

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0010.16
(405 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
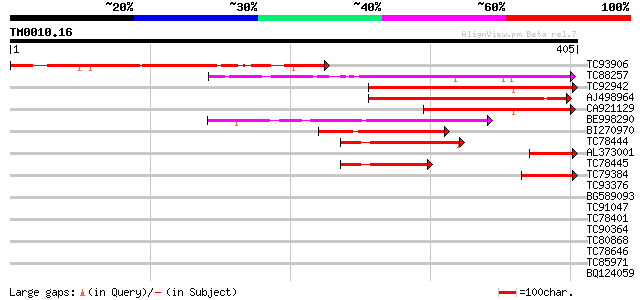
Score E
Sequences producing significant alignments: (bits) Value
TC93906 similar to GP|21741691|emb|CAD40593. oj000126_13.15 {Ory... 231 5e-61
TC88257 weakly similar to PIR|G96668|G96668 protein F1N19.7 [imp... 166 2e-41
TC92942 similar to GP|15451042|gb|AAK96792.1 putative protein {A... 145 2e-35
AJ498964 weakly similar to GP|10177031|db emb|CAB72177.1~gene_id... 125 3e-29
CA921129 similar to GP|14532460|gb AT5g13810/MAC12_24 {Arabidops... 114 5e-26
BE998290 weakly similar to GP|14532460|gb| AT5g13810/MAC12_24 {A... 96 2e-20
BI270970 weakly similar to PIR|T48164|T481 hypothetical protein ... 87 1e-17
TC78444 similar to GP|14532460|gb|AAK63958.1 AT5g13810/MAC12_24 ... 73 2e-13
AL373001 69 4e-12
TC78445 similar to GP|14532460|gb|AAK63958.1 AT5g13810/MAC12_24 ... 60 1e-09
TC79384 similar to GP|14532460|gb|AAK63958.1 AT5g13810/MAC12_24 ... 51 9e-07
TC93376 37 0.014
BG589093 similar to PIR|F71608|F716 hypothetical protein PFB0700... 37 0.018
TC91047 similar to GP|22136346|gb|AAM91251.1 unknown protein {Ar... 36 0.023
TC78401 similar to GP|6735386|emb|CAB69043.1 glutaredoxin {Arabi... 35 0.052
TC90364 similar to GP|5442102|gb|AAD43253.1| peptide methionine ... 35 0.068
TC80868 similar to PIR|H86410|H86410 protein F3M18.8 [imported] ... 34 0.12
TC78646 similar to GP|11994338|dbj|BAB02297. contains similarity... 32 0.34
TC85971 similar to GP|19424073|gb|AAL87350.1 unknown protein {Ar... 32 0.44
BQ124059 similar to PIR|T05236|T052 hypothetical protein F18A5.6... 30 1.3
>TC93906 similar to GP|21741691|emb|CAD40593. oj000126_13.15 {Oryza sativa},
partial (8%)
Length = 707
Score = 231 bits (588), Expect = 5e-61
Identities = 146/255 (57%), Positives = 168/255 (65%), Gaps = 27/255 (10%)
Frame = +2
Query: 1 MGCVSSKLIRKDIKQEHVIIDNCGGGGRYLSHVVSLTSSTYGALKLDK-----DNDQPVI 55
MGCVSSK I+K+I QE Y++HVVSLTSSTYGALKLD D+ ++
Sbjct: 23 MGCVSSKHIKKEINQEK---------NPYINHVVSLTSSTYGALKLDNNSNNNDSSNSIV 175
Query: 56 A----------------AKPREEPEAITTINAWELMEGLEDGVPISNQPKKSPKSSSPFL 99
+ + P ++PE + INAWELMEGLE+GVPISN PKKSPK S+PFL
Sbjct: 176 SETETQTESKTEPKSNPSPPHKDPETVFNINAWELMEGLEEGVPISNFPKKSPK-SAPFL 352
Query: 100 RGFMNSDTRSPLKFLNQLGSPK-TVKKSTGKENKVQVNGVRTGGVRRLDYSNSPKGILK- 157
RGFM SDTRSPLKFL+Q GSPK T+KK GKENKVQV + GGVRRLDY SPKGILK
Sbjct: 353 RGFMASDTRSPLKFLSQYGSPKSTLKKPLGKENKVQVTNMVRGGVRRLDY--SPKGILKS 526
Query: 158 PSNLSPNASKNSGIPAKGSPICARRKSFGGNEKDTLQCSSRRKS----QSPLFDPELVAS 213
+N SP KN KGSP ARR SFG S RKS SPLFDPE++AS
Sbjct: 527 TTNSSP---KN----LKGSPFSARRNSFGN--------ESVRKSPGSVPSPLFDPEIIAS 661
Query: 214 YEKELSEEEEQVKRM 228
YEKELSEEEEQ+KR+
Sbjct: 662 YEKELSEEEEQIKRI 706
>TC88257 weakly similar to PIR|G96668|G96668 protein F1N19.7 [imported] -
Arabidopsis thaliana, partial (29%)
Length = 1283
Score = 166 bits (419), Expect = 2e-41
Identities = 100/271 (36%), Positives = 154/271 (55%), Gaps = 9/271 (3%)
Frame = +2
Query: 143 VRRLDYSNSPKGILKPSNLSPNASKNSGIPAKGSPICARRKSFGGNEKDTLQCSSRRKSQ 202
+ LD +SPK K + SP + P K + G+ + S K +
Sbjct: 398 LEELDAKSSPKK--KNESQSPKPPQPQAQPTKNKVVDV------GSVRTLKPSLSSSKLK 553
Query: 203 SPLFDPELVASYEKELSEEEEQVKRMVWATPKTRRARKSLDSLSLIQMFEKKCPPGGENS 262
+F + +KE E+E +R+ R R+ D LS + +KCPP G ++
Sbjct: 554 DNIFIKRDILEKQKE--EKESNFERI-------ERLRR--DPLS---RYPEKCPPNGNDA 691
Query: 263 VVIYTTTLRGIRKTFEDCNKVRAIIESYCVQMRERDVSMDSGFKEELRKLMGAEQ----- 317
VVIYTT+LRG+RKTF++CN+ R ++E++ V ERDV++ F +E+++L+ E+
Sbjct: 692 VVIYTTSLRGVRKTFDNCNQARDLLENHRVVFDERDVALHGEFLKEVKELLLTEEELGVG 871
Query: 318 VKVPVVFVKGRLVGGVDEVVKLEDEEKLGVLLEG--IPKAVG--GCEGCGGVRFVMCVEC 373
V +P VFVKGR +GG+DE+++L + +LG +L + + VG GC GCGGVRFV C +C
Sbjct: 872 VVLPRVFVKGRYLGGLDELMELNETGRLGRILNATRVERGVGRQGCVGCGGVRFVPCWDC 1051
Query: 374 NGSCKVLDEDKKKTVKCGKCNENGIMQCPIC 404
GSCK++ D ++C KCNENG++ CP C
Sbjct: 1052GGSCKLVTSDAGGEIRCPKCNENGLVHCPTC 1144
>TC92942 similar to GP|15451042|gb|AAK96792.1 putative protein {Arabidopsis
thaliana}, partial (38%)
Length = 650
Score = 145 bits (367), Expect = 2e-35
Identities = 69/152 (45%), Positives = 101/152 (66%), Gaps = 3/152 (1%)
Frame = +1
Query: 257 PGGENSVVIYTTTLRGIRKTFEDCNKVRAIIESYCVQMRERDVSMDSGFKEELRKLMGAE 316
PG EN +V+Y T+LRGIRKT+EDC VR I+ Y V + ERD+SMDS +++EL+ +G +
Sbjct: 73 PGTENRIVVYCTSLRGIRKTYEDCCAVRMILRGYRVAVDERDISMDSSYRKELQNALGGK 252
Query: 317 Q-VKVPVVFVKGRLVGGVDEVVKLEDEEKLGVLLEGIPKAVGG--CEGCGGVRFVMCVEC 373
V +P VF++G+ VG D++ +L + +L +L+G P C+ CG RFV C C
Sbjct: 253 SVVTLPQVFIRGKHVGNADDLKQLNESGELARMLKGFPTQDPWFVCDKCGDARFVPCNNC 432
Query: 374 NGSCKVLDEDKKKTVKCGKCNENGIMQCPICC 405
NGS KV +E++ K +C CNENG+++C CC
Sbjct: 433 NGSRKVFEEEQGKLKRCVHCNENGLIRCSSCC 528
>AJ498964 weakly similar to GP|10177031|db
emb|CAB72177.1~gene_id:MQJ2.15~similar to unknown
protein {Arabidopsis thaliana}, partial (50%)
Length = 764
Score = 125 bits (314), Expect = 3e-29
Identities = 68/147 (46%), Positives = 93/147 (63%), Gaps = 2/147 (1%)
Frame = +2
Query: 257 PGGENSVVIYTTTLRGIRKTFEDCNKVRAIIESYCVQMRERDVSMDSGFKEELRKLMGAE 316
P + VVIY T+LR +R T+EDC VR+I+ + + + ERDVSMDSGF ELR + G +
Sbjct: 323 PSSQQRVVIYFTSLRVVRTTYEDCKTVRSILRGFKIHLDERDVSMDSGFLSELRLVTGHK 502
Query: 317 Q-VKVPVVFVKGRLVGGVDEVVKLEDEEKLGVLLEGIPKAVG-GCEGCGGVRFVMCVECN 374
+K+P VF+ GR +GG EV L + +L LLEG+P A C CG RFV+C EC+
Sbjct: 503 TGLKLPRVFINGRYIGGAQEVTWLHENGELKKLLEGLPVADSLVCHVCGDHRFVLCGECS 682
Query: 375 GSCKVLDEDKKKTVKCGKCNENGIMQC 401
G+ KV E K C CNE+G+++C
Sbjct: 683 GARKVYAE-KGGFKTCMDCNESGLIRC 760
>CA921129 similar to GP|14532460|gb AT5g13810/MAC12_24 {Arabidopsis
thaliana}, partial (40%)
Length = 729
Score = 114 bits (286), Expect = 5e-26
Identities = 52/111 (46%), Positives = 75/111 (66%), Gaps = 2/111 (1%)
Frame = -2
Query: 296 ERDVSMDSGFKEELRKLMGAEQVKVPVVFVKGRLVGGVDEVVKLEDEEKLGVLLEGIPKA 355
ERDVSMD F++EL+ +MG E V +P VFV+G+ +GG D + L + +L +LEG P+
Sbjct: 680 ERDVSMDRAFRKELQSVMGEENVTLPQVFVRGKYIGGADVIKSLFETGELKRILEGFPRM 501
Query: 356 VGG--CEGCGGVRFVMCVECNGSCKVLDEDKKKTVKCGKCNENGIMQCPIC 404
G CE CG RF+ C C+GS K+ DED+ + +C +CNENG+++CP C
Sbjct: 500 KPGFVCESCGDARFIPCENCSGSRKLFDEDEGLSKRCLECNENGLVRCPCC 348
>BE998290 weakly similar to GP|14532460|gb| AT5g13810/MAC12_24 {Arabidopsis
thaliana}, partial (19%)
Length = 584
Score = 96.3 bits (238), Expect = 2e-20
Identities = 68/206 (33%), Positives = 104/206 (50%), Gaps = 2/206 (0%)
Frame = +1
Query: 142 GVRRLDYSNSPKGILKPSN--LSPNASKNSGIPAKGSPICARRKSFGGNEKDTLQCSSRR 199
G L + + PK ++ P N SP+ +NS P P+ R S CSS +
Sbjct: 1 GRNTLCFLSPPKSVMWPPNWLASPSRHRNSPPPPSPLPLHCRTHSIN------FTCSSFK 162
Query: 200 KSQSPLFDPELVASYEKELSEEEEQVKRMVWATPKTRRARKSLDSLSLIQMFEKKCPPGG 259
++ + + E L + +R+ +T R S + Q PPG
Sbjct: 163 DIETLIQEKP-----EPNLPKSPSLFRRIRISTTFLRALSASRTTAPPPQTIS--LPPGL 321
Query: 260 ENSVVIYTTTLRGIRKTFEDCNKVRAIIESYCVQMRERDVSMDSGFKEELRKLMGAEQVK 319
EN VVIY T+LR IR+T+ DC VR+I+ ++ V ERDVS+D F++EL +++ + V
Sbjct: 322 ENRVVIYFTSLRVIRRTYNDCRAVRSILRNFRVITDERDVSIDDRFRDELNEILNRKNVT 501
Query: 320 VPVVFVKGRLVGGVDEVVKLEDEEKL 345
+P VFV G +GGVDEV +L + +L
Sbjct: 502 LPRVFVGGVYIGGVDEVKQLHESGEL 579
>BI270970 weakly similar to PIR|T48164|T481 hypothetical protein T10O8.130 -
Arabidopsis thaliana, partial (15%)
Length = 654
Score = 87.0 bits (214), Expect = 1e-17
Identities = 44/94 (46%), Positives = 60/94 (63%)
Frame = +2
Query: 221 EEEQVKRMVWATPKTRRARKSLDSLSLIQMFEKKCPPGGENSVVIYTTTLRGIRKTFEDC 280
EE Q AT K K SL + FE+KCPPGG NS+++YTT+LRGIRKTF+DC
Sbjct: 371 EELQEFNFKLATTKQVACIKGYPSL---KDFEEKCPPGGSNSIILYTTSLRGIRKTFQDC 541
Query: 281 NKVRAIIESYCVQMRERDVSMDSGFKEELRKLMG 314
N + ++ S V ERDVS+D +++EL ++G
Sbjct: 542 NTIHFLLRSLRVMYHERDVSLDLEYRQELXNILG 643
>TC78444 similar to GP|14532460|gb|AAK63958.1 AT5g13810/MAC12_24
{Arabidopsis thaliana}, partial (26%)
Length = 708
Score = 72.8 bits (177), Expect = 2e-13
Identities = 38/92 (41%), Positives = 62/92 (67%), Gaps = 3/92 (3%)
Frame = +3
Query: 237 RARKSLDSLSLIQMFEKKCPPGGENSVVIYTTTLRGIRKTFEDCNKVRAIIESYCVQMRE 296
++ KS+DS+ +I++ G E+ +V+Y T+LRGIR+T+EDC VR I+ + V + E
Sbjct: 450 KSSKSIDSVPVIKLL------GTEDRIVVYFTSLRGIRRTYEDCYAVRMILRGFRVWVDE 611
Query: 297 RDVSMDSGFKEELRKLMGAEQVK---VPVVFV 325
RDVSMD +++EL +MG + +K +P VF+
Sbjct: 612 RDVSMDICYRKELMSVMGEKSMKNVTLPQVFI 707
>AL373001
Length = 416
Score = 68.6 bits (166), Expect = 4e-12
Identities = 27/34 (79%), Positives = 31/34 (90%)
Frame = +2
Query: 372 ECNGSCKVLDEDKKKTVKCGKCNENGIMQCPICC 405
ECNGSCKVLDE +KKTVKCG CNENGI++C +CC
Sbjct: 5 ECNGSCKVLDEKQKKTVKCGYCNENGIIRCSLCC 106
>TC78445 similar to GP|14532460|gb|AAK63958.1 AT5g13810/MAC12_24
{Arabidopsis thaliana}, partial (19%)
Length = 733
Score = 60.5 bits (145), Expect = 1e-09
Identities = 30/66 (45%), Positives = 46/66 (69%)
Frame = +1
Query: 237 RARKSLDSLSLIQMFEKKCPPGGENSVVIYTTTLRGIRKTFEDCNKVRAIIESYCVQMRE 296
++ KS+DS+ +I++ G E+ +V+Y T+LRGIR+T+EDC VR I+ + V + E
Sbjct: 544 KSSKSIDSVPVIKLL------GTEDRIVVYFTSLRGIRRTYEDCYAVRMILRGFKVWVDE 705
Query: 297 RDVSMD 302
RDVSMD
Sbjct: 706 RDVSMD 723
>TC79384 similar to GP|14532460|gb|AAK63958.1 AT5g13810/MAC12_24
{Arabidopsis thaliana}, partial (14%)
Length = 873
Score = 50.8 bits (120), Expect = 9e-07
Identities = 20/40 (50%), Positives = 27/40 (67%)
Frame = +3
Query: 366 RFVMCVECNGSCKVLDEDKKKTVKCGKCNENGIMQCPICC 405
RF+ C C GS K+ DED+ +C CNENG+++CP CC
Sbjct: 6 RFLPCGNCCGSKKIFDEDEGLLKRCLVCNENGLIRCPNCC 125
>TC93376
Length = 450
Score = 37.0 bits (84), Expect = 0.014
Identities = 18/69 (26%), Positives = 27/69 (39%), Gaps = 19/69 (27%)
Frame = +2
Query: 355 AVGGCEGCGGVRFVMCVECNGSCKVLD-------------------EDKKKTVKCGKCNE 395
++G C+ CG VR C +C G+ + + E + K + C KC
Sbjct: 110 SLGLCKRCGNVRRTACAKCKGTGTIKEGGLLSFNLVDDLYETIGNRESQVKKIACDKCQA 289
Query: 396 NGIMQCPIC 404
G CP C
Sbjct: 290 KGYFPCPEC 316
>BG589093 similar to PIR|F71608|F716 hypothetical protein PFB0700c - malaria
parasite (Plasmodium falciparum), partial (2%)
Length = 657
Score = 36.6 bits (83), Expect = 0.018
Identities = 23/58 (39%), Positives = 27/58 (45%)
Frame = +1
Query: 257 PGGENSVVIYTTTLRGIRKTFEDCNKVRAIIESYCVQMRERDVSMDSGFKEELRKLMG 314
PG E VVIY T+L K R+RDVSMDS F EL ++MG
Sbjct: 448 PGTEQRVVIYFTSLARGSTNLRRLQKRTRNSTRLPRSSRQRDVSMDSSFLTELNRIMG 621
>TC91047 similar to GP|22136346|gb|AAM91251.1 unknown protein {Arabidopsis
thaliana}, partial (45%)
Length = 1034
Score = 36.2 bits (82), Expect = 0.023
Identities = 16/53 (30%), Positives = 25/53 (46%), Gaps = 6/53 (11%)
Frame = +2
Query: 358 GCEGCGGVRFVMCVECNGSCKVLDEDKKKT------VKCGKCNENGIMQCPIC 404
GC CGG ++C C G+ K ++K+ +C C G + CP+C
Sbjct: 308 GCRNCGGSGNIICDMCGGTGKWKALNRKRAQDVYEFTECPNCYGRGKLVCPVC 466
>TC78401 similar to GP|6735386|emb|CAB69043.1 glutaredoxin {Arabidopsis
thaliana}, partial (68%)
Length = 706
Score = 35.0 bits (79), Expect = 0.052
Identities = 28/93 (30%), Positives = 46/93 (49%), Gaps = 4/93 (4%)
Frame = +3
Query: 261 NSVVIYTTTL----RGIRKTFEDCNKVRAIIESYCVQMRERDVSMDSGFKEELRKLMGAE 316
N + I++ T R + F++ N+V Y V++ ERD S ++ L ++G
Sbjct: 174 NKIAIFSKTYCPYCRRAKAVFKELNQV-----PYVVELDERDDG--SKIQDVLVNIVGKR 332
Query: 317 QVKVPVVFVKGRLVGGVDEVVKLEDEEKLGVLL 349
V P VF+ G+ +GG DE V+ + L LL
Sbjct: 333 TV--PQVFINGKHLGGSDETVEAYESGLLAKLL 425
>TC90364 similar to GP|5442102|gb|AAD43253.1| peptide methionine sulfoxide
reductase {Gracilaria gracilis}, partial (12%)
Length = 473
Score = 34.7 bits (78), Expect = 0.068
Identities = 21/73 (28%), Positives = 39/73 (52%), Gaps = 3/73 (4%)
Frame = +1
Query: 280 CNKVRAIIESYCVQ---MRERDVSMDSGFKEELRKLMGAEQVKVPVVFVKGRLVGGVDEV 336
C K +A ++ + V+ + D S S +E L++ Q VP +F+KG+ VGG D++
Sbjct: 130 CKKAKATLKEFNVEPGVVELDDDSDGSAIQEYLKEK--TSQSTVPNIFIKGQHVGGCDDL 303
Query: 337 VKLEDEEKLGVLL 349
+ +D L ++
Sbjct: 304 LAAKDNGSLSKMI 342
>TC80868 similar to PIR|H86410|H86410 protein F3M18.8 [imported] -
Arabidopsis thaliana, partial (52%)
Length = 812
Score = 33.9 bits (76), Expect = 0.12
Identities = 15/45 (33%), Positives = 27/45 (59%)
Frame = +3
Query: 293 QMRERDVSMDSGFKEELRKLMGAEQVKVPVVFVKGRLVGGVDEVV 337
++ E+D + G +EL + E+V+ P VF+ G L GG+D ++
Sbjct: 354 EVEEKD---EVGLVKELESIANEEKVQFPAVFIGGNLFGGLDRIM 479
>TC78646 similar to GP|11994338|dbj|BAB02297. contains similarity to
thioredoxin~gene_id:MSJ11.6 {Arabidopsis thaliana},
partial (68%)
Length = 807
Score = 32.3 bits (72), Expect = 0.34
Identities = 30/121 (24%), Positives = 55/121 (44%), Gaps = 3/121 (2%)
Frame = +1
Query: 235 TRRARKSLDSLSLIQMFEKKCPPGGENSVVIYTTTLRGIRKTFEDC---NKVRAIIESYC 291
T + S SLS I + K +N V+IY ++G+ F C + +++ Y
Sbjct: 169 TNKVENSGTSLSSIIEQDVK-----DNPVMIY---MKGV-PDFPQCGFSSLAVKVLKQYD 321
Query: 292 VQMRERDVSMDSGFKEELRKLMGAEQVKVPVVFVKGRLVGGVDEVVKLEDEEKLGVLLEG 351
V + R++ D K+ ++ + P VF+KG +GG D V+ + +L L+
Sbjct: 322 VPLSARNILQDPEVKDAVKAF--SHWPTFPQVFIKGEFIGGSDIVLSMHQSGELKEKLKD 495
Query: 352 I 352
+
Sbjct: 496 V 498
>TC85971 similar to GP|19424073|gb|AAL87350.1 unknown protein {Arabidopsis
thaliana}, partial (77%)
Length = 1540
Score = 32.0 bits (71), Expect = 0.44
Identities = 19/57 (33%), Positives = 25/57 (43%), Gaps = 2/57 (3%)
Frame = +3
Query: 350 EGIP--KAVGGCEGCGGVRFVMCVECNGSCKVLDEDKKKTVKCGKCNENGIMQCPIC 404
E IP + + C GC G ++C CN DE+ G EN + QCP C
Sbjct: 618 EKIPHSEVIEKCSGCAGRGGIVCATCNA-----DEEP------GHYKENQVTQCPSC 755
>BQ124059 similar to PIR|T05236|T052 hypothetical protein F18A5.60 -
Arabidopsis thaliana, partial (16%)
Length = 579
Score = 30.4 bits (67), Expect = 1.3
Identities = 15/51 (29%), Positives = 24/51 (46%), Gaps = 5/51 (9%)
Frame = +3
Query: 359 CEGCGGVRFVMCVECNGSCKVLDEDK-----KKTVKCGKCNENGIMQCPIC 404
C C G+ ++C EC+GS + E + + KC C G + C +C
Sbjct: 291 CLQCSGLGVLLCTECDGSGEPNIEPQFMEWVGEDTKCPYCEGLGHITCDLC 443
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.314 0.134 0.391
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 11,795,939
Number of Sequences: 36976
Number of extensions: 161101
Number of successful extensions: 729
Number of sequences better than 10.0: 57
Number of HSP's better than 10.0 without gapping: 704
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 712
length of query: 405
length of database: 9,014,727
effective HSP length: 98
effective length of query: 307
effective length of database: 5,391,079
effective search space: 1655061253
effective search space used: 1655061253
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (22.0 bits)
S2: 59 (27.3 bits)
Lotus: description of TM0010.16


