used: 2089630911
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.9 bits)
S2: 60 (27.7 bits)
Lotus: description of TM0010.6


