

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
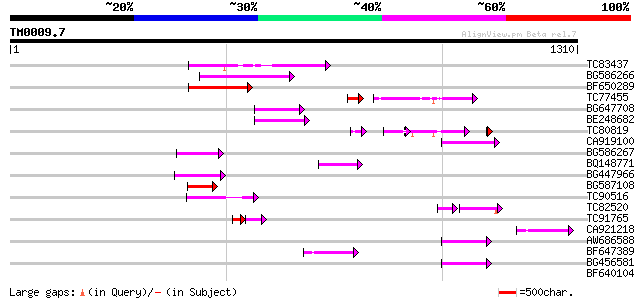
Query= TM0009.7
(1310 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC83437 weakly similar to PIR|D86384|D86384 unknown protein [imp... 174 2e-43
BG586266 similar to GP|7267666|em RNA-directed DNA polymerase-li... 161 1e-39
BF650289 weakly similar to GP|9049283|dbj| orf129a {Beta vulgari... 114 2e-25
TC77455 similar to GP|22335695|dbj|BAC10549. nine-cis-epoxycarot... 86 8e-22
BG647708 weakly similar to GP|13786450|gb| putative reverse tran... 84 3e-16
BE248682 similar to GP|18568269|gb putative gag-pol polyprotein ... 80 4e-15
TC80819 weakly similar to GP|10140689|gb|AAG13524.1 putative non... 72 4e-13
CA919100 homologue to PIR|G90291|G902 endoglucanase precursor [i... 70 7e-12
BG586267 weakly similar to PIR|A84888|A8 hypothetical protein At... 65 2e-10
BQ148771 65 2e-10
BG447966 weakly similar to PIR|G96509|G96 protein F27F5.21 [impo... 64 5e-10
BG587108 similar to PIR|G96509|G9 protein F27F5.21 [imported] - ... 62 1e-09
TC90516 59 1e-08
TC82520 46 2e-08
TC91765 weakly similar to GP|19881779|gb|AAM01180.1 Putative ret... 44 6e-08
CA921218 56 8e-08
AW686588 54 3e-07
BF647389 54 3e-07
BG456581 46 1e-04
BF640104 42 0.002
>TC83437 weakly similar to PIR|D86384|D86384 unknown protein [imported] -
Arabidopsis thaliana, partial (6%)
Length = 951
Score = 174 bits (442), Expect = 2e-43
Identities = 110/339 (32%), Positives = 170/339 (49%), Gaps = 11/339 (3%)
Frame = +2
Query: 413 DFFSSGRISRGMNAAFITLIPKKENASSLNHYRPISLIRSSYKLIAKVLATRLQKVMPKL 472
+F + ++ +G+N+ FI LIPK +N LN +RPISL+ S YK++ K+LA RL+ V+ +
Sbjct: 17 EFHRNRKLFKGINSTFIALIPKVDNPQRLNDFRPISLVGSLYKILGKLLANRLRVVIGSV 196
Query: 473 LSQNQFAFTKERQIADCILLAN-----------EVADFLSKKSEGGLMLKVDLAKAYDQV 521
+S Q AF K RQI + + L ++ L + + L + L V
Sbjct: 197 ISDAQSAFVKNRQILEMVFL*QMRLWMRLRN*RKIFCCLRWILKRLITLSIGLIWILF*V 376
Query: 522 DWEFLINMLHEMNFGDKWISWMKSCISSASLAILVNGSPTDFFSIEKGLRQGDPLSPLLF 581
FL+ W W+K C+S+A+ ++LVNGSPT+ + K L Q + F
Sbjct: 377 GMSFLV----------LWRKWIKECVSTATTSVLVNGSPTNV--LMKSLVQTQLFTRYSF 520
Query: 582 NLCLNGLSCLLNQFLGEENYCGVRISRNLTINHLLFADDTLLFCENNELQLETLCNVLIT 641
+ + ++HL FA+DTLL N + L L+
Sbjct: 521 G-----------------------VVNPVVVSHLQFANDTLLLETKNWANIRALRAALVI 631
Query: 642 FLFASGLKVNNQKSELIGCNVQTVEVERLAHSFGWSVGTLPISYLGAPLGGNPRRIVFWN 701
F SGLKVN KS L+ N+ + A W VG +P YLG P+ GN RR+ FW
Sbjct: 632 F*AMSGLKVNFHKSGLVCVNIAPSWLSEAASVLSWKVGKVPFLYLGMPIEGNSRRLSFWE 811
Query: 702 PMLEKLKCKTRSYNSKFMSLSGRLVITKSVLCSIPLFLM 740
P++ ++K + +NS+F+S GRLV+ KSVL S+ ++ +
Sbjct: 812 PIVNRIKARLTGWNSRFLSFGGRLVLLKSVLTSLSVYAL 928
>BG586266 similar to GP|7267666|em RNA-directed DNA polymerase-like protein
{Arabidopsis thaliana}, partial (18%)
Length = 789
Score = 161 bits (408), Expect = 1e-39
Identities = 86/224 (38%), Positives = 133/224 (58%), Gaps = 4/224 (1%)
Frame = -3
Query: 438 ASSLNHYRPISLIRSSYKLIAKVLATRLQKVMPKLLSQNQFAFTKERQIADCILLANEVA 497
A ++ YR I+ + YK+IAK+L+ R+Q ++ ++S +Q AF R I+D +L+ +++
Sbjct: 787 AKRVSEYRTIAPCNTQYKIIAKILSKRMQPLLRSIISPSQSAFVPGRAISDNVLITHKIL 608
Query: 498 DFLSK---KSEGGLMLKVDLAKAYDQVDWEFLINMLHEMNFGDKWISWMKSCISSASLAI 554
+L + K + +K D+ KAYD++ W FL +L + F WISW+ C+S+ S +
Sbjct: 607 HYLRQSGAKKHVSMAVKTDMTKAYDRIAWNFLREVLTRLGFHGIWISWIMECVSTVSYSF 428
Query: 555 LVNGSPTDFFSIEKGLRQGDPLSPLLFNLCLNGLSCLLNQFLGEENYCGVRISRNL-TIN 613
L+NG P +GLRQGDPLSP LF LC LS L Q L + GV+++RN IN
Sbjct: 427 LINGGPQGRVLPSRGLRQGDPLSPYLFILCTEVLSGLCQQALRKGTLPGVKVARNCPPIN 248
Query: 614 HLLFADDTLLFCENNELQLETLCNVLITFLFASGLKVNNQKSEL 657
HLLFADDT+ F ++N L +++ + ASG +N KS +
Sbjct: 247 HLLFADDTMFFGKSNASSCAILLSIMDKYRAASGRCIN*TKSAI 116
>BF650289 weakly similar to GP|9049283|dbj| orf129a {Beta vulgaris}, partial
(69%)
Length = 616
Score = 114 bits (286), Expect = 2e-25
Identities = 55/150 (36%), Positives = 98/150 (64%), Gaps = 1/150 (0%)
Frame = +3
Query: 413 DFFSSGRISRGMNAAFITLIPKKENASSLNHYRPISLIRSSYKLIAKVLATRLQKVMPKL 472
DFF +G + + +N ++TL+PK+ N +S+ ++RPI+ YK+I+K+L +R+Q V+ +
Sbjct: 159 DFFKTGFMPKIINCTYVTLLPKEVNVTSVKNFRPIACCSVIYKIISKILTSRMQGVLNSV 338
Query: 473 LSQNQFAFTKERQIADCILLANEVADFLSKKS-EGGLMLKVDLAKAYDQVDWEFLINMLH 531
+S+NQ AF K R I D I+L++E+ S+K M+K+DL KAYD +W F+ +++
Sbjct: 339 VSENQSAFVKGRVIFDNIILSHELVKSYSRKGISPRCMVKIDLXKAYDSXEWPFIKHLML 518
Query: 532 EMNFGDKWISWMKSCISSASLAILVNGSPT 561
E+ F K+++W+ + +++AS NG T
Sbjct: 519 ELGFPYKFVNWVMAXLTTASYTFNXNGDLT 608
>TC77455 similar to GP|22335695|dbj|BAC10549. nine-cis-epoxycarotenoid
dioxygenase1 {Pisum sativum}, partial (43%)
Length = 1865
Score = 85.5 bits (210), Expect(2) = 8e-22
Identities = 70/256 (27%), Positives = 112/256 (43%), Gaps = 15/256 (5%)
Frame = -2
Query: 840 PAMGSRAIQDIWRVLNEDHSLSAGFKENLLISIGDGSRVKFWRDPWVSNVALCHLFPRLY 899
PA SR +D+ + E+ F L+ +GDG FW+D W S+V L FPR +
Sbjct: 790 PAYASRWWKDLMSL--EEVGRVRWFPRELIRKVGDGRSSFFWKDAWDSSVPLRESFPRAF 617
Query: 900 AMSENKEARIAEVGTYSRGKWGWSLTFRG-QFFAWEEELYQQFLASINHIIPAAVGGDQL 958
+ ++ + W L +R + F WE+E + L + ++ D
Sbjct: 616 FPYRLLKMGCGDLWDMNAEGVRWRLYWRRLELFEWEKERLLELLGRLEGVV-LRYWADIW 440
Query: 959 IWCANQTGLFSVKELCYWL-------------EDHLFTHQMWQVPKSIAKLLPPKVALFF 1005
+W ++ G+FSV CY+L E+ + ++W KS A P KV F
Sbjct: 439 VWKPDKEGVFSVNS-CYFLLQNLRLLEDRLSYEEEVIFRELW---KSKA---PAKVLAFS 281
Query: 1006 WQAQNDKIATRANL-LARGIDLNGSTQCLFCNAFDESCNHIFLHCHSIWSLWCVIVNRER 1064
W D+I T NL R + + S +C+FC DE+ H+FLHC I + ++
Sbjct: 280 WTLFLDRIPTMVNLGKRRLLRVEDSKRCVFCGCQDETVVHLFLHCDVISKV*REVMRWLN 101
Query: 1065 FLWVMPASLADLILEW 1080
F + P +L + W
Sbjct: 100 FNLISPPNLLIHAICW 53
Score = 37.7 bits (86), Expect(2) = 8e-22
Identities = 18/38 (47%), Positives = 23/38 (60%)
Frame = -3
Query: 780 CLNRKAGGLGLGFIEWRNRGLLLKWAWRFGREINSLWR 817
CL R GGLG+ I N LL KW WR ++ +SLW+
Sbjct: 969 CLPRCKGGLGVRDIRLVNVSLLAKWWWRLLQDQSSLWK 856
>BG647708 weakly similar to GP|13786450|gb| putative reverse transcriptase
{Oryza sativa}, partial (9%)
Length = 708
Score = 84.3 bits (207), Expect = 3e-16
Identities = 52/116 (44%), Positives = 69/116 (58%), Gaps = 2/116 (1%)
Frame = +1
Query: 567 EKGLRQGDPLSPLLFNLCLNGLSCLLNQFLGEENYCGVRISR-NLTINHLLFADDTLLFC 625
EKGLRQGDPLSP LF LC N LS LL + ++N G++++R + I HLLFADD+LLF
Sbjct: 7 EKGLRQGDPLSPYLFILCANVLSGLLKREGNKQNLHGIQVARSDPKITHLLFADDSLLFA 186
Query: 626 ENNELQLETLCNVLITFLFASGLKVNNQKSEL-IGCNVQTVEVERLAHSFGWSVGT 680
N + T+ VL ++ ASG VN +KSE+ NV E E + G+
Sbjct: 187 RANLTEAATIMQVLHSYQSASGQLVNFEKSEVSYSQNVPNQEKEMICQQIAIKTGS 354
>BE248682 similar to GP|18568269|gb putative gag-pol polyprotein {Zea mays},
partial (1%)
Length = 441
Score = 80.5 bits (197), Expect = 4e-15
Identities = 51/128 (39%), Positives = 68/128 (52%), Gaps = 1/128 (0%)
Frame = +3
Query: 565 SIEKGLRQGDPLSPLLFNLCLNGLSCLLNQFLGEENYCGVRISRNLT-INHLLFADDTLL 623
S+++GL+QGDPL+P LF L G+S L+ + + G + R T ++HL +ADDTL
Sbjct: 15 SVQRGLKQGDPLAPFLFLLVAEGISGLMKNAVNRNLFQGFDVKRGGTRVSHLQYADDTLC 194
Query: 624 FCENNELQLETLCNVLITFLFASGLKVNNQKSELIGCNVQTVEVERLAHSFGWSVGTLPI 683
L TL +L F ASGLKVN KS LIG NV +E ++P
Sbjct: 195 IGMPTVDNLWTLKALLQGFEMASGLKVNFHKSSLIGINVPRDFMEAACRFLNCREESIPF 374
Query: 684 SYLGAPLG 691
YLG P G
Sbjct: 375 IYLGLPGG 398
>TC80819 weakly similar to GP|10140689|gb|AAG13524.1 putative non-LTR
retroelement reverse transcriptase {Oryza sativa
(japonica cultivar-group)}, partial (2%)
Length = 1262
Score = 71.6 bits (174), Expect(2) = 4e-13
Identities = 45/161 (27%), Positives = 75/161 (45%), Gaps = 12/161 (7%)
Frame = +2
Query: 913 GTYSRGKWGWSLTFRG-----QFFAWEEELYQQFLASINHIIPAAVGGDQLIWCANQTGL 967
G G+ GW++ R + F WEEE ++ +N+ + D+ W +
Sbjct: 374 GGADMGRLGWTVDGRAWEWTRRLFVWEEECVRECCILLNNFVLQDNVNDKWRWLLDPVNG 553
Query: 968 FSVKELCYWL------EDHLFTHQMWQVPKSIAKLLPPKVALFFWQAQNDKIATRANLLA 1021
+SVK ++ D +W K +P KV+LF W+ +++ T+ NL+
Sbjct: 554 YSVKVFYRYITSTGHISDRSLVDDVWH------KHIPSKVSLFVWRLLRNRLPTKDNLVH 715
Query: 1022 RGIDLNGSTQCLFCNAFD-ESCNHIFLHCHSIWSLWCVIVN 1061
RG+ L + C+ C D ES H+FLHC+ SLW ++ N
Sbjct: 716 RGVLLATNAACV-CGCVDSESTTHLFLHCNVFCSLWSLVRN 835
Score = 45.1 bits (105), Expect(2) = 8e-07
Identities = 22/62 (35%), Positives = 34/62 (54%)
Frame = +3
Query: 864 FKENLLISIGDGSRVKFWRDPWVSNVALCHLFPRLYAMSENKEARIAEVGTYSRGKWGWS 923
F+EN+ + +GDG FW D W +V L +PRL ++ +KE ++ + G WG
Sbjct: 240 FEENIRMVVGDGRDAFFWYDTWAGDVPLRLKYPRLLDLAMDKECKVG----LTWGGWGGR 407
Query: 924 LT 925
LT
Sbjct: 408 LT 413
Score = 27.3 bits (59), Expect(2) = 8e-07
Identities = 15/38 (39%), Positives = 18/38 (46%)
Frame = +1
Query: 787 GLGLGFIEWRNRGLLLKWAWRFGREINSLWRFFVIERY 824
GLG+G N LL KW WR + LW + RY
Sbjct: 31 GLGVGAF---NLSLLGKWCWRLLVDKEGLWHRVLKARY 135
Score = 22.3 bits (46), Expect(2) = 4e-13
Identities = 8/14 (57%), Positives = 9/14 (64%)
Frame = +1
Query: 1101 WSIWIERNNICFNH 1114
W IW ERNN F +
Sbjct: 946 WVIWKERNNRLFQN 987
>CA919100 homologue to PIR|G90291|G902 endoglucanase precursor [imported] -
Sulfolobus solfataricus, partial (3%)
Length = 789
Score = 69.7 bits (169), Expect = 7e-12
Identities = 43/140 (30%), Positives = 72/140 (50%), Gaps = 4/140 (2%)
Frame = -2
Query: 997 LPPKVALFFWQAQNDKIATRANLLARGIDLNGSTQCLF-CNAFDESCNHIFLHCHSIWSL 1055
+P KV++F W+ D++ T++NL+ RG+ + C+ C A ES H+FL C SL
Sbjct: 569 VPLKVSVFAWRLLRDRLPTKSNLIYRGVIPTEAGLCVSGCGAL-ESAQHLFLSCSYFASL 393
Query: 1056 WCVIVNRERFLWVMPASLADLILEWDFLP--KISDKSLWSLIPYALSWSIWIERNNICFN 1113
W ++ + F+ V L+D +++ + +S LI +W +W ERNN+CFN
Sbjct: 392 WSLVRDWIGFVGVDTNVLSDHFVQFVHSTGGNKASQSFLQLIWLLCAWVLWTERNNMCFN 213
Query: 1114 HKALNISEIWD-FHLLRIGW 1132
+ + D L +GW
Sbjct: 212 DSITPLPRLLDKVKYLSLGW 153
>BG586267 weakly similar to PIR|A84888|A8 hypothetical protein At2g45230
[imported] - Arabidopsis thaliana, partial (16%)
Length = 794
Score = 65.1 bits (157), Expect = 2e-10
Identities = 39/114 (34%), Positives = 64/114 (55%), Gaps = 4/114 (3%)
Frame = +1
Query: 385 TAIKPQGLTALI*NSFRNSGAWFEWMF----EDFFSSGRISRGMNAAFITLIPKKENASS 440
T Q TA + +SF N G +E + ++ G + R L+PKK A
Sbjct: 451 THTSAQARTA*MCSSFNNFGTQWEMISHLWRKNSSEQGNLKRESTKQTSGLVPKKLEAKR 630
Query: 441 LNHYRPISLIRSSYKLIAKVLATRLQKVMPKLLSQNQFAFTKERQIADCILLAN 494
L +RPISL +YK+++KVL+ RL+ V+P ++++ Q AF + + I+D IL+A+
Sbjct: 631 LVEFRPISLCNVAYKIVSKVLSKRLKSVLPWIITETQAAFGRRQLISDNILIAH 792
>BQ148771
Length = 680
Score = 64.7 bits (156), Expect = 2e-10
Identities = 30/103 (29%), Positives = 58/103 (56%)
Frame = -3
Query: 713 SYNSKFMSLSGRLVITKSVLCSIPLFLMVVFKAPKGVILEVEKLFHAFIWGQEQGRKKIN 772
++ + +SL+ R+ + KSV+ ++PL+ M+ PK I E++KL F+WG + ++ +
Sbjct: 567 NWKANHLSLARRVTLAKSVIEAVPLYPMMTTIIPKACIEEIQKLQRKFVWGDTEVSRRYH 388
Query: 773 WIGWDQICLNRKAGGLGLGFIEWRNRGLLLKWAWRFGREINSL 815
+GW+ + + GLGL ++ N+ ++K W NSL
Sbjct: 387 AVGWETMSKPKTIYGLGLRRLDVMNKACIMKLGWSIYSGSNSL 259
>BG447966 weakly similar to PIR|G96509|G96 protein F27F5.21 [imported] -
Arabidopsis thaliana, partial (7%)
Length = 687
Score = 63.5 bits (153), Expect = 5e-10
Identities = 39/118 (33%), Positives = 63/118 (53%), Gaps = 1/118 (0%)
Frame = +2
Query: 381 LNPLTAIKPQGLTALI*NSFRNS-GAWFEWMFEDFFSSGRISRGMNAAFITLIPKKENAS 439
++P+ A P GL AL + + G + M ++ + +N FI LIPK +N +
Sbjct: 293 MHPVKAPGPDGLPALFFQKYWHIVGKEVQQMVLQVLNNSMETEELNKTFIVLIPKGKNPN 472
Query: 440 SLNHYRPISLIRSSYKLIAKVLATRLQKVMPKLLSQNQFAFTKERQIADCILLANEVA 497
+ YRPISL K+I KV+A R+++ +P ++ Q AF + R I D L+A V+
Sbjct: 473 TPKDYRPISLCNVVMKIITKVIANRVKQTLPDVIDVEQSAFVQGRLITDNALIAWSVS 646
>BG587108 similar to PIR|G96509|G9 protein F27F5.21 [imported] - Arabidopsis
thaliana, partial (17%)
Length = 677
Score = 62.0 bits (149), Expect = 1e-09
Identities = 30/71 (42%), Positives = 43/71 (60%)
Frame = +3
Query: 410 MFEDFFSSGRISRGMNAAFITLIPKKENASSLNHYRPISLIRSSYKLIAKVLATRLQKVM 469
M F +SG++ +N I LIPKK+ + + RPISL YK+I+KVL RL+ +
Sbjct: 456 MVNSFLASGKLDTRLNTTNICLIPKKKRPTRMTELRPISLCNVGYKIISKVLCQRLKVCL 635
Query: 470 PKLLSQNQFAF 480
P L+S+ Q AF
Sbjct: 636 PSLISETQSAF 668
>TC90516
Length = 983
Score = 58.9 bits (141), Expect = 1e-08
Identities = 51/167 (30%), Positives = 73/167 (43%)
Frame = -2
Query: 408 EWMFEDFFSSGRISRGMNAAFITLIPKKENASSLNHYRPISLIRSSYKLIAKVLATRLQK 467
E MFE F++ + + + F+TLI K E L + +S + S KL+AKVLA RL
Sbjct: 400 EIMFEQFYTPANLLKIFS*YFLTLISKVEYPILLGDFSLMSFLGSL*KLMAKVLALRLAH 221
Query: 468 VMPKLLSQNQFAFTKERQIADCILLANEVADFLSKKSEGGLMLKVDLAKAYDQVDWEFLI 527
+M K++ NQ F + RQ D ++ NE+ +
Sbjct: 220 IMEKIIFVNQSTFVRGRQHVDGVVAINEIIVLM--------------------------- 122
Query: 528 NMLHEMNFGDKWISWMKSCISSASLAILVNGSPTDFFSIEKGLRQGD 574
GD + C+ AILVN SPT I+KGL+QGD
Sbjct: 121 --------GDGVVD-*SVCVYK*P-AILVNCSPT*EIDIQKGLKQGD 11
>TC82520
Length = 833
Score = 45.8 bits (107), Expect(2) = 2e-08
Identities = 30/118 (25%), Positives = 50/118 (41%), Gaps = 19/118 (16%)
Frame = +2
Query: 1040 ESCNHIFLHCHSIWSLWCVIVNRERFLWVMPASLADLILEWDFL---PKISDKSLWSLIP 1096
E+ H+FLHC SLW ++ L V+PA + +++ + P+ + S ++
Sbjct: 329 ETATHLFLHCDIFGSLWSHVLRWLHLLLVLPADIRQFFIQFTSMAGSPRFT-HSFLQIMW 505
Query: 1097 YALSWSIWIERNNICFNHKALNISE----------IW------DFHLLRIGWWTHELL 1138
+A W +W +RNN F + + S +W F WW H LL
Sbjct: 506 FASVWVLWKKRNNRVFQNSLSDPSTFVEQVKMHSFLWLKFQQATFSFSYHDWWKHPLL 679
Score = 32.0 bits (71), Expect(2) = 2e-08
Identities = 14/46 (30%), Positives = 26/46 (56%)
Frame = +3
Query: 988 QVPKSIAKLLPPKVALFFWQAQNDKIATRANLLARGIDLNGSTQCL 1033
QV K +P KV++F W+ ++++ T+ NL+ R + T C+
Sbjct: 174 QVDDVWQKNIPSKVSMFVWRLFHNRLPTKVNLMQRHVLQQTHTACI 311
>TC91765 weakly similar to GP|19881779|gb|AAM01180.1 Putative retroelement
{Oryza sativa (japonica cultivar-group)}, partial (1%)
Length = 625
Score = 43.5 bits (101), Expect(2) = 6e-08
Identities = 20/47 (42%), Positives = 28/47 (59%)
Frame = +2
Query: 546 CISSASLAILVNGSPTDFFSIEKGLRQGDPLSPLLFNLCLNGLSCLL 592
C+ S +LVN D +GL+QGD LSP +F +C+ GLS L+
Sbjct: 104 CVESNDYYVLVNNDAVDPIIPSRGLQQGDHLSPYIFIICVEGLSFLI 244
Score = 32.7 bits (73), Expect(2) = 6e-08
Identities = 11/30 (36%), Positives = 22/30 (72%)
Frame = +1
Query: 514 LAKAYDQVDWEFLINMLHEMNFGDKWISWM 543
++K Y++VD ++L ++ +M F ++WI WM
Sbjct: 7 ISKVYNRVD*DYLKEIMIKMGFNNRWIYWM 96
>CA921218
Length = 707
Score = 56.2 bits (134), Expect = 8e-08
Identities = 39/130 (30%), Positives = 62/130 (47%)
Frame = -3
Query: 1172 PDAGLIKINVDGAAKGSPGRCGIGGILRNNANVILGFFSKSVGEGYAFEAEVLAIHEALL 1231
P+ I N +G+A + CG G RN+ L F+++ G G A A++ A+
Sbjct: 594 PEPLWINCNTNGSANINTSACG--GTFRNSNAYFLLCFAENTGNGNALHAKLSGAMRAIE 421
Query: 1232 FCVQRQLRNIIIESDSSLAVGWVNNRCNRPWKLLNKLNQIDRWLVETNCLKVNHIYREAN 1291
R ++ +E DSSL V ++ W L N+ N +L+ + V+H++RE N
Sbjct: 420 IAAARNWSHLWLELDSSLVVNAFKSKSVIHWNLRNRWNNC-LFLISSMNFFVSHVFREGN 244
Query: 1292 GEADKLAKRG 1301
AD LA G
Sbjct: 243 QCADGLANFG 214
>AW686588
Length = 567
Score = 54.3 bits (129), Expect = 3e-07
Identities = 34/119 (28%), Positives = 56/119 (46%), Gaps = 2/119 (1%)
Frame = +1
Query: 997 LPPKVALFFWQAQNDKIATRANLLARGIDLNGSTQCLFCNAFDESCNHIFLHCHSIWSLW 1056
+P KV++ W+ D++ T+ANL+ R + C+ E+ NH+FLHC + ++W
Sbjct: 142 VPLKVSILAWRLIRDRLPTKANLVRRRCLAVEAAGCVVGCGIAETANHLFLHCATFGAVW 321
Query: 1057 CVIVNRERFLWVMPASLADLILEWDFLP--KISDKSLWSLIPYALSWSIWIERNNICFN 1113
I P L+D +++ + +S LI W +W ERNN FN
Sbjct: 322 QHIRAWIGVSGADPHDLSDHFIQFITCTGHTRARRSFMQLIWLLCVWMVWNERNNRLFN 498
>BF647389
Length = 404
Score = 54.3 bits (129), Expect = 3e-07
Identities = 36/128 (28%), Positives = 60/128 (46%)
Frame = +2
Query: 679 GTLPISYLGAPLGGNPRRIVFWNPMLEKLKCKTRSYNSKFMSLSGRLVITKSVLCSIPLF 738
G LP YLG PL ++ ++K+ C+ + +SK +S +G L + K VL + +
Sbjct: 26 GKLPSRYLGVPLSSKKLYVI---QRVKKIICRIEN*SSKLLSYAGSLQLIKIVLFGVQPY 196
Query: 739 LMVVFKAPKGVILEVEKLFHAFIWGQEQGRKKINWIGWDQICLNRKAGGLGLGFIEWRNR 798
VF P VI ++ F+ + G K I + ICL + AGG + ++ N+
Sbjct: 197 WSQVFVLP*KVIKLIQTTCRIFL*TGKSGTSKRALIAREHICLPKTAGGWNVIDLKVXNQ 376
Query: 799 GLLLKWAW 806
+ K W
Sbjct: 377 TAICKLXW 400
>BG456581
Length = 683
Score = 45.8 bits (107), Expect = 1e-04
Identities = 31/117 (26%), Positives = 56/117 (47%)
Frame = +2
Query: 997 LPPKVALFFWQAQNDKIATRANLLARGIDLNGSTQCLFCNAFDESCNHIFLHCHSIWSLW 1056
+PP + W+ +D++ T NL +RG L + C FC E+ +H+FL C + +LW
Sbjct: 95 IPPSHSFICWRLAHDRLPTDDNLSSRGCAL--VSMCSFCLEQVETSDHLFLRCKFVVTLW 268
Query: 1057 CVIVNRERFLWVMPASLADLILEWDFLPKISDKSLWSLIPYALSWSIWIERNNICFN 1113
+ ++ R + + +S L+ + L+ + SIW RNN+ F+
Sbjct: 269 SWLCSQLR-VGLDFSSFKALLSSLPRHCSSQVRDLYVAAVVHMVHSIWWARNNVRFS 436
>BF640104
Length = 344
Score = 42.0 bits (97), Expect = 0.002
Identities = 26/89 (29%), Positives = 48/89 (53%), Gaps = 2/89 (2%)
Frame = -1
Query: 1169 WVTPDAGLIKINVDGAAKGSPGRCGIGGILRNNANVIL--GFFSKSVGEGYAFEAEVLAI 1226
W+ P G+IKIN D A S GIG I RN+ +++ G +++ G AE +
Sbjct: 338 WIKPHQGVIKINCD-ANLTSEDVWGIGVITRNDNGIVMASGTWNRP-GFMCPITAEAWGV 165
Query: 1227 HEALLFCVQRQLRNIIIESDSSLAVGWVN 1255
++A LF + + +N++ E+D+ + ++
Sbjct: 164 YQAALFALDQGFQNVLFENDNEKLISMLS 78
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.335 0.145 0.488
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 48,734,024
Number of Sequences: 36976
Number of extensions: 821888
Number of successful extensions: 6505
Number of sequences better than 10.0: 73
Number of HSP's better than 10.0 without gapping: 3541
Number of HSP's successfully gapped in prelim test: 355
Number of HSP's that attempted gapping in prelim test: 2770
Number of HSP's gapped (non-prelim): 4189
length of query: 1310
length of database: 9,014,727
effective HSP length: 108
effective length of query: 1202
effective length of database: 5,021,319
effective search space: 6035625438
effective search space used: 6035625438
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (21.6 bits)
S2: 64 (29.3 bits)
Lotus: description of TM0009.7


