

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
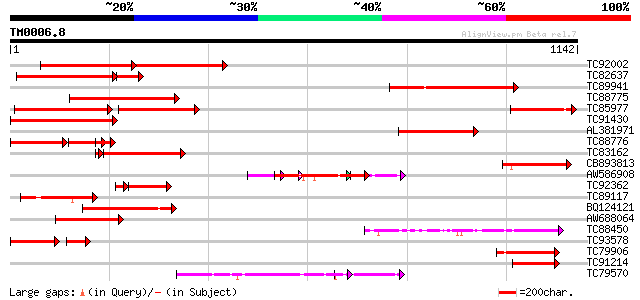
Query= TM0006.8
(1142 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC92002 similar to GP|4104929|gb|AAD02218.1| auxin response fact... 375 0.0
TC82637 similar to PIR|G86331|G86331 IAA24 [imported] - Arabidop... 323 e-104
TC89941 similar to GP|19310546|gb|AAL85006.1 unknown protein {Ar... 370 e-102
TC88775 homologue to PIR|F86427|F86427 auxin response factor 6 (... 358 8e-99
TC85977 similar to GP|10176918|dbj|BAB10162. auxin response fact... 216 2e-98
TC91430 similar to PIR|F86427|F86427 auxin response factor 6 (AR... 305 7e-83
AL381971 similar to GP|1711205|gb|A IAA23 {Arabidopsis thaliana}... 244 1e-64
TC88776 homologue to PIR|F86427|F86427 auxin response factor 6 (... 151 2e-64
TC83162 homologue to PIR|F86427|F86427 auxin response factor 6 (... 214 6e-60
CB893813 similar to GP|19352047|d auxin response factor 7b {Oryz... 192 6e-49
AW586908 weakly similar to GP|2104814|emb| pleiotropic effects o... 182 6e-46
TC92362 similar to GP|19352047|dbj|BAB85917. auxin response fact... 148 3e-42
TC89117 similar to GP|12484199|gb|AAG53998.1 auxin response tran... 158 1e-38
BQ124121 similar to PIR|G86331|G863 IAA24 [imported] - Arabidops... 153 4e-37
AW688064 homologue to GP|13272405|gb unknown protein {Arabidopsi... 147 2e-35
TC88450 similar to PIR|F86427|F86427 auxin response factor 6 (AR... 138 1e-32
TC93578 similar to GP|9757747|dbj|BAB08228.1 auxin response fact... 87 8e-32
TC79906 similar to PIR|F86427|F86427 auxin response factor 6 (AR... 127 2e-29
TC91214 similar to GP|2708484|gb|AAB92476.1| IAA24 {Arabidopsis ... 126 5e-29
TC79570 similar to PIR|F86427|F86427 auxin response factor 6 (AR... 124 2e-28
>TC92002 similar to GP|4104929|gb|AAD02218.1| auxin response factor 7
{Arabidopsis thaliana}, partial (31%)
Length = 1138
Score = 375 bits (964), Expect(2) = 0.0
Identities = 182/194 (93%), Positives = 193/194 (98%)
Frame = +3
Query: 62 DFIPSYPNLPSKLICMLHNVALHADPETDEVYAQMTLQPVNKYDKDAILASDMGLKQNRQ 121
D IPSYPNLPSKLICMLHNVALHADPETDEVYAQMTLQPVNKYDK+A+LASDMGLKQN+Q
Sbjct: 3 DIIPSYPNLPSKLICMLHNVALHADPETDEVYAQMTLQPVNKYDKEAMLASDMGLKQNQQ 182
Query: 122 PTEFFCKTLTASDTSTHGGFSVPRRAAEKIFPPLDYSMQPPAQELVAKDLHDTTWSFRHI 181
P+EFFCKTLTASDTSTHGGFSVPRRAAEKIFPPLD+SMQPPAQE+VAKDL+DTTW+FRHI
Sbjct: 183 PSEFFCKTLTASDTSTHGGFSVPRRAAEKIFPPLDFSMQPPAQEIVAKDLYDTTWTFRHI 362
Query: 182 YRGQPKRHLLTTGWSVFVSTKRLFAGDSVLFIRDEKQQLLLGIRRANRQQPALSSSVISS 241
YRGQPKRHLLTTGWSVFVSTKRLFAGDSVLFIRDEKQQLLLGI+R++RQQPALSSSVISS
Sbjct: 363 YRGQPKRHLLTTGWSVFVSTKRLFAGDSVLFIRDEKQQLLLGIKRSSRQQPALSSSVISS 542
Query: 242 DSMHIGILAAAAHA 255
DSMHIGILAAAAHA
Sbjct: 543 DSMHIGILAAAAHA 584
Score = 342 bits (878), Expect(2) = 0.0
Identities = 161/193 (83%), Positives = 173/193 (89%), Gaps = 1/193 (0%)
Frame = +1
Query: 247 GILAAAAHAAANNSPFTIFYNPRASPSEFVIPLAKFNKAMYTQVSLGMRFRMMFETEESG 306
G L A+NNSPFTIFYNPR SP EFVIPLAK+NKA+YT VSLGMRFRMMFETE+SG
Sbjct: 559 GFLLLRLMLASNNSPFTIFYNPRTSPCEFVIPLAKYNKALYTHVSLGMRFRMMFETEDSG 738
Query: 307 VRRHMGTITGISDLDSARWKSSQWRNLQVGWDESTAGERPSRVSIWEIEPVVTPFYICPP 366
VRR+MGTITGISDLD RWK+SQWRNLQVGWDESTAGERPSRVSIW+IEPVVTPFYICPP
Sbjct: 739 VRRYMGTITGISDLDPVRWKNSQWRNLQVGWDESTAGERPSRVSIWDIEPVVTPFYICPP 918
Query: 367 PFFRPKFPRQPGMPDDESDIENAFKRAMPWLGDDFGMKDASGSIFPGLSLVQWMSM-QQN 425
PFFRPKFP+QPGMP+DES+IENAFKRAMPWLGD+ GMKD S S+FPGLSLVQWMSM N
Sbjct: 919 PFFRPKFPKQPGMPEDESEIENAFKRAMPWLGDELGMKDTSSSVFPGLSLVQWMSMXXNN 1098
Query: 426 NQFSAAQSGCFPP 438
NQFSAAQSG PP
Sbjct: 1099NQFSAAQSGLLPP 1137
>TC82637 similar to PIR|G86331|G86331 IAA24 [imported] - Arabidopsis
thaliana, partial (28%)
Length = 889
Score = 323 bits (828), Expect(2) = e-104
Identities = 153/204 (75%), Positives = 177/204 (86%), Gaps = 1/204 (0%)
Frame = +1
Query: 15 GERKTINSELWHACAGPLVSLPPVGSLVVYFPQGHSEQVAASMQK-ETDFIPSYPNLPSK 73
G RKT+NSELWHACAGPLVSLP VG LV YFPQGHSEQVAAS ++ T IP+YP+LPS+
Sbjct: 115 GVRKTLNSELWHACAGPLVSLPQVGGLVYYFPQGHSEQVAASTRRIATSQIPNYPSLPSQ 294
Query: 74 LICMLHNVALHADPETDEVYAQMTLQPVNKYDKDAILASDMGLKQNRQPTEFFCKTLTAS 133
L+C + NV LHAD ETDE+YAQMTLQP+N +K+ S+ GL +++ PTEFFCKTLTAS
Sbjct: 295 LLCQVQNVTLHADKETDEIYAQMTLQPLNS-EKEVFPVSEFGLSKSKHPTEFFCKTLTAS 471
Query: 134 DTSTHGGFSVPRRAAEKIFPPLDYSMQPPAQELVAKDLHDTTWSFRHIYRGQPKRHLLTT 193
DTSTHGGFSVPRRAAEK+FPPLDY++QPP QELV +DLHD TW+FRHIYRGQPKRHLLTT
Sbjct: 472 DTSTHGGFSVPRRAAEKLFPPLDYTIQPPTQELVVRDLHDNTWTFRHIYRGQPKRHLLTT 651
Query: 194 GWSVFVSTKRLFAGDSVLFIRDEK 217
GWS+FV +KRL AGDSVLFIRDE+
Sbjct: 652 GWSLFVGSKRLRAGDSVLFIRDEE 723
Score = 76.3 bits (186), Expect(2) = e-104
Identities = 37/54 (68%), Positives = 43/54 (79%)
Frame = +2
Query: 216 EKQQLLLGIRRANRQQPALSSSVISSDSMHIGILAAAAHAAANNSPFTIFYNPR 269
+K QL +G+RRANRQQ L SSV+S+DS G+LAAAAHAAAN FTIFYNPR
Sbjct: 719 KKSQLRVGVRRANRQQTTLPSSVLSADSCTFGVLAAAAHAAANRKSFTIFYNPR 880
>TC89941 similar to GP|19310546|gb|AAL85006.1 unknown protein {Arabidopsis
thaliana}, partial (9%)
Length = 799
Score = 370 bits (951), Expect = e-102
Identities = 192/260 (73%), Positives = 212/260 (80%), Gaps = 1/260 (0%)
Frame = +2
Query: 766 EKVLNSNSFSSSLMQPHQLPVNHPQNTQKSLAITRAPSTLTDGDAPSCSTSPSTNNCQIS 825
EK+LN+N SS LM P QLPVNH QNTQKS TRAPS TDGDAPSCSTSPSTNNCQ S
Sbjct: 5 EKLLNNNYSSSPLMHPQQLPVNHLQNTQKSPTNTRAPSAFTDGDAPSCSTSPSTNNCQTS 184
Query: 826 PSNLLKRNQQVPATLGGSLVVESTSNLIQELQSKSDMQIKNEFSNVKGFDQLKYKGTITD 885
P N LKRNQ P T GG +VE+T+NL+QELQSKSDMQ K+E VKG D+ K+KG I D
Sbjct: 185 PPNPLKRNQ--PDTFGGPSMVETTNNLMQELQSKSDMQSKHELHGVKGPDKQKHKGAIND 358
Query: 886 QMEASSSGTSYCLDPGNVQQSLPLSNFCMEGDVQSHS-RSNLPFDSNLDGLTPDPVLMRG 944
MEASS GTSY +DPGN+ Q+ PLSNF M+GDV S RSN+PF SNLDGLTPDP+L RG
Sbjct: 359 HMEASS-GTSYGIDPGNIHQNFPLSNFYMDGDVHSQQPRSNIPFASNLDGLTPDPLLSRG 535
Query: 945 YDSQKDLQNMLSNYGGAPRDIETELSTADISSQSFGVPNMTFNPGCSGDVGINDTGVLNN 1004
YDSQKDLQN+LSNYGGAPRDIETELSTADISSQSFGVPNM+F PGCS ++ I DTGVLNN
Sbjct: 536 YDSQKDLQNLLSNYGGAPRDIETELSTADISSQSFGVPNMSFKPGCSNEIPIADTGVLNN 715
Query: 1005 GLRANQTQRMRTYTKVQKRG 1024
GL +NQ QRMRTYTK QK G
Sbjct: 716 GLWSNQNQRMRTYTKGQKCG 775
>TC88775 homologue to PIR|F86427|F86427 auxin response factor 6 (ARF6)
[imported] - Arabidopsis thaliana, partial (23%)
Length = 678
Score = 358 bits (918), Expect = 8e-99
Identities = 173/222 (77%), Positives = 197/222 (87%), Gaps = 1/222 (0%)
Frame = +3
Query: 121 QPTEFFCKTLTASDTSTHGGFSVPRRAAEKIFPPLDYSMQPPAQELVAKDLHDTTWSFRH 180
+ T +FCKTLTASDTSTHGGFSVPRRAAEK+FPPLD+S QPPAQEL+A+DLH W FRH
Sbjct: 9 EATNYFCKTLTASDTSTHGGFSVPRRAAEKVFPPLDFSQQPPAQELIARDLHGNEWKFRH 188
Query: 181 IYRGQPKRHLLTTGWSVFVSTKRLFAGDSVLFIRDEKQQLLLGIRRANRQQPALSSSVIS 240
I+RGQPKRHLLTTGWSVFVS KRL AGDSVLFI +EK QLLLGIRRA+R Q + SSV+S
Sbjct: 189 IFRGQPKRHLLTTGWSVFVSAKRLVAGDSVLFIWNEKNQLLLGIRRASRPQTVMPSSVLS 368
Query: 241 SDSMHIGILAAAAHAAANNSPFTIFYNPRASPSEFVIPLAKFNKAMY-TQVSLGMRFRMM 299
SDSMH+G+LAAAAHAAA NS FTIFYNPRA PSEFVIPLAK+ KA+Y T+VS+GMRFRM+
Sbjct: 369 SDSMHLGLLAAAAHAAATNSRFTIFYNPRACPSEFVIPLAKYVKAVYHTRVSVGMRFRML 548
Query: 300 FETEESGVRRHMGTITGISDLDSARWKSSQWRNLQVGWDEST 341
FETEES VRR+MGTITGI DL+S RW +S WR+++VGWDEST
Sbjct: 549 FETEESSVRRYMGTITGICDLNSVRWPNSHWRSVKVGWDEST 674
>TC85977 similar to GP|10176918|dbj|BAB10162. auxin response factor-like
protein {Arabidopsis thaliana}, partial (65%)
Length = 3162
Score = 216 bits (550), Expect(2) = 2e-98
Identities = 110/197 (55%), Positives = 137/197 (68%), Gaps = 1/197 (0%)
Frame = +3
Query: 11 NSGEGERKTINSELWHACAGPLVSLPPVGSLVVYFPQGHSEQVAASMQKETD-FIPSYPN 69
++G + ELWHACAGPLV++P G LV YFPQGH EQV AS + ++ +P Y +
Sbjct: 270 STGREAEAALYRELWHACAGPLVTVPREGELVFYFPQGHIEQVEASTNQASEQHMPVY-D 446
Query: 70 LPSKLICMLHNVALHADPETDEVYAQMTLQPVNKYDKDAILASDMGLKQNRQPTEFFCKT 129
L K++C + NV L A+P+TDEV+AQ+TL P D++A+ R FCKT
Sbjct: 447 LRPKILCRVINVMLKAEPDTDEVFAQVTLVPEPNQDENAVEKEAPPAPPPRFHVHSFCKT 626
Query: 130 LTASDTSTHGGFSVPRRAAEKIFPPLDYSMQPPAQELVAKDLHDTTWSFRHIYRGQPKRH 189
LTASDTSTHGGFSV +R A++ PPLD S QPP QELVAKDLH W FRHI+RGQP+RH
Sbjct: 627 LTASDTSTHGGFSVLKRHADECLPPLDMSKQPPTQELVAKDLHGNEWRFRHIFRGQPRRH 806
Query: 190 LLTTGWSVFVSTKRLFA 206
LL +GWSVFVS+KRL A
Sbjct: 807 LLQSGWSVFVSSKRLVA 857
Score = 163 bits (412), Expect(2) = 2e-98
Identities = 81/165 (49%), Positives = 110/165 (66%), Gaps = 3/165 (1%)
Frame = +2
Query: 220 LLLGIRRANRQQPALSSSVISSDSMHIGILAAAAHAAANNSPFTIFYNPRASPSEFVIPL 279
L +G+RRA RQQ + SSVISS SMH+G+LA A HA + FT++Y PR SP+EF++P
Sbjct: 899 LRVGVRRAMRQQGNVPSSVISSHSMHLGVLATAWHAVLTGTMFTVYYKPRTSPAEFIVPY 1078
Query: 280 AKFNKAMYTQVSLGMRFRMMFETEESGVRRHMGTITGISDLDSARWKSSQWRNLQVGWDE 339
++ +++ ++GMRF+M FE EE+ +R GTI GI D DS RW +S+WR L+V WDE
Sbjct: 1079 DQYMESLKNNYTIGMRFKMRFEGEEAPEQRFTGTIVGIEDSDSKRWPTSKWRCLKVRWDE 1258
Query: 340 STAGERPSRVSIWEIEPVVTPFYICPPPFFRPKFPRQ---PGMPD 381
++ RP RVS W+IEP + P + P P RPK PR P PD
Sbjct: 1259 TSNIPRPERVSPWKIEPALAPPALNPLPMPRPKRPRANVVPSSPD 1393
Score = 97.1 bits (240), Expect = 4e-20
Identities = 56/133 (42%), Positives = 82/133 (61%), Gaps = 1/133 (0%)
Frame = +2
Query: 1010 QTQRMRTYTKVQKRG-SVGRCIDVTRYKGYDELRNDLARMFGIEGQLEDPQRTDWKLVYV 1068
Q+ R+ TKV K+G ++GR +D+T++ YDEL +L ++F G+L PQ+ DW +V+
Sbjct: 2246 QSGSARSCTKVHKKGIALGRSVDLTKFSDYDELTAELDQLFEFRGELISPQK-DWLVVFT 2422
Query: 1069 DHENDILLVGDDPWEEFVSCVQSIKILSYTEVQQMSLDGDLGNVPIPNQACSGTDSGNAW 1128
D+E D++LVGDDPW+EF S V+ I I E+Q+MS G L + N S TD G+A
Sbjct: 2423 DNEGDMMLVGDDPWQEFCSMVRKIYIYPKEEIQKMS-PGTLSSKNEENH--SATDGGDAQ 2593
Query: 1129 RGPYDDNSAASFN 1141
N +AS N
Sbjct: 2594 ETKSQLNQSASDN 2632
>TC91430 similar to PIR|F86427|F86427 auxin response factor 6 (ARF6)
[imported] - Arabidopsis thaliana, partial (23%)
Length = 728
Score = 305 bits (781), Expect = 7e-83
Identities = 148/218 (67%), Positives = 174/218 (78%), Gaps = 1/218 (0%)
Frame = +3
Query: 1 MKAPSNCYLPNSGEGERKTINSELWHACAGPLVSLPPVGSLVVYFPQGHSEQVAASMQKE 60
MK S + P EGE++ ++SELWHACAGPLV LP GS VVYFPQGHSEQVA S +E
Sbjct: 63 MKLSSAGFSPPPQEGEKRVLDSELWHACAGPLVCLPADGSRVVYFPQGHSEQVAVSTNRE 242
Query: 61 TDFIPSYPNLPSKLICMLHNVALHADPETDEVYAQMTLQPVNKYD-KDAILASDMGLKQN 119
D P+LP +LIC LHN+ +HAD ETDEVYAQMTLQP+N + K+A L +++G N
Sbjct: 243 VDTYIPNPSLPPQLICQLHNLTMHADTETDEVYAQMTLQPLNPEEQKEAYLPAELGTA-N 419
Query: 120 RQPTEFFCKTLTASDTSTHGGFSVPRRAAEKIFPPLDYSMQPPAQELVAKDLHDTTWSFR 179
+QPT +FCK LTASDTSTHGGFSVPRRAAEK+FPPLD++ QPPAQEL+A+DLH W F+
Sbjct: 420 KQPTNYFCKILTASDTSTHGGFSVPRRAAEKVFPPLDFTQQPPAQELIARDLHGNDWKFK 599
Query: 180 HIYRGQPKRHLLTTGWSVFVSTKRLFAGDSVLFIRDEK 217
HI+RGQPKRHLLTTGWSVFVS KRL AGDSVLFI +EK
Sbjct: 600 HIFRGQPKRHLLTTGWSVFVSAKRLVAGDSVLFIWNEK 713
>AL381971 similar to GP|1711205|gb|A IAA23 {Arabidopsis thaliana}, partial
(4%)
Length = 482
Score = 244 bits (624), Expect = 1e-64
Identities = 124/161 (77%), Positives = 135/161 (83%)
Frame = +2
Query: 783 QLPVNHPQNTQKSLAITRAPSTLTDGDAPSCSTSPSTNNCQISPSNLLKRNQQVPATLGG 842
QL VN +NTQK L ITR PSTLTDG+APSCSTSPSTNNC+I+ NLLKRNQQVP T+GG
Sbjct: 2 QLSVNQTRNTQKPLTITRVPSTLTDGEAPSCSTSPSTNNCRITQPNLLKRNQQVPTTIGG 181
Query: 843 SLVVESTSNLIQELQSKSDMQIKNEFSNVKGFDQLKYKGTITDQMEASSSGTSYCLDPGN 902
L VE TSNLI +LQSKSDM I +EFSNVKG DQL YKGT TDQ+EA SSGTSYCLDPGN
Sbjct: 182 ILAVEPTSNLIHDLQSKSDMHIXHEFSNVKGSDQLXYKGTTTDQLEA-SSGTSYCLDPGN 358
Query: 903 VQQSLPLSNFCMEGDVQSHSRSNLPFDSNLDGLTPDPVLMR 943
VQQ+LPLSNFCMEGDVQS+ R NLPFDS LDGL D +L R
Sbjct: 359 VQQNLPLSNFCMEGDVQSNPRINLPFDSYLDGLMSDTMLSR 481
>TC88776 homologue to PIR|F86427|F86427 auxin response factor 6 (ARF6)
[imported] - Arabidopsis thaliana, partial (22%)
Length = 1278
Score = 151 bits (381), Expect(2) = 2e-64
Identities = 74/117 (63%), Positives = 91/117 (77%), Gaps = 2/117 (1%)
Frame = +3
Query: 1 MKAPSNCYLPNSGEGERKTINSELWHACAGPLVSLPPVGSLVVYFPQGHSEQVAASMQKE 60
M+ S + P EGE++ ++SELWHACAGPLVSLP VGS VVYFPQGHSEQVA S KE
Sbjct: 516 MRLSSASFSPPPQEGEKRVLDSELWHACAGPLVSLPAVGSRVVYFPQGHSEQVAVSTNKE 695
Query: 61 TD-FIPSYPNLPSKLICMLHNVALHADPETDEVYAQMTLQPVNKYD-KDAILASDMG 115
D IP+YP+LP +LIC LHN+ +HAD ETDEVYAQMTLQP+N + K+A L +++G
Sbjct: 696 VDAHIPNYPSLPPQLICQLHNLTMHADVETDEVYAQMTLQPLNAQEQKEAYLPAELG 866
Score = 114 bits (286), Expect(2) = 2e-64
Identities = 52/78 (66%), Positives = 61/78 (77%)
Frame = +1
Query: 118 QNRQPTEFFCKTLTASDTSTHGGFSVPRRAAEKIFPPLDYSMQPPAQELVAKDLHDTTWS 177
Q +QPT +FCKTLTASDTSTHGGFSVPRRAAEK+FPPLD+S QPPAQEL+A+DLH W
Sbjct: 871 QGKQPTNYFCKTLTASDTSTHGGFSVPRRAAEKVFPPLDFSQQPPAQELIARDLHGNEWK 1050
Query: 178 FRHIYRGQPKRHLLTTGW 195
F+ Y +P + T W
Sbjct: 1051FQ-TYLPRPAQKTSTYNW 1101
Score = 73.6 bits (179), Expect = 4e-13
Identities = 34/40 (85%), Positives = 36/40 (90%)
Frame = +2
Query: 174 TTWSFRHIYRGQPKRHLLTTGWSVFVSTKRLFAGDSVLFI 213
T +FRHI+RGQPKRHLLTTGWSVFVS KRL AGDSVLFI
Sbjct: 1040 TNGNFRHIFRGQPKRHLLTTGWSVFVSAKRLVAGDSVLFI 1159
>TC83162 homologue to PIR|F86427|F86427 auxin response factor 6 (ARF6)
[imported] - Arabidopsis thaliana, partial (18%)
Length = 569
Score = 214 bits (545), Expect(2) = 6e-60
Identities = 113/167 (67%), Positives = 137/167 (81%), Gaps = 2/167 (1%)
Frame = +2
Query: 190 LLTTGWSVFVSTKRLFAGDSVLFIRDEKQQLLLGIRRANRQQPALSSSVISSDSMHIGIL 249
LLTTGWSVFVS KRL AGDSVLFI +EK QLLLGIRRA+R Q + SSV+SSDSMHIG+L
Sbjct: 62 LLTTGWSVFVSAKRLVAGDSVLFIWNEKNQLLLGIRRASRPQTVMPSSVLSSDSMHIGLL 241
Query: 250 AAAAHAAANNSPFTIFYNPRASPSEFVIPLAKFNKAMY-TQVSLGM-RFRMMFETEESGV 307
AAAAHAAA NS FT+F+NPRASPSEFVIPL+K+ KA+Y T+ L + RFRM+FETEES V
Sbjct: 242 AAAAHAAATNSCFTVFFNPRASPSEFVIPLSKYIKAVYHTRPFLSVCRFRMLFETEESSV 421
Query: 308 RRHMGTITGISDLDSARWKSSQWRNLQVGWDESTAGERPSRVSIWEI 354
RR+MGTIT ISD+D RW +S WR+++VG + G + + VS+WE+
Sbjct: 422 RRYMGTITSISDMDPVRWPNSHWRSVRVGGMNHSWG-KTATVSLWEM 559
Score = 36.6 bits (83), Expect(2) = 6e-60
Identities = 13/17 (76%), Positives = 14/17 (81%)
Frame = +1
Query: 173 DTTWSFRHIYRGQPKRH 189
D W FRHI+RGQPKRH
Sbjct: 10 DVEWKFRHIFRGQPKRH 60
>CB893813 similar to GP|19352047|d auxin response factor 7b {Oryza sativa},
partial (10%)
Length = 787
Score = 192 bits (488), Expect = 6e-49
Identities = 98/143 (68%), Positives = 112/143 (77%), Gaps = 5/143 (3%)
Frame = +1
Query: 993 DVGINDTGVLNNGLRA-----NQTQRMRTYTKVQKRGSVGRCIDVTRYKGYDELRNDLAR 1047
D IND+G L++G A +Q QR+RTYTKV KRG+VGR ID+TRY GYDEL++DLAR
Sbjct: 1 DSTINDSGFLDSGPWAPRPPPHQFQRIRTYTKVYKRGAVGRSIDITRYSGYDELKHDLAR 180
Query: 1048 MFGIEGQLEDPQRTDWKLVYVDHENDILLVGDDPWEEFVSCVQSIKILSYTEVQQMSLDG 1107
FGIEGQLED QR WKLVYVDHEND+LLVGDDPWEEFV+CV+ IKILS EVQQMSLDG
Sbjct: 181 RFGIEGQLEDRQRVGWKLVYVDHENDVLLVGDDPWEEFVNCVRCIKILSPQEVQQMSLDG 360
Query: 1108 DLGNVPIPNQACSGTDSGNAWRG 1130
D GN +P QA S +D G A G
Sbjct: 361 DFGNGGLPRQASSSSDGGIA*MG 429
>AW586908 weakly similar to GP|2104814|emb| pleiotropic effects on cellular
differentiation and slug behaviour {Dictyostelium
discoideum}, partial (8%)
Length = 668
Score = 182 bits (462), Expect = 6e-46
Identities = 121/219 (55%), Positives = 138/219 (62%), Gaps = 27/219 (12%)
Frame = +2
Query: 534 QQQQQQLQQQQSQQQQLQSLLQIPMNQLQQP--RQQQLPEPKNLPLPQQQLPQQLGQQS- 590
QQ QQQ QQQ++QSLL +P++QLQQ RQQQL +NLP PQQQ PQ L QQS
Sbjct: 2 QQPPASWSQQQQQQQKMQSLLPMPLDQLQQQQQRQQQLAGSQNLPQPQQQQPQ-LTQQSP 178
Query: 591 -------------QKAIMSNGVVASNQITNQLVLQ-------QQQLLSGGISPQQSIHSA 630
Q M+NG + SNQI NQ V Q QQQLLSG + QQ++ SA
Sbjct: 179 QQPQQQQQHQQPCQNTTMNNGTIGSNQIPNQCVQQPVTYSQLQQQLLSGSMQSQQNLQSA 358
Query: 631 SKTTFPLTSLPQESQFLQQID-QQASLLQRQQLPQQTQLQQSSLHLLQQNQPQRAPQQPP 689
K +TSLPQ+SQF QQID QQA LLQRQQ QQ QLQQSSL LLQQ+ QR PQQP
Sbjct: 359 GKNGLMMTSLPQDSQFQQQIDQQQAGLLQRQQ--QQNQLQQSSLQLLQQSMLQRGPQQPQ 532
Query: 690 VATMSQQTPSEQQLHLQLLQKLQQHQQHQQL---LSTSS 725
++ Q S+QQ LQLLQKLQQ QQ QQ LSTSS
Sbjct: 533 MSQTVPQNISDQQQQLQLLQKLQQQQQQQQQQQPLSTSS 649
Score = 68.2 bits (165), Expect = 2e-11
Identities = 50/121 (41%), Positives = 64/121 (52%), Gaps = 7/121 (5%)
Frame = +2
Query: 479 NQI-NQLHQSPVSWPQQQQQQQQQQQQQQQQLQLQHQQQQQQLQQQQQQQQQQQLQQQQQ 537
NQI NQ Q PV++ Q QQQ Q QQ LQ + Q Q QQQ+ QQQ
Sbjct: 254 NQIPNQCVQQPVTYSQLQQQLLSGSMQSQQNLQSAGKNGLMMTSLPQDSQFQQQIDQQQA 433
Query: 538 QQLQQQQSQQQQLQSLLQIPMNQLQQ--PRQQQLPE--PKNLPLPQQQLP--QQLGQQSQ 591
LQ+QQ Q Q QS LQ+ + Q P+Q Q+ + P+N+ QQQL Q+L QQ Q
Sbjct: 434 GLLQRQQQQNQLQQSSLQLLQQSMLQRGPQQPQMSQTVPQNISDQQQQLQLLQKLQQQQQ 613
Query: 592 K 592
+
Sbjct: 614 Q 616
Score = 60.1 bits (144), Expect = 5e-09
Identities = 69/209 (33%), Positives = 79/209 (37%)
Frame = +2
Query: 480 QINQLHQSPVSWPQQQQQQQQQQQQQQQQLQLQHQQQQQQLQQQQQQQQQQQLQQQQQQQ 539
Q Q Q ++ Q Q QQQQ Q QQ Q QQQQQ QQ Q Q
Sbjct: 86 QQQQQRQQQLAGSQNLPQPQQQQPQLTQQSPQQPQQQQQ--HQQPCQNTTMNNGTIGSNQ 259
Query: 540 LQQQQSQQQQLQSLLQIPMNQLQQPRQQQLPEPKNLPLPQQQLPQQLGQQSQKAIMSNGV 599
+ Q QQ S LQ + QQ L L LPQ Q Q G+
Sbjct: 260 IPNQCVQQPVTYSQLQQQLLSGSMQSQQNLQSAGKNGLMMTSLPQDSQFQQQIDQQQAGL 439
Query: 600 VASNQITNQLVLQQQQLLSGGISPQQSIHSASKTTFPLTSLPQESQFLQQIDQQASLLQR 659
+ Q NQL QLL QQS+ L PQ+ Q Q + Q S
Sbjct: 440 LQRQQQQNQLQQSSLQLL------QQSM---------LQRGPQQPQMSQTVPQNIS---- 562
Query: 660 QQLPQQTQLQQSSLHLLQQNQPQRAPQQP 688
QQ QLQ L LQQ Q Q+ QQP
Sbjct: 563 ---DQQQQLQ--LLQKLQQQQQQQQQQQP 634
Score = 57.0 bits (136), Expect = 4e-08
Identities = 44/115 (38%), Positives = 56/115 (48%), Gaps = 3/115 (2%)
Frame = +2
Query: 686 QQPPVATMSQQTPSEQQLHLQL---LQKLQQHQQHQQLLSTSSPLLQPQLLQQQSTPQQN 742
QQPP A+ SQQ +Q++ L L +LQQ QQ QQ L+ S L QP QQ PQ
Sbjct: 2 QQPP-ASWSQQQQQQQKMQSLLPMPLDQLQQQQQRQQQLAGSQNLPQP----QQQQPQLT 166
Query: 743 QQLTQLPASQHQTQQLGNNAFSTEKVLNSNSFSSSLMQPHQLPVNHPQNTQKSLA 797
QQ Q P Q Q QQ N + SN + +Q PV + Q Q+ L+
Sbjct: 167 QQSPQQPQQQQQHQQPCQNTTMNNGTIGSNQIPNQCVQQ---PVTYSQLQQQLLS 322
Score = 50.1 bits (118), Expect = 5e-06
Identities = 33/75 (44%), Positives = 36/75 (48%)
Frame = +2
Query: 480 QINQLHQSPVSWPQQQQQQQQQQQQQQQQLQLQHQQQQQQLQQQQQQQQQQQLQQQQQQQ 539
QI+Q + QQQ Q QQ Q QQ LQ QQ Q+ Q Q Q QQ Q Q
Sbjct: 413 QIDQQQAGLLQRQQQQNQLQQSSLQLLQQSMLQRGPQQPQMSQTVPQNISDQQQQLQLLQ 592
Query: 540 LQQQQSQQQQLQSLL 554
QQQ QQQQ Q L
Sbjct: 593 KLQQQQQQQQQQQPL 637
Score = 36.2 bits (82), Expect = 0.074
Identities = 34/102 (33%), Positives = 45/102 (43%), Gaps = 2/102 (1%)
Frame = +2
Query: 420 MSMQQNNQFSAAQSGCFPPMLSPNTLHSNLGTDDPSKLLSFQAPA--LSAPSLQFNKPNL 477
M QQN Q SA ++G L ++ + LL Q L SLQ + ++
Sbjct: 329 MQSQQNLQ-SAGKNGLMMTSLPQDSQFQQQIDQQQAGLLQRQQQQNQLQQSSLQLLQQSM 505
Query: 478 PNQINQLHQSPVSWPQQQQQQQQQQQQQQQQLQLQHQQQQQQ 519
+ Q Q + PQ QQQQ Q Q+ Q Q QQQQQQ
Sbjct: 506 LQRGPQQPQMSQTVPQNISDQQQQLQLLQKLQQQQQQQQQQQ 631
>TC92362 similar to GP|19352047|dbj|BAB85917. auxin response factor 7b
{Oryza sativa}, partial (11%)
Length = 399
Score = 148 bits (373), Expect(2) = 3e-42
Identities = 71/89 (79%), Positives = 82/89 (91%), Gaps = 1/89 (1%)
Frame = +2
Query: 239 ISSDSMHIGILAAAAHAAANNSPFTIFYNPRASPSEFVIPLAKFNKAMYT-QVSLGMRFR 297
+SSDSMHIGILAAAAHA+ANNSPFT+FYNPRASPSEFVIPLAK+ +A+Y+ Q+S GMRFR
Sbjct: 128 LSSDSMHIGILAAAAHASANNSPFTVFYNPRASPSEFVIPLAKYYRAVYSHQISPGMRFR 307
Query: 298 MMFETEESGVRRHMGTITGISDLDSARWK 326
MMFETE+SG RR+MGT G+SDLDS RWK
Sbjct: 308 MMFETEDSGTRRYMGTXIGVSDLDSVRWK 394
Score = 43.5 bits (101), Expect(2) = 3e-42
Identities = 22/26 (84%), Positives = 23/26 (87%)
Frame = +1
Query: 214 RDEKQQLLLGIRRANRQQPALSSSVI 239
RDEKQQLLLGIRRANRQ LSSSV+
Sbjct: 52 RDEKQQLLLGIRRANRQPTNLSSSVL 129
>TC89117 similar to GP|12484199|gb|AAG53998.1 auxin response transcription
factor 3 {Arabidopsis thaliana}, partial (26%)
Length = 861
Score = 158 bits (400), Expect = 1e-38
Identities = 84/163 (51%), Positives = 100/163 (60%), Gaps = 9/163 (5%)
Frame = +1
Query: 23 ELWHACAGPLVSLPPVGSLVVYFPQGHSEQVAASMQKETDFIPSYPNLPSKLICMLHNVA 82
ELWHACAGPL+SLP GS+VVY PQGH EQ DF S N+P + C + +V
Sbjct: 370 ELWHACAGPLISLPKKGSIVVYVPQGHFEQAH-------DFPVSACNIPPHVFCRVLDVK 528
Query: 83 LHADPETDEVYAQMTLQPVNKYDKDAILASDMGLKQNRQPTE---------FFCKTLTAS 133
LHA+ +DEVY Q+ L P N+ + + + + TE FCKTLTAS
Sbjct: 529 LHAEEGSDEVYCQVLLVPENQQLEQNVREGVIDADAEEEDTEAIVKSTTPHMFCKTLTAS 708
Query: 134 DTSTHGGFSVPRRAAEKIFPPLDYSMQPPAQELVAKDLHDTTW 176
DTSTHGGFSVPR AAE FPPLDY Q P+QELV KDLH + W
Sbjct: 709 DTSTHGGFSVPRXAAEDCFPPLDYGQQRPSQELVXKDLHGSKW 837
>BQ124121 similar to PIR|G86331|G863 IAA24 [imported] - Arabidopsis thaliana,
partial (2%)
Length = 569
Score = 153 bits (386), Expect = 4e-37
Identities = 78/190 (41%), Positives = 116/190 (61%), Gaps = 2/190 (1%)
Frame = +2
Query: 148 AEKIFPPLDYSMQPPAQELVAKDLHDTTWSFRHIYRGQPKRHLLTTGWSVFVSTKRLFAG 207
A++ FP LD ++QPP Q+LV KDL W+FRHIY + HLLT GW+ FV++K L G
Sbjct: 5 ADRCFP*LDMTLQPPMQDLVVKDLQGIEWNFRHIYCDHERAHLLTNGWNTFVNSKNLRPG 184
Query: 208 DSVLFIRDEKQQLLLGIRRANRQQPALSSSVI--SSDSMHIGILAAAAHAAANNSPFTIF 265
DS +F+ E ++ +GIRRA +Q + + + SS ++ +G LAA HA + S F +
Sbjct: 185 DSCIFVSGENGEIGIGIRRAMKQHSHICTKLCQQSSQNIQLGALAAVVHAVSIGSLFHLQ 364
Query: 266 YNPRASPSEFVIPLAKFNKAMYTQVSLGMRFRMMFETEESGVRRHMGTITGISDLDSARW 325
Y+P +P EF+IPL + +++ S+G R M+ +E G R GTI G D+D RW
Sbjct: 365 YHPWIAPFEFMIPLKTYVESIEKDYSIGTRVHML--SEVGGCPRRYGTIVGNEDIDPIRW 538
Query: 326 KSSQWRNLQV 335
S+WR ++V
Sbjct: 539 PGSEWRCIKV 568
>AW688064 homologue to GP|13272405|gb unknown protein {Arabidopsis thaliana},
partial (15%)
Length = 576
Score = 147 bits (372), Expect = 2e-35
Identities = 71/137 (51%), Positives = 95/137 (68%)
Frame = +1
Query: 93 YAQMTLQPVNKYDKDAILASDMGLKQNRQPTEFFCKTLTASDTSTHGGFSVPRRAAEKIF 152
++++TL P+ + + + G + + +P F KTLT SD + GGFSVPR AE IF
Sbjct: 1 FSKITLIPLRNSELENDDSDGDGSENSEKPASF-AKTLTQSDANNGGGFSVPRYCAETIF 177
Query: 153 PPLDYSMQPPAQELVAKDLHDTTWSFRHIYRGQPKRHLLTTGWSVFVSTKRLFAGDSVLF 212
P LDYS +PP Q ++AKD+H W FRHIYRG P+RHLLTTGWS FV+ K+L AGDS++F
Sbjct: 178 PRLDYSAEPPVQTVIAKDVHGEVWKFRHIYRGTPRRHLLTTGWSSFVNQKKLVAGDSIVF 357
Query: 213 IRDEKQQLLLGIRRANR 229
+R E +L +GIRRA R
Sbjct: 358 LRAESGELFVGIRRAKR 408
>TC88450 similar to PIR|F86427|F86427 auxin response factor 6 (ARF6)
[imported] - Arabidopsis thaliana, partial (17%)
Length = 1422
Score = 138 bits (347), Expect = 1e-32
Identities = 136/434 (31%), Positives = 203/434 (46%), Gaps = 33/434 (7%)
Frame = +3
Query: 714 HQQHQQLLSTSSPLLQPQLLQQQSTPQQ--------NQQLTQLPASQHQTQQ--LGNNAF 763
+ Q Q LS S P Q Q+ PQQ N QL Q QTQQ + NN
Sbjct: 6 NNQENQNLSQSQP-------QAQTNPQQHPQHQHSFNNQLHHHSQQQQQTQQQVVDNNQQ 164
Query: 764 STEKVLNSNSFSSSLMQPHQLPVNHPQNTQKSLAITRAPSTLTDGDAPSCSTSPSTNNCQ 823
+ V + F S+ QP P P SL ++ +D + S +T
Sbjct: 165 ISGSVSTMSQFVSAT-QPQSPP---PMQALSSLCHQQS---FSDSNVNSSTTI------- 302
Query: 824 ISPSNLLKRNQQVPATLGGSLVVESTSNLIQELQSKSDMQIKNEFSNVKGFDQLKYKGTI 883
+SP + + +G S + +S L+ ++ S + ++N G+ K
Sbjct: 303 VSPLHSI---------MGSSFPHDESSLLMSLPRTSSWVPVQNS----TGWPS---KRIA 434
Query: 884 TDQMEASSSGTSYCLDP-----GNVQQSL----------PLSNFCMEGDVQSHSRSNLPF 928
D + SSG S C+ P G + S+ P ++ + + +SNL F
Sbjct: 435 VDPL--LSSGASQCILPQVEQLGQARNSMSQNAITLPPFPGRECSIDQEGSNDPQSNLLF 608
Query: 929 DSNLDGLTPDPVLMRGYDSQKDLQNMLSNYGGAPRDIETELSTADISSQSF---GVPNMT 985
N+D P +L L N +SN+ G + + + S S+ + +
Sbjct: 609 GVNID---PSSLL---------LHNGMSNFKGISGNNNDSSTMSYHQSSSYMNTAGADSS 752
Query: 986 FNPGCSGDVGINDTGVLN---NGLRANQTQRMRTYTKVQKRGSVGRCIDVTRYKGYDELR 1042
N G + +G ++G L+ NG + N +T+ KV K GS GR +D+T++ Y+ELR
Sbjct: 753 LNHGVTPSIG--ESGFLHTQENGEQGNNPLN-KTFVKVYKSGSFGRSLDITKFSSYNELR 923
Query: 1043 NDLARMFGIEGQLEDPQRTDWKLVYVDHENDILLVGDDPWEEFVSCVQSIKILSYTEVQQ 1102
++LARMFG+EG+LEDP R+ W+LV+VD END+LL+GD PW EFV+ V IKILS EVQQ
Sbjct: 924 SELARMFGLEGELEDPVRSGWQLVFVDRENDVLLLGDGPWPEFVNSVWCIKILSPEEVQQ 1103
Query: 1103 MSLD--GDLGNVPI 1114
M G L +VPI
Sbjct: 1104 MGNTGLGLLNSVPI 1145
Score = 37.4 bits (85), Expect = 0.033
Identities = 28/77 (36%), Positives = 29/77 (37%), Gaps = 4/77 (5%)
Frame = +3
Query: 496 QQQQQQQQQQQQQLQLQHQQQQQQLQQQQQQQQQQQLQQQQQQQLQQQQSQQQQLQSLLQ 555
Q Q Q Q QQ Q QH Q QQQQQ QQ QQ+ S Q S Q
Sbjct: 33 QSQPQAQTNPQQHPQHQHSFNNQLHHHSQQQQQTQQQVVDNNQQISGSVSTMSQFVSATQ 212
Query: 556 ----IPMNQLQQPRQQQ 568
PM L QQ
Sbjct: 213 PQSPPPMQALSSLCHQQ 263
Score = 37.4 bits (85), Expect = 0.033
Identities = 77/327 (23%), Positives = 114/327 (34%), Gaps = 4/327 (1%)
Frame = +3
Query: 494 QQQQQQQQQQQQQQQLQLQHQQQQQQLQQQQQQQQQQQLQQQQQQQLQQQQSQQQQLQSL 553
Q+ Q Q Q Q Q QH Q Q Q QQQ QQ QQQ Q S
Sbjct: 12 QENQNLSQSQPQAQTNPQQHPQHQHSFNNQLHHHSQQQ------QQTQQQVVDNNQQISG 173
Query: 554 LQIPMNQLQQPRQQQLPEPKNLPLPQQQLPQQLGQQSQKAIMSNGVVASNQITNQLVLQQ 613
M+Q Q Q P P Q L QQS + V +S I
Sbjct: 174 SVSTMSQFVSATQPQSPP------PMQALSSLCHQQS---FSDSNVNSSTTI-------- 302
Query: 614 QQLLSGGISPQQSIHSASKTTFPLTSLPQESQFLQQIDQQASLLQRQQLPQQTQLQQSSL 673
+SP +HS ++FP ES L + + +S +P Q S
Sbjct: 303 -------VSP---LHSIMGSSFP----HDESSLLMSLPRTSS-----WVPVQNSTGWPS- 422
Query: 674 HLLQQNQPQRAPQQPPVATMSQQTPSEQQLHLQLLQKLQQHQQHQQLLSTSSPLLQPQLL 733
+R P +++ + Q +L +++Q Q + +S ++ L P
Sbjct: 423 --------KRIAVDPLLSSGASQC---------ILPQVEQLGQARNSMSQNAITLPPFPG 551
Query: 734 QQQSTPQQNQQLTQ----LPASQHQTQQLGNNAFSTEKVLNSNSFSSSLMQPHQLPVNHP 789
++ S Q+ Q + + L +N S K ++ N+ SS M HQ
Sbjct: 552 RECSIDQEGSNDPQSNLLFGVNIDPSSLLLHNGMSNFKGISGNNNDSSTMSYHQ------ 713
Query: 790 QNTQKSLAITRAPSTLTDGDAPSCSTS 816
+ + A S+L G PS S
Sbjct: 714 --SSSYMNTAGADSSLNHGVTPSIGES 788
Score = 33.9 bits (76), Expect = 0.37
Identities = 24/53 (45%), Positives = 24/53 (45%), Gaps = 1/53 (1%)
Frame = +3
Query: 479 NQINQ-LHQSPVSWPQQQQQQQQQQQQQQQQLQLQHQQQQQQLQQQQQQQQQQ 530
NQ NQ L QS QQ Q Q QL H QQQQQ QQQ QQ
Sbjct: 9 NQENQNLSQSQPQAQTNPQQHPQHQHSFNNQLH-HHSQQQQQTQQQVVDNNQQ 164
>TC93578 similar to GP|9757747|dbj|BAB08228.1 auxin response factor 4
{Arabidopsis thaliana}, partial (18%)
Length = 634
Score = 86.7 bits (213), Expect(2) = 8e-32
Identities = 47/98 (47%), Positives = 61/98 (61%)
Frame = +1
Query: 3 APSNCYLPNSGEGERKTINSELWHACAGPLVSLPPVGSLVVYFPQGHSEQVAASMQKETD 62
+PS C +S + ELWHACAGPL SLP G++VVYFPQGH EQ A+ +
Sbjct: 124 SPSTCSSSSSTSPLVSSSYLELWHACAGPLTSLPKKGNVVVYFPQGHLEQFASFSPFKQL 303
Query: 63 FIPSYPNLPSKLICMLHNVALHADPETDEVYAQMTLQP 100
IP+Y +L ++ C + NV L A+ E DEVY Q+TL P
Sbjct: 304 EIPNY-DLQPQIFCRVVNVQLLANKENDEVYTQVTLLP 414
Score = 70.1 bits (170), Expect(2) = 8e-32
Identities = 32/48 (66%), Positives = 34/48 (70%)
Frame = +2
Query: 115 GLKQNRQPTEFFCKTLTASDTSTHGGFSVPRRAAEKIFPPLDYSMQPP 162
G + FCKTLT SDTSTHGGFSVPRRAAE FPPLDY +Q P
Sbjct: 488 GGSPTKSTPHMFCKTLTVSDTSTHGGFSVPRRAAEDCFPPLDYKLQRP 631
>TC79906 similar to PIR|F86427|F86427 auxin response factor 6 (ARF6)
[imported] - Arabidopsis thaliana, partial (12%)
Length = 896
Score = 127 bits (319), Expect = 2e-29
Identities = 64/128 (50%), Positives = 88/128 (68%), Gaps = 1/128 (0%)
Frame = +1
Query: 981 VPNMTFNPGCSGDVGINDTGVLNNGLRANQ-TQRMRTYTKVQKRGSVGRCIDVTRYKGYD 1039
V + + NPG + VG ++G + Q + +T+ KV K GS GR +D+T++ Y
Sbjct: 82 VLDSSLNPGLTHGVG--ESGFMQTPENGGQGNPQNKTFVKVYKSGSFGRSLDITKFSNYP 255
Query: 1040 ELRNDLARMFGIEGQLEDPQRTDWKLVYVDHENDILLVGDDPWEEFVSCVQSIKILSYTE 1099
ELR++LARMFG+EG+LEDP R+ W+LV+VD END+LL+GD PW EFV+ V IKILS E
Sbjct: 256 ELRSELARMFGLEGELEDPVRSGWQLVFVDQENDVLLLGDGPWPEFVNSVWCIKILSPQE 435
Query: 1100 VQQMSLDG 1107
VQQM +G
Sbjct: 436 VQQMGTNG 459
>TC91214 similar to GP|2708484|gb|AAB92476.1| IAA24 {Arabidopsis thaliana},
partial (11%)
Length = 562
Score = 126 bits (316), Expect = 5e-29
Identities = 59/94 (62%), Positives = 76/94 (80%)
Frame = +3
Query: 1014 MRTYTKVQKRGSVGRCIDVTRYKGYDELRNDLARMFGIEGQLEDPQRTDWKLVYVDHEND 1073
+RTYTKVQK GSVGR IDVT +K Y+EL + MFG++G L D + + WKLVYVD+E+D
Sbjct: 144 IRTYTKVQKAGSVGRSIDVTTFKNYEELIRAIECMFGLDGLLNDTKGSGWKLVYVDYESD 323
Query: 1074 ILLVGDDPWEEFVSCVQSIKILSYTEVQQMSLDG 1107
+LLVGDDPWEEFV CV+ I+ILS +EV++MS +G
Sbjct: 324 VLLVGDDPWEEFVGCVRCIRILSPSEVKEMSEEG 425
>TC79570 similar to PIR|F86427|F86427 auxin response factor 6 (ARF6)
[imported] - Arabidopsis thaliana, partial (15%)
Length = 1143
Score = 124 bits (312), Expect = 2e-28
Identities = 122/371 (32%), Positives = 163/371 (43%), Gaps = 18/371 (4%)
Frame = +2
Query: 337 WDESTAGERPSRVSIWEIEPVVTPFYICPPPF-FRPKFPRQPGMPDDESDIENAFKRAMP 395
WDESTAGER RVS+WEIEP+ T F + P PF R K P PG+P ++ F P
Sbjct: 2 WDESTAGERQPRVSLWEIEPLTT-FPMYPSPFPLRLKRPWPPGLPSFHGLKDDDFGMNSP 178
Query: 396 --WLGD-DFGMKDASGSIFPGLSLVQWMSMQQNNQFSAAQSGCFPPMLSPNTLHSNLGTD 452
WL D D G++ + F G+ + WM + + Q + M + ++ T
Sbjct: 179 LLWLRDADRGLQTLN---FQGMGVNPWMQARLDPSMMNMQPDMYQAMAAAAL--QDMRTL 343
Query: 453 DPSK-----LLSFQAPALSAPSLQFNKPNLPNQINQLHQSPVSWPQQQQQQ---QQQQQQ 504
DPSK LL FQ P + N + NQ+ Q Q ++P+ Q+ Q Q Q Q
Sbjct: 344 DPSKQHPASLLQFQQPQNFSNG---NAALMQNQMLQQSQPHQAFPKNQESQHPSQSQAQT 514
Query: 505 QQQQLQLQHQQQQQQLQQQQQQQQQQQLQQQQQQQLQQQQSQQQQLQSLLQIPMNQLQQP 564
QQ Q LQHQ QQQQQQQ QQQQQQQ Q QQ QQQQ Q Q+ +Q
Sbjct: 515 QQFQQLLQHQHSFTNQNYHMQQQQQQQQQQQQQQQQQHQQQQQQQSQQQQQVVDHQQMSN 694
Query: 565 RQQQLPEPKNLPLPQQQLPQQLGQQSQKAIMSNGVVASNQITNQLVLQQQQLLSGGISPQ 624
+ + + P Q P Q AI S G + +N + + G S
Sbjct: 695 AVSTMSQFVSAPPSQSTQPMQ-------AISSIGHQQNFSDSNGNPVSPLHNMLGSFSND 853
Query: 625 QSIHSASKTTFPLTSLPQESQFLQQIDQQA------SLLQRQQLPQQTQLQQSSLHLLQQ 678
++ H + P + +P +S + + A S LPQ QL QS Q
Sbjct: 854 ETSHLLNFPR-PNSWVPVQSSTARPSKRVAVDPLLSSGASHYVLPQVEQLGQS-----QT 1015
Query: 679 NQPQRAPQQPP 689
Q A PP
Sbjct: 1016TMSQNAITLPP 1048
Score = 46.2 bits (108), Expect = 7e-05
Identities = 51/151 (33%), Positives = 62/151 (40%), Gaps = 9/151 (5%)
Frame = +2
Query: 654 ASLLQRQQLPQQTQLQQSSL---HLLQQNQPQRAPQQPPVATMSQQTPSEQQLHLQLLQK 710
ASLLQ QQ PQ ++L +LQQ+QP +A + Q PS+ Q Q Q+
Sbjct: 365 ASLLQFQQ-PQNFSNGNAALMQNQMLQQSQPHQAFPK----NQESQHPSQSQAQTQQFQQ 529
Query: 711 LQQHQQ----HQQLLSTSSPLLQPQLLQQQSTPQQNQQLTQLPASQHQTQQLGNNAFSTE 766
L QHQ + Q Q QQQ QQ QQ Q Q +NA ST
Sbjct: 530 LLQHQHSFTNQNYHMQQQQQQQQQQQQQQQQQHQQQQQQQSQQQQQVVDHQQMSNAVSTM 709
Query: 767 KVLNSNSFSSSLMQPHQL--PVNHPQNTQKS 795
S S S QP Q + H QN S
Sbjct: 710 SQFVSAPPSQS-TQPMQAISSIGHQQNFSDS 799
Score = 41.2 bits (95), Expect = 0.002
Identities = 61/249 (24%), Positives = 86/249 (34%), Gaps = 24/249 (9%)
Frame = +2
Query: 578 PQQQLPQQLGQQSQKAIMSNGVVASNQITNQLVLQQQQLLSGGISPQQSIHSASKTTFPL 637
P +Q P L Q Q SNG A Q NQ++ Q Q P Q+ ++ P
Sbjct: 347 PSKQHPASLLQFQQPQNFSNGNAALMQ--NQMLQQSQ--------PHQAFPKNQESQHPS 496
Query: 638 TSLPQESQFLQ------------------QIDQQASLLQRQQLPQQTQLQQSSLHLLQQN 679
S Q QF Q Q QQ Q+QQ QQ Q QQS QQ
Sbjct: 497 QSQAQTQQFQQLLQHQHSFTNQNYHMQQQQQQQQQQQQQQQQQHQQQQQQQSQ---QQQQ 667
Query: 680 QPQRAPQQPPVATMSQ---QTPSEQQLHLQLLQKLQQHQQHQQLLSTSSPLLQPQLLQQQ 736
V+TMSQ PS+ +Q + + Q S +P+ +
Sbjct: 668 VVDHQQMSNAVSTMSQFVSAPPSQSTQPMQAISSIGHQQNFSD--SNGNPVSPLHNMLGS 841
Query: 737 STPQQNQQLTQLPASQHQTQQLGNNAFSTEKVLNS---NSFSSSLMQPHQLPVNHPQNTQ 793
+ + L P + A +++V +S +S + P + Q T
Sbjct: 842 FSNDETSHLLNFPRPNSWVPVQSSTARPSKRVAVDPLLSSGASHYVLPQVEQLGQSQTTM 1021
Query: 794 KSLAITRAP 802
AIT P
Sbjct: 1022SQNAITLPP 1048
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.314 0.128 0.372
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 33,888,116
Number of Sequences: 36976
Number of extensions: 516587
Number of successful extensions: 11860
Number of sequences better than 10.0: 366
Number of HSP's better than 10.0 without gapping: 3651
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 6398
length of query: 1142
length of database: 9,014,727
effective HSP length: 107
effective length of query: 1035
effective length of database: 5,058,295
effective search space: 5235335325
effective search space used: 5235335325
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (22.0 bits)
S2: 64 (29.3 bits)
Lotus: description of TM0006.8


