

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0010.3.5
(80 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
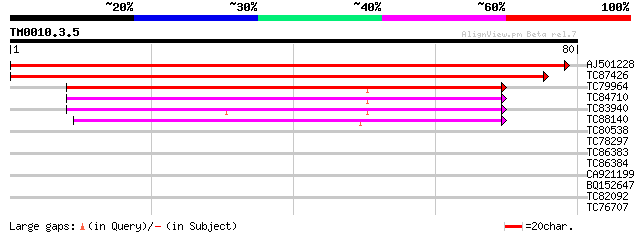
AJ501228 homologue to SP|O82221|RUXG_A Probable small nuclear ri... 150 9e-38
TC87426 homologue to PIR|S22826|S22826 small nuclear ribonucleop... 148 4e-37
TC79964 homologue to PIR|B84453|B84453 probable snRNP splicing f... 59 3e-10
TC84710 homologue to PIR|B84453|B84453 probable snRNP splicing f... 54 8e-09
TC83940 homologue to PIR|B84453|B84453 probable snRNP splicing f... 45 4e-06
TC88140 similar to PIR|T47805|T47805 U6 snRNA-associated Sm-like... 39 5e-04
TC80538 homologue to PIR|D96797|D96797 Sm-like protein [imported... 36 0.002
TC78297 homologue to GP|9758886|dbj|BAB09440.1 U6 snRNA-associat... 36 0.003
TC86383 similar to GP|21593343|gb|AAM65292.1 small nuclear ribon... 35 0.004
TC86384 similar to GP|21593343|gb|AAM65292.1 small nuclear ribon... 35 0.004
CA921199 similar to PIR|T51950|T51 DNA-directed RNA polymerase (... 32 0.044
BQ152647 homologue to PIR|B84453|B8445 probable snRNP splicing f... 28 0.49
TC82092 similar to PIR|H86164|H86164 hypothetical protein AAC721... 27 1.1
TC76707 similar to SP|Q07202|CORA_MEDSA Cold and drought-regulat... 25 7.1
>AJ501228 homologue to SP|O82221|RUXG_A Probable small nuclear
ribonucleoprotein G (snRNP-G) (Sm protein G) (Sm-G)
(SmG). [Mouse-ear cress], partial (98%)
Length = 658
Score = 150 bits (379), Expect = 9e-38
Identities = 74/79 (93%), Positives = 77/79 (96%)
Frame = +2
Query: 1 MSRSGQPPDLKKYMDKNLQIKLNANRMIVGTLRGYDQFMNMVVDNTVEVNGNEKNDIGMV 60
MSRSGQPPDLKKYMDK LQIKLNANRMIVGTLRG+DQFMN+VVDNTVEVNGNEKNDIGMV
Sbjct: 191 MSRSGQPPDLKKYMDKQLQIKLNANRMIVGTLRGFDQFMNLVVDNTVEVNGNEKNDIGMV 370
Query: 61 VIRGNSVVTVEALEPVTRT 79
VIRGNSVVTVEALEPV R+
Sbjct: 371 VIRGNSVVTVEALEPVNRS 427
>TC87426 homologue to PIR|S22826|S22826 small nuclear ribonucleoprotein E
homolog - alfalfa, complete
Length = 609
Score = 148 bits (373), Expect = 4e-37
Identities = 73/76 (96%), Positives = 75/76 (98%)
Frame = +2
Query: 1 MSRSGQPPDLKKYMDKNLQIKLNANRMIVGTLRGYDQFMNMVVDNTVEVNGNEKNDIGMV 60
MSRSGQPPDLKKYMDK LQIKLNANRMIVGTLRG+DQFMN+VVDNTVEVNGNEKNDIGMV
Sbjct: 71 MSRSGQPPDLKKYMDKQLQIKLNANRMIVGTLRGFDQFMNLVVDNTVEVNGNEKNDIGMV 250
Query: 61 VIRGNSVVTVEALEPV 76
VIRGNSVVTVEALEPV
Sbjct: 251 VIRGNSVVTVEALEPV 298
>TC79964 homologue to PIR|B84453|B84453 probable snRNP splicing factor
[imported] - Arabidopsis thaliana, partial (93%)
Length = 607
Score = 58.9 bits (141), Expect = 3e-10
Identities = 27/71 (38%), Positives = 43/71 (60%), Gaps = 9/71 (12%)
Frame = +1
Query: 9 DLKKYMDKNLQIKLNANRMIVGTLRGYDQFMNMVVDNTVEV---------NGNEKNDIGM 59
DL K++DK +Q+KL R + GTL+GYDQ +N+V+D VE ++ +G+
Sbjct: 91 DLAKFVDKGVQVKLTGGRQVTGTLKGYDQLLNLVLDEAVEFLRDPDDPLKTTDQTRSLGL 270
Query: 60 VVIRGNSVVTV 70
+V RG +V+ V
Sbjct: 271 IVCRGTAVMLV 303
>TC84710 homologue to PIR|B84453|B84453 probable snRNP splicing factor
[imported] - Arabidopsis thaliana, partial (93%)
Length = 543
Score = 54.3 bits (129), Expect = 8e-09
Identities = 25/71 (35%), Positives = 41/71 (57%), Gaps = 9/71 (12%)
Frame = +3
Query: 9 DLKKYMDKNLQIKLNANRMIVGTLRGYDQFMNMVVDNTVEV---------NGNEKNDIGM 59
DL K++DK +Q+KL R + GTL+GYDQ +N+ +D VE ++ +G+
Sbjct: 45 DLAKFVDKGVQVKLTGGRQVTGTLKGYDQLLNLFLDEAVEFLRDPDDPLKTTDQTRSLGL 224
Query: 60 VVIRGNSVVTV 70
+ RG +V+ V
Sbjct: 225 IGCRGTAVMLV 257
>TC83940 homologue to PIR|B84453|B84453 probable snRNP splicing factor
[imported] - Arabidopsis thaliana, partial (93%)
Length = 632
Score = 45.4 bits (106), Expect = 4e-06
Identities = 28/99 (28%), Positives = 44/99 (44%), Gaps = 37/99 (37%)
Frame = +1
Query: 9 DLKKYMDKNLQIKLNANRMIV----------------------------GTLRGYDQFMN 40
DL K++DK +Q+KL R +V GTL+GYDQ +N
Sbjct: 94 DLAKFVDKGVQVKLTGGRQVVLVLSLLEV*VWFQNVTGVISLIHR*KLTGTLKGYDQLLN 273
Query: 41 MVVDNTVEV---------NGNEKNDIGMVVIRGNSVVTV 70
+V+D VE ++ +G++V RG +V+ V
Sbjct: 274 LVLDEAVEFLRDPDDPLKTTDQTRSLGLIVCRGTAVMLV 390
>TC88140 similar to PIR|T47805|T47805 U6 snRNA-associated Sm-like protein -
Arabidopsis thaliana, complete
Length = 649
Score = 38.5 bits (88), Expect = 5e-04
Identities = 23/62 (37%), Positives = 34/62 (54%), Gaps = 1/62 (1%)
Frame = +2
Query: 10 LKKYMDKNLQIKLNANRMIVGTLRGYDQFMNMVVDNTVE-VNGNEKNDIGMVVIRGNSVV 68
LK + + +KLN+ G L D +MN+ ++ T E VNG KN G IRGN+V+
Sbjct: 161 LKSIRGRPVVVKLNSGVDYRGILACLDGYMNIAMEQTEEYVNGQLKNKYGDAFIRGNNVL 340
Query: 69 TV 70
+
Sbjct: 341 YI 346
>TC80538 homologue to PIR|D96797|D96797 Sm-like protein [imported] -
Arabidopsis thaliana, partial (95%)
Length = 554
Score = 36.2 bits (82), Expect = 0.002
Identities = 25/90 (27%), Positives = 46/90 (50%), Gaps = 15/90 (16%)
Frame = +2
Query: 6 QPPDLKKY-MDKNLQIKLNANRMIVGTLRGYDQFMNMVVDN------TVEVNG------- 51
+P DL + +D+ + +KL ++R + G L YDQ +NM++ + TVE++
Sbjct: 74 EPLDLIRLSLDERIYVKLRSDRELRGKLHAYDQHLNMILGDVEEIVTTVEIDDETYEEIV 253
Query: 52 -NEKNDIGMVVIRGNSVVTVEALEPVTRTS 80
K + + +RG+ V+ V P RT+
Sbjct: 254 RTTKRTVPFLFVRGDGVILV---SPPLRTA 334
>TC78297 homologue to GP|9758886|dbj|BAB09440.1 U6 snRNA-associated Sm-like
protein-like {Arabidopsis thaliana}, partial (95%)
Length = 693
Score = 35.8 bits (81), Expect = 0.003
Identities = 14/62 (22%), Positives = 35/62 (55%), Gaps = 4/62 (6%)
Frame = +1
Query: 10 LKKYMDKNLQIKLNANRMIVGTLRGYDQFMNMVVDNTVE----VNGNEKNDIGMVVIRGN 65
+ + + + + + ++ +VGTLRG+D ++NMV+++ E G + +++ GN
Sbjct: 205 IDRCIGSKIWVIMKGDKELVGTLRGFDVYVNMVLEDVTEYEITAEGRRITKLDQILLNGN 384
Query: 66 SV 67
++
Sbjct: 385 NI 390
>TC86383 similar to GP|21593343|gb|AAM65292.1 small nuclear
ribonucleoprotein homolog {Arabidopsis thaliana},
complete
Length = 684
Score = 35.4 bits (80), Expect = 0.004
Identities = 14/42 (33%), Positives = 31/42 (73%), Gaps = 2/42 (4%)
Frame = +2
Query: 28 IVGTLRGYDQFMNMVVDNTVEVNGNEKN--DIGMVVIRGNSV 67
I G + G+D++MN+V+D+ EVN +K+ +G ++++G+++
Sbjct: 191 IEGRIIGFDEYMNLVLDDAEEVNVKKKSKKTLGRILLKGDNI 316
>TC86384 similar to GP|21593343|gb|AAM65292.1 small nuclear
ribonucleoprotein homolog {Arabidopsis thaliana},
complete
Length = 658
Score = 35.4 bits (80), Expect = 0.004
Identities = 14/42 (33%), Positives = 31/42 (73%), Gaps = 2/42 (4%)
Frame = +1
Query: 28 IVGTLRGYDQFMNMVVDNTVEVNGNEKN--DIGMVVIRGNSV 67
I G + G+D++MN+V+D+ EVN +K+ +G ++++G+++
Sbjct: 223 IEGRIIGFDEYMNLVLDDAEEVNVKKKSKKTLGRILLKGDNI 348
>CA921199 similar to PIR|T51950|T51 DNA-directed RNA polymerase (EC 2.7.7.6)
23K chain [imported] - Arabidopsis thaliana, partial
(49%)
Length = 756
Score = 32.0 bits (71), Expect = 0.044
Identities = 11/39 (28%), Positives = 26/39 (66%)
Frame = +2
Query: 10 LKKYMDKNLQIKLNANRMIVGTLRGYDQFMNMVVDNTVE 48
+ + + + + + ++ +VGTLRG+D ++NMV+++ E
Sbjct: 617 IDRCIGSKIWVIMKGDKELVGTLRGFDVYVNMVLEDVTE 733
>BQ152647 homologue to PIR|B84453|B8445 probable snRNP splicing factor
[imported] - Arabidopsis thaliana, partial (73%)
Length = 495
Score = 28.5 bits (62), Expect = 0.49
Identities = 10/19 (52%), Positives = 14/19 (73%)
Frame = +3
Query: 19 QIKLNANRMIVGTLRGYDQ 37
++KL R + GTL+GYDQ
Sbjct: 30 RVKLTGGRQVTGTLKGYDQ 86
>TC82092 similar to PIR|H86164|H86164 hypothetical protein AAC72111.1
[imported] - Arabidopsis thaliana, partial (80%)
Length = 311
Score = 27.3 bits (59), Expect = 1.1
Identities = 11/36 (30%), Positives = 22/36 (60%)
Frame = +1
Query: 11 KKYMDKNLQIKLNANRMIVGTLRGYDQFMNMVVDNT 46
K + + + ++L + I GTL DQ++N+ ++NT
Sbjct: 49 KDLVGREVTVELKNDLAIRGTLHSVDQYLNIKLENT 156
>TC76707 similar to SP|Q07202|CORA_MEDSA Cold and drought-regulated protein
CORA. [Alfalfa] {Medicago sativa}, partial (87%)
Length = 780
Score = 24.6 bits (52), Expect = 7.1
Identities = 16/60 (26%), Positives = 28/60 (46%)
Frame = +2
Query: 18 LQIKLNANRMIVGTLRGYDQFMNMVVDNTVEVNGNEKNDIGMVVIRGNSVVTVEALEPVT 77
+ +++ ++V T M +VV TVEV ++ MVV+ + V +E VT
Sbjct: 197 IMVEVITTAVVVTTTVEVVTTMVVVVTTTVEVATTMVEEVTMVVVTTTAEVVTTTVEVVT 376
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.314 0.133 0.360
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 1,496,639
Number of Sequences: 36976
Number of extensions: 10508
Number of successful extensions: 48
Number of sequences better than 10.0: 28
Number of HSP's better than 10.0 without gapping: 47
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 47
length of query: 80
length of database: 9,014,727
effective HSP length: 56
effective length of query: 24
effective length of database: 6,944,071
effective search space: 166657704
effective search space used: 166657704
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (22.0 bits)
S2: 51 (24.3 bits)
Lotus: description of TM0010.3.5


