

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0009.6
(309 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
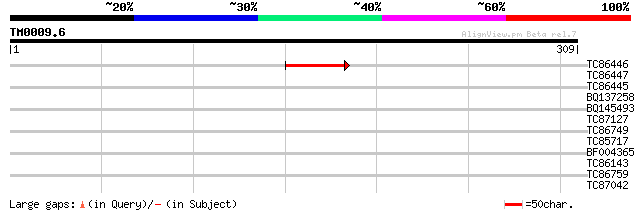
Score E
Sequences producing significant alignments: (bits) Value
TC86446 53 1e-07
TC86447 weakly similar to GP|19911585|dbj|BAB86896. syringolide-... 40 0.001
TC86445 weakly similar to GP|19911585|dbj|BAB86896. syringolide-... 40 0.001
BQ137258 similar to GP|5917666|gb| extensin-like protein {Zea ma... 33 0.18
BQ145493 similar to GP|9759068|dbj| gene_id:MGN6.10~unknown prot... 32 0.31
TC87127 similar to GP|12324745|gb|AAG52327.1 unknown protein; 38... 31 0.69
TC86749 similar to GP|9758060|dbj|BAB08639.1 cysteine-tRNA ligas... 29 2.6
TC85717 similar to SP|Q01899|HS7M_PHAVU Heat shock 70 kDa protei... 28 3.4
BF004365 weakly similar to GP|21539547|gb| unknown protein {Arab... 28 3.4
TC86143 similar to GP|1370182|emb|CAA98168.1 RAB7A {Lotus japoni... 28 5.8
TC86759 similar to GP|16930405|gb|AAL31888.1 At2g32080/F22D22.17... 27 10.0
TC87042 similar to GP|13605827|gb|AAK32899.1 AT5g48380/MJE7_1 {A... 27 10.0
>TC86446
Length = 203
Score = 53.1 bits (126), Expect = 1e-07
Identities = 26/35 (74%), Positives = 30/35 (85%)
Frame = -2
Query: 151 TTINHALITRSVFINQTLVPVSLELKDTVVRAYRT 185
+ INHALI S+FI QT VPVSLE+KDTVVR+YRT
Sbjct: 106 SAINHALIAWSLFIIQTYVPVSLEIKDTVVRSYRT 2
>TC86447 weakly similar to GP|19911585|dbj|BAB86896. syringolide-induced
protein 13-1-1 {Glycine max}, partial (36%)
Length = 1223
Score = 39.7 bits (91), Expect = 0.001
Identities = 18/21 (85%), Positives = 20/21 (94%)
Frame = +1
Query: 169 VPVSLELKDTVVRAYRTDPVQ 189
V VSLE+KDTVVR+YRTDPVQ
Sbjct: 181 VSVSLEIKDTVVRSYRTDPVQ 243
>TC86445 weakly similar to GP|19911585|dbj|BAB86896. syringolide-induced
protein 13-1-1 {Glycine max}, partial (22%)
Length = 588
Score = 39.7 bits (91), Expect = 0.001
Identities = 18/21 (85%), Positives = 20/21 (94%)
Frame = +1
Query: 169 VPVSLELKDTVVRAYRTDPVQ 189
V VSLE+KDTVVR+YRTDPVQ
Sbjct: 106 VSVSLEIKDTVVRSYRTDPVQ 168
>BQ137258 similar to GP|5917666|gb| extensin-like protein {Zea mays}, partial
(3%)
Length = 1053
Score = 32.7 bits (73), Expect = 0.18
Identities = 18/38 (47%), Positives = 20/38 (52%)
Frame = -1
Query: 93 PPGCQSSRSGTVLLRSRCRVDPYLVPLKHTGFAFRRRS 130
PP S S +VL R R P LVP + GF RRRS
Sbjct: 180 PPATPSLPSSSVLYRPPSRPPPPLVPARPPGFGARRRS 67
>BQ145493 similar to GP|9759068|dbj| gene_id:MGN6.10~unknown protein
{Arabidopsis thaliana}, partial (6%)
Length = 153
Score = 32.0 bits (71), Expect = 0.31
Identities = 14/18 (77%), Positives = 16/18 (88%)
Frame = -2
Query: 153 INHALITRSVFINQTLVP 170
+NHALITRS FI+QT VP
Sbjct: 59 VNHALITRSWFIHQTYVP 6
>TC87127 similar to GP|12324745|gb|AAG52327.1 unknown protein; 3866-2463
{Arabidopsis thaliana}, partial (76%)
Length = 747
Score = 30.8 bits (68), Expect = 0.69
Identities = 18/46 (39%), Positives = 25/46 (54%), Gaps = 3/46 (6%)
Frame = -1
Query: 240 ILKCYTHSLIPIQD-CSIHGPPKQFPENFLLKQPIKSTES--EGMF 282
ILK + +P+ D C++ PP QFP+ L Q ES EG+F
Sbjct: 561 ILKGQQFTFVPLIDLCNLEHPPTQFPQPILHSQHFSILESFLEGLF 424
>TC86749 similar to GP|9758060|dbj|BAB08639.1 cysteine-tRNA ligase
{Arabidopsis thaliana}, partial (84%)
Length = 2041
Score = 28.9 bits (63), Expect = 2.6
Identities = 20/60 (33%), Positives = 27/60 (44%)
Frame = -2
Query: 37 FSSTLGGSFTQISGFVLVFLLGHHHLPLTAATLSSTAIPVAAAATLASSSLFRRHLPPGC 96
FSSTLG + + LG H L + + S +AIP A + L S L + L C
Sbjct: 1734 FSSTLGALLSVETACATSGSLGMHGLQIVSLPTSISAIPFAVRSVLTLSLLAKSLLILAC 1555
>TC85717 similar to SP|Q01899|HS7M_PHAVU Heat shock 70 kDa protein
mitochondrial precursor. [Kidney bean French bean]
{Phaseolus vulgaris}, partial (5%)
Length = 677
Score = 28.5 bits (62), Expect = 3.4
Identities = 15/45 (33%), Positives = 21/45 (46%)
Frame = +2
Query: 121 HTGFAFRRRSTTSAEPFCCKEGARVKNVNRTTINHALITRSVFIN 165
HTG RR S+ FC +RV+ V + NH + +IN
Sbjct: 68 HTGNRRRRSSSACVVVFCSSSRSRVRRVLNRSRNHRRRSSGTWIN 202
>BF004365 weakly similar to GP|21539547|gb| unknown protein {Arabidopsis
thaliana}, partial (6%)
Length = 649
Score = 28.5 bits (62), Expect = 3.4
Identities = 17/50 (34%), Positives = 26/50 (52%), Gaps = 1/50 (2%)
Frame = +1
Query: 167 TLVPVSLELKDTVVRAYRTDPVQRFLYQRVP-KNESHIFRPWIYKREEGK 215
T +P +E T V+ + T+ + FLY++VP N H + REE K
Sbjct: 235 TQIPSMVENLTTKVKTWETNEEKSFLYEKVPLLNSLHEYNVQRQLREEEK 384
>TC86143 similar to GP|1370182|emb|CAA98168.1 RAB7A {Lotus japonicus},
partial (96%)
Length = 1314
Score = 27.7 bits (60), Expect = 5.8
Identities = 21/75 (28%), Positives = 33/75 (44%), Gaps = 17/75 (22%)
Frame = -3
Query: 31 LPTSGGFSSTLGGSFTQISGFVLVFLLGHHHLPLTAATL-----------------SSTA 73
L T SSTL SF ++S + ++ L HH L + TL +++
Sbjct: 913 LATHKNASSTL*PSFAEVSKYGMLPLEAHHSLAFFSVTLLLFPPSTSTLFPSSTKGNASG 734
Query: 74 IPVAAAATLASSSLF 88
+PV+A +S LF
Sbjct: 733 LPVSACLRKSSCQLF 689
>TC86759 similar to GP|16930405|gb|AAL31888.1 At2g32080/F22D22.17
{Arabidopsis thaliana}, partial (94%)
Length = 1467
Score = 26.9 bits (58), Expect = 10.0
Identities = 28/89 (31%), Positives = 43/89 (47%), Gaps = 6/89 (6%)
Frame = -3
Query: 8 IDQRMKKIHNSKDKSNPTILGRFLPTSGGFSSTLGGSFTQISGFVLVFLLGHH------H 61
I + +KK N++++++ GR P+S GF L + T+I + V L H
Sbjct: 469 IIEEIKKPGNARERNDD---GR--PSSRGFFRNLKVTTTRILFQIEVKQLVFHL*SLTKK 305
Query: 62 LPLTAATLSSTAIPVAAAATLASSSLFRR 90
L ++ T +STA AAA T SL R
Sbjct: 304 LHVSTTTTTSTATTTAAATTTPGVSLHLR 218
>TC87042 similar to GP|13605827|gb|AAK32899.1 AT5g48380/MJE7_1 {Arabidopsis
thaliana}, partial (51%)
Length = 2206
Score = 26.9 bits (58), Expect = 10.0
Identities = 13/30 (43%), Positives = 22/30 (73%), Gaps = 2/30 (6%)
Frame = -2
Query: 67 ATLSSTAI--PVAAAATLASSSLFRRHLPP 94
AT+SST + P+ ++ T SSS+++ +LPP
Sbjct: 1986 ATISSTMLLSPIISSGTNISSSVWKLYLPP 1897
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.323 0.137 0.419
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 11,387,997
Number of Sequences: 36976
Number of extensions: 184289
Number of successful extensions: 1030
Number of sequences better than 10.0: 24
Number of HSP's better than 10.0 without gapping: 1024
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1029
length of query: 309
length of database: 9,014,727
effective HSP length: 96
effective length of query: 213
effective length of database: 5,465,031
effective search space: 1164051603
effective search space used: 1164051603
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.5 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (22.0 bits)
S2: 58 (26.9 bits)
Lotus: description of TM0009.6


