

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0005.6
(566 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
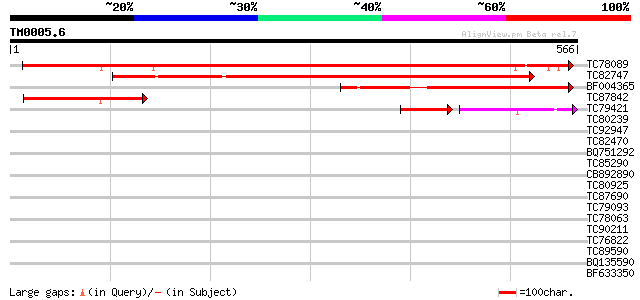
Sequences producing significant alignments: (bits) Value
TC78089 similar to GP|4522009|gb|AAD21782.1| unknown protein {Ar... 451 e-127
TC82747 similar to GP|9758202|dbj|BAB08676.1 emb|CAB82688.1~gene... 339 2e-93
BF004365 weakly similar to GP|21539547|gb| unknown protein {Arab... 243 1e-64
TC87842 similar to GP|9758120|dbj|BAB08592.1 gene_id:MCO15.18~pi... 141 8e-34
TC79421 similar to GP|11558256|emb|CAC17796. microtubule-associa... 77 5e-31
TC80239 similar to GP|7208779|emb|CAB76912.1 hypothetical protei... 36 0.045
TC92947 similar to GP|15294212|gb|AAK95283.1 AT5g17550/K10A8_30 ... 33 0.38
TC82470 homologue to GP|23497460|gb|AAN37003.1 hypothetical prot... 32 0.65
BQ751292 homologue to GP|13129509|gb| unknown protein {Oryza sat... 32 0.65
TC85290 similar to GP|7767653|gb|AAF69150.1| F27F5.2 {Arabidopsi... 32 0.85
CB892890 weakly similar to GP|9758141|dbj| disease resistance pr... 32 0.85
TC80925 similar to GP|22022575|gb|AAM83244.1 AT3g03330/T21P5_25 ... 32 0.85
TC87690 similar to GP|15028227|gb|AAK76610.1 putative oligopepti... 31 1.1
TC79093 similar to GP|4102190|gb|AAD01431.1| 35 kDa seed maturat... 29 4.2
TC78063 similar to GP|14194099|gb|AAK56244.1 AT3g52990/F8J2_160 ... 29 4.2
TC90211 similar to SP|P93768|PSD3_TOBAC Probable 26S proteasome ... 29 5.5
TC76822 similar to PIR|T06459|T06459 62K sucrose-binding protein... 29 5.5
TC89590 similar to GP|17473836|gb|AAL38344.1 unknown protein {Ar... 28 7.2
BQ135590 similar to GP|23499166|em hypothetical protein {Plasmod... 28 7.2
BF633350 similar to GP|17473640|gb| unknown protein {Arabidopsis... 28 7.2
>TC78089 similar to GP|4522009|gb|AAD21782.1| unknown protein {Arabidopsis
thaliana}, partial (85%)
Length = 2853
Score = 451 bits (1161), Expect = e-127
Identities = 245/582 (42%), Positives = 376/582 (64%), Gaps = 31/582 (5%)
Frame = +1
Query: 13 TTCASLLSQLQMIWDEIGESDSDRDNMLLQLEQECLDIYRRKVDATRKHKAELCQWLADA 72
TTC SLL +L+ IW++IGES+ D+D ML++LE+ECLD+YRRKVD KA Q LA
Sbjct: 718 TTCTSLLKELEQIWNDIGESEKDKDRMLMELERECLDVYRRKVDEAANIKARFHQSLAAK 897
Query: 73 EAELINLVSSLGESSSFS-----RGKGTLKQQLATIRPVLEDLRSKKDERVKEFSKIKSQ 127
EAE+ L+++LGE S + +LK++LA+I P++E+L+ KK+ER+K+ + IK+Q
Sbjct: 898 EAEIATLIAALGEHDMSSPIKMEKRSASLKEKLASITPLVEELKKKKEERLKQLADIKTQ 1077
Query: 128 ISQICVEIAGYEQPK-----SSSEEVDQSDLTFKKLGELKSHLQDLQNEKILRQQKVKSH 182
I +I EI G SS+ VD+ DL+ ++L E ++HL+ LQ EK R QKV
Sbjct: 1078 IEKISGEICGIHSVNDDVSVSSTGGVDEQDLSLRRLNEYQTHLRSLQKEKSDRLQKVLQC 1257
Query: 183 ISTISELSAVMSIDFWKTLSEIHPSLGDSSKGTPQSISNDTLASLTGYIHSLKQEKQQRL 242
++ + L V+ +DF +T+ ++HPSL + +ISN TL L I LK E++ R+
Sbjct: 1258 VNEVHSLCGVLGLDFGQTVDDVHPSLHGTQVEQSTNISNSTLEGLEKTILKLKTERKARI 1437
Query: 243 LKVQELTKFLVDLWDLIETPIDEQKAFNHVTRLISASVDEVSTHGCLSSDVIEQVEAEVQ 302
K++++ L +LW+L++T +E+ F +++ S E++ G LS+++IE+ AEV+
Sbjct: 1438 QKLKDIVANLFELWNLMDTSKEERNYFLRNNSIVTTSESEITERGALSTEIIEKASAEVE 1617
Query: 303 RLNALKASKMKDLVFKRQNELEEIYRGVHMDVDSEAARQILTSLIESGNIDLSDLLQSMD 362
RL LKAS+MK+LVFK+++ELEEI R H++ D+ A + ++LI+SG +D +LL +++
Sbjct: 1618 RLAELKASRMKELVFKKRSELEEICRLTHIEPDTSTAAEKASALIDSGLVDPCELLANIE 1797
Query: 363 DQIRQAKEQAQSRREILDRVEKWRFAAEEEKWLDEYERDENRYSAVRGAHKNLKRAEKAR 422
QI +AK++A SRRE+ DR++KW FA EEE WLDEY +D NRYSA RGAH NLKRAE+AR
Sbjct: 1798 AQIVKAKDEALSRREVTDRIDKWLFACEEENWLDEYSQDNNRYSAGRGAHINLKRAERAR 1977
Query: 423 IVVSKIPSIVENLTTKVKAWEMEKGIPFLYEKAPLLQSLEEYHVQRQLREEEKRKSREQK 482
+ ++KIP++V+NL +K AWE EK FLY+ L++ L++Y + RQ REEEKR+ R+QK
Sbjct: 1978 VTITKIPAMVDNLISKTLAWEDEKKTCFLYDGVRLVELLDDYKLTRQQREEEKRRQRDQK 2157
Query: 483 RLQEQHAVEQEALFGSRSATK------KPLGQSTTANTIVGTPNGRRMLTPSSRYGTSGG 536
++Q+ ++EA++GS+ + K K G N+I G NG + P+ R + G
Sbjct: 2158 KMQDLLLNQKEAIYGSKPSPKKTNSFRKTNGYRANGNSI-GYGNGNGSMPPTPRRNSMSG 2334
Query: 537 ------KERRESVRGN---------NIIPVNYVALPKDDSVS 563
R S R N + P+N+VA+PK+D++S
Sbjct: 2335 TTSELHTPRSYSGRHNGYFNEMRRLSTAPLNFVAIPKEDTMS 2460
>TC82747 similar to GP|9758202|dbj|BAB08676.1
emb|CAB82688.1~gene_id:K17N15.15~strong similarity to
unknown protein {Arabidopsis thaliana}, partial (45%)
Length = 1329
Score = 339 bits (869), Expect = 2e-93
Identities = 183/422 (43%), Positives = 270/422 (63%)
Frame = +3
Query: 103 IRPVLEDLRSKKDERVKEFSKIKSQISQICVEIAGYEQPKSSSEEVDQSDLTFKKLGELK 162
I P LE++ K R +F++++ QI + +EI G ++ S VD++DL+ +K +L
Sbjct: 6 ILPQLEEMIIYKFYRPNQFTEVQEQIQSLSMEIYGPKEYGPSV--VDETDLSLRKFEDLH 179
Query: 163 SHLQDLQNEKILRQQKVKSHISTISELSAVMSIDFWKTLSEIHPSLGDSSKGTPQSISND 222
L LQ+EKI R + V+ H+ T++ L V+ DF +T+ IHPSLGD P+S+SND
Sbjct: 180 RQLNALQSEKIGRLKTVQEHLCTLNSLCLVLGFDFKQTVLGIHPSLGDEG---PKSVSND 350
Query: 223 TLASLTGYIHSLKQEKQQRLLKVQELTKFLVDLWDLIETPIDEQKAFNHVTRLISASVDE 282
T+ L I L++ K QR+ K+Q+L +++LW+L++TPI+EQ+ F +VT I+A DE
Sbjct: 351 TIQQLAAAIQQLREIKLQRMQKLQDLATTMLELWNLMDTPIEEQQMFQNVTCNIAA*EDE 530
Query: 283 VSTHGCLSSDVIEQVEAEVQRLNALKASKMKDLVFKRQNELEEIYRGVHMDVDSEAARQI 342
++ LS D I VE EV RL LK+SKMK+LV K++ ELEEI R H+ + + A +
Sbjct: 531 ITEPNTLSEDFINCVEVEVSRLEELKSSKMKELVLKKRTELEEICRKTHLVPEIDGAVEY 710
Query: 343 LTSLIESGNIDLSDLLQSMDDQIRQAKEQAQSRREILDRVEKWRFAAEEEKWLDEYERDE 402
IESG +D + +L+ + Q+ Q KE+A R++IL++VEKW A EE WL+EY RD+
Sbjct: 711 AVEAIESGIVDPACVLEQFERQVAQVKEEALGRKDILEKVEKWLAACNEESWLEEYNRDD 890
Query: 403 NRYSAVRGAHKNLKRAEKARIVVSKIPSIVENLTTKVKAWEMEKGIPFLYEKAPLLQSLE 462
NRY+A RGAH LKRAEKAR +V+KIP +V+ LT+K AWE EKG F Y+ LL +E
Sbjct: 891 NRYNAGRGAHLTLKRAEKARGLVNKIPGMVDALTSKTIAWEKEKGFEFTYDGVRLLSMIE 1070
Query: 463 EYHVQRQLREEEKRKSREQKRLQEQHAVEQEALFGSRSATKKPLGQSTTANTIVGTPNGR 522
EY++ R+ +E+E+R+ R+ K+LQ Q EQEAL+GS+ + KP G R
Sbjct: 1071EYNILREEKEQERRRQRDLKKLQGQMIAEQEALYGSKPSPSKPQSVKKGPRMSTGGAASR 1250
Query: 523 RM 524
R+
Sbjct: 1251RV 1256
>BF004365 weakly similar to GP|21539547|gb| unknown protein {Arabidopsis
thaliana}, partial (6%)
Length = 649
Score = 243 bits (620), Expect = 1e-64
Identities = 139/233 (59%), Positives = 162/233 (68%)
Frame = +1
Query: 331 HMDVDSEAARQILTSLIESGNIDLSDLLQSMDDQIRQAKEQAQSRREILDRVEKWRFAAE 390
HMD+D++AARQILT LI GNID+S+LLQ MD QIR+AKEQAQSRR+I+DRV +W+ AAE
Sbjct: 4 HMDMDAKAARQILTDLI--GNIDMSELLQGMDKQIRKAKEQAQSRRDIIDRVAEWKSAAE 177
Query: 391 EEKWLDEYERDENRYSAVRGAHKNLKRAEKARIVVSKIPSIVENLTTKVKAWEMEKGIPF 450
EEKWLDEYER RAEKARI+V++IPS+VENLTTKVK WE + F
Sbjct: 178 EEKWLDEYER----------------RAEKARILVTQIPSMVENLTTKVKTWETNEEKSF 309
Query: 451 LYEKAPLLQSLEEYHVQRQLREEEKRKSREQKRLQEQHAVEQEALFGSRSATKKPLGQST 510
LYEK PLL SL EY+VQRQLREEEKRKSREQK++ EQ AVEQEA+ GS SATKKP
Sbjct: 310 LYEKVPLLNSLHEYNVQRQLREEEKRKSREQKQVNEQLAVEQEAVSGSGSATKKPSEPEH 489
Query: 511 TANTIVGTPNGRRMLTPSSRYGTSGGKERRESVRGNNIIPVNYVALPKDDSVS 563
+ + N G K + R NNI PVNYV+LPK DSVS
Sbjct: 490 SC*HHIWDTNWALNAYSFKPLWNLGYKGSQRKCRVNNITPVNYVSLPKVDSVS 648
>TC87842 similar to GP|9758120|dbj|BAB08592.1
gene_id:MCO15.18~pir||T04799~strong similarity to
unknown protein {Arabidopsis thaliana}, partial (31%)
Length = 835
Score = 141 bits (355), Expect = 8e-34
Identities = 71/127 (55%), Positives = 92/127 (71%), Gaps = 3/127 (2%)
Frame = +2
Query: 14 TCASLLSQLQMIWDEIGESDSDRDNMLLQLEQECLDIYRRKVDATRKHKAELCQWLADAE 73
TC SLL +LQ IWDE+GESD +RD MLLQLEQECLD+Y+RKV+ K +A+L Q L+DA+
Sbjct: 296 TCGSLLKKLQEIWDEVGESDEERDKMLLQLEQECLDVYKRKVEHAAKSRAQLLQALSDAK 475
Query: 74 AELINLVSSLGESSSF---SRGKGTLKQQLATIRPVLEDLRSKKDERVKEFSKIKSQISQ 130
EL L+S+LGE GT+K+QLA I P LE L +K+ER+K+FS ++SQI +
Sbjct: 476 LELSTLLSALGEKGFAGIPDNTSGTIKEQLAAIAPALEQLWQQKEERIKDFSDVQSQIQK 655
Query: 131 ICVEIAG 137
IC EI G
Sbjct: 656 ICGEITG 676
>TC79421 similar to GP|11558256|emb|CAC17796. microtubule-associated protein
MAP65-1c {Nicotiana tabacum}, partial (30%)
Length = 1030
Score = 77.4 bits (189), Expect(2) = 5e-31
Identities = 33/52 (63%), Positives = 43/52 (82%)
Frame = +2
Query: 391 EEKWLDEYERDENRYSAVRGAHKNLKRAEKARIVVSKIPSIVENLTTKVKAW 442
EE WL++Y RD+NRY+A RGAH LKRAEKARI+V+KIP++V+ L K +AW
Sbjct: 29 EESWLEDYNRDDNRYNASRGAHLTLKRAEKARILVNKIPALVDTLVAKTRAW 184
Score = 75.5 bits (184), Expect(2) = 5e-31
Identities = 47/123 (38%), Positives = 64/123 (51%), Gaps = 6/123 (4%)
Frame = +3
Query: 450 FLYEKAPLLQSLEEYHVQRQLREEEKRKSREQKRLQEQHAVEQEALFGSRSATKKP---- 505
F Y+ PLL L+EY + R REEEKR+ R+QK+ QE + E EA FGS+ + +P
Sbjct: 207 FTYDGVPLLAMLDEYAMLRHEREEEKRRMRDQKKFQELQSTEPEAPFGSKPSPARPVSAA 386
Query: 506 --LGQSTTANTIVGTPNGRRMLTPSSRYGTSGGKERRESVRGNNIIPVNYVALPKDDSVS 563
+G GTPN R L S K+ + R + PVNYVA+ K+D+ S
Sbjct: 387 KKVGGPRANGGANGTPNRRLSLNAHQNGSRSISKDGKRDNR--PVAPVNYVAMSKEDAAS 560
Query: 564 RGS 566
S
Sbjct: 561 HVS 569
>TC80239 similar to GP|7208779|emb|CAB76912.1 hypothetical protein {Cicer
arietinum}, partial (43%)
Length = 930
Score = 35.8 bits (81), Expect = 0.045
Identities = 29/140 (20%), Positives = 65/140 (45%)
Frame = +3
Query: 380 DRVEKWRFAAEEEKWLDEYERDENRYSAVRGAHKNLKRAEKARIVVSKIPSIVENLTTKV 439
DRV +W ++EEK +E R + + A + K+ E+A++ K +E +
Sbjct: 18 DRVSRWGDESDEEKEPNEPVRTKEEDEQILKAEQARKKEEEAKL---KEKKRLEEIEKAK 188
Query: 440 KAWEMEKGIPFLYEKAPLLQSLEEYHVQRQLREEEKRKSREQKRLQEQHAVEQEALFGSR 499
+A + +K ++ L ++ +E + + RE+ +K ++K + + AVE +
Sbjct: 189 EALQRKKRNAEKAQQRALYKAQKEAEQKEKEREKRAKKKGKRKTVTTEDAVENTEQDAAA 368
Query: 500 SATKKPLGQSTTANTIVGTP 519
S + + L ++ + + P
Sbjct: 369 SPSSETLTRTLEESDQIEKP 428
>TC92947 similar to GP|15294212|gb|AAK95283.1 AT5g17550/K10A8_30
{Arabidopsis thaliana}, partial (84%)
Length = 792
Score = 32.7 bits (73), Expect = 0.38
Identities = 15/65 (23%), Positives = 35/65 (53%)
Frame = +3
Query: 349 SGNIDLSDLLQSMDDQIRQAKEQAQSRREILDRVEKWRFAAEEEKWLDEYERDENRYSAV 408
+G+ D+ ++++M Q+ + A+ +EI +R KW + +EY+R ++Y +
Sbjct: 435 AGSQDMESIVETMMQQLLSKEILAEPMKEIGERYPKWLEEHKANLSKEEYDRYSHQYELI 614
Query: 409 RGAHK 413
R ++
Sbjct: 615 RNLNE 629
>TC82470 homologue to GP|23497460|gb|AAN37003.1 hypothetical protein
{Plasmodium falciparum 3D7}, partial (8%)
Length = 1064
Score = 32.0 bits (71), Expect = 0.65
Identities = 21/84 (25%), Positives = 38/84 (45%), Gaps = 1/84 (1%)
Frame = +2
Query: 85 ESSSFSRGKGTLKQQLATIRPVLEDLRSKKDERVKEFSKIKSQISQICVEIAGYEQPKSS 144
+ S + TLK++L + ED+RSK + IK I + C ++ E K
Sbjct: 86 DDMSTDKSFSTLKEKLVLVEKSFEDIRSKTQVEERRLQSIKRDIEECCEDL---ENKKKE 256
Query: 145 SEEVDQSDLTFKKL-GELKSHLQD 167
+V + KK+ G++ ++D
Sbjct: 257 IRDVGRIIEARKKMQGKIDECVKD 328
>BQ751292 homologue to GP|13129509|gb| unknown protein {Oryza sativa
(japonica cultivar-group)}, partial (10%)
Length = 668
Score = 32.0 bits (71), Expect = 0.65
Identities = 33/140 (23%), Positives = 62/140 (43%), Gaps = 1/140 (0%)
Frame = -2
Query: 27 DEIGESDSDRDNMLLQLEQECLDIYRRKVDATRKHKAELC-QWLADAEAELINLVSSLGE 85
D GESD R + QE +++ R + + + L+ E + V S+ E
Sbjct: 532 DGDGESDEVRFERRIAEAQERQKRLNERLENVRTYVGKAATRELSVKERAFVEEVKSM-E 356
Query: 86 SSSFSRGKGTLKQQLATIRPVLEDLRSKKDERVKEFSKIKSQISQICVEIAGYEQPKSSS 145
+S G +Q ++ +ED++ +DE V + K++ + V+ + P SS
Sbjct: 355 ASLVGSENGAAARQQNQLKKRIEDVQRLRDELVADVEKLQKPVDGKDVQ----DSPSRSS 188
Query: 146 EEVDQSDLTFKKLGELKSHL 165
E S++ KL +++S L
Sbjct: 187 ELKIPSEIRKAKLQQVRSLL 128
>TC85290 similar to GP|7767653|gb|AAF69150.1| F27F5.2 {Arabidopsis
thaliana}, partial (32%)
Length = 1558
Score = 31.6 bits (70), Expect = 0.85
Identities = 55/293 (18%), Positives = 115/293 (38%), Gaps = 21/293 (7%)
Frame = +3
Query: 221 NDTLASLTGYIHSL-KQEKQQRLLKVQELTKFLVDLWDLIETPIDEQKAFNHVT-----R 274
ND L YI L K+E++Q+ ++ + + + D ++E A +T R
Sbjct: 54 NDRLLVFQDYIRDLEKEEEEQKRIQKERVRRGERKNRDAFRKLLEEHIADGVLTAKTQWR 233
Query: 275 LISASVDEV--------STHGCLSSDVIEQVEAEVQRLNALKASKMKDLVFKRQNELEEI 326
V E+ +T G D+ E V +++ + +KD++ + ++
Sbjct: 234 DYCLKVKELPQYQAVASNTSGSTPKDLFEDVFENLEKQYHEDKTLIKDIL--KSGKITVA 407
Query: 327 YRGVHMDVDSEAARQILTSLIESGNIDLSDLLQSMDDQIRQAKE-QAQSRREILDRVEKW 385
V D S + + I N+ L L + + ++ ++ +E +A+ R+ + D
Sbjct: 408 TTSVFEDFKSAVSEEATCKTISEINLKL--LFEELLERAKEKEEKEAKKRQRLADDFTNL 581
Query: 386 RFAAEEEKWLDEYER------DENRYSAVRGAHKNLKRAEKARIVVSKIPSIVENLTTKV 439
+ ++ +E D Y ++ S I E T +
Sbjct: 582 LYTLKDIITSSTWEECKALFEDTQEYISIGNE--------------SYSKEIFEEYITYL 719
Query: 440 KAWEMEKGIPFLYEKAPLLQSLEEYHVQRQLREEEKRKSREQKRLQEQHAVEQ 492
K EK EKA + EE +++ ++EK + RE+++ +E+H ++
Sbjct: 720 KEKAKEKERKREEEKAKKEKEREEKEKRKEKEKKEKEREREKEKSKERHKKDE 878
>CB892890 weakly similar to GP|9758141|dbj| disease resistance protein-like
{Arabidopsis thaliana}, partial (4%)
Length = 852
Score = 31.6 bits (70), Expect = 0.85
Identities = 28/103 (27%), Positives = 51/103 (49%), Gaps = 4/103 (3%)
Frame = +2
Query: 233 SLKQEKQQRLLKVQELTKFLVDLWDLIETPIDEQKAFNHVTRLISASVDEVSTHGCLSSD 292
S+ Q +Q+ EL K L+ +D+ E D+Q N + R +S S D++ T + +
Sbjct: 287 SIHQARQEPF----ELRKRLI--FDVNENSWDQQNQQNTIARTLSISPDKILTSDWSNVE 448
Query: 293 VIEQVEAEVQRLNALKASKMKDLVFKRQNELEEI----YRGVH 331
I+QVE + L+ K + + K+ +L+ + Y+G H
Sbjct: 449 KIKQVEVLILNLHTEKYTLPE--CIKKMTKLKVLIITNYKGFH 571
>TC80925 similar to GP|22022575|gb|AAM83244.1 AT3g03330/T21P5_25
{Arabidopsis thaliana}, partial (70%)
Length = 854
Score = 31.6 bits (70), Expect = 0.85
Identities = 55/188 (29%), Positives = 80/188 (42%), Gaps = 7/188 (3%)
Frame = +2
Query: 65 LCQWLADAEAELINLVSSLGESSSFSRGKGTLKQQLATIRPVLE-DLRSKKDERVKEFSK 123
L Q LA A+LI S + + +R K LK + A +L DL S +D K +
Sbjct: 206 LAQQLASLGAKLI---LSARDEADLNRVKSQLKGKHADEAKILPLDLTSGEDSLRKVVDE 376
Query: 124 IKSQISQICVEI----AGYEQPKSSSEEVDQSDL--TFKKLGELKSHLQDLQNEKILRQQ 177
+S V+ A YE+PKSS +V + L TF L L +LR+
Sbjct: 377 AESLFPDSGVDYMIHNAAYERPKSSVLDVTEESLKATFDVNVFGTITLTRLLTPFMLRRG 556
Query: 178 KVKSHISTISELSAVMSIDFWKTLSEIHPSLGDSSKGTPQSISNDTLASLTGYIHSLKQE 237
K H VMS KT P+ G Q++ + + +L GY HSL+ E
Sbjct: 557 K--GHF-------VVMSSAAGKT-----PAPG-------QAVYSASKYALNGYFHSLRSE 673
Query: 238 KQQRLLKV 245
Q+ ++V
Sbjct: 674 LCQKGIQV 697
>TC87690 similar to GP|15028227|gb|AAK76610.1 putative oligopeptidase A
{Arabidopsis thaliana}, partial (30%)
Length = 1019
Score = 31.2 bits (69), Expect = 1.1
Identities = 25/114 (21%), Positives = 53/114 (45%)
Frame = +2
Query: 156 KKLGELKSHLQDLQNEKILRQQKVKSHISTISELSAVMSIDFWKTLSEIHPSLGDSSKGT 215
K EL+S ++D+Q EK+ Q ++ + A+ WKTLS+ + ++
Sbjct: 500 KDNSELRSAIEDVQAEKVKFQLRLGQSKPLYNAFKAIQDSPDWKTLSDARKRIVENQ--I 673
Query: 216 PQSISNDTLASLTGYIHSLKQEKQQRLLKVQELTKFLVDLWDLIETPIDEQKAF 269
+++ N SL+ +K+++ K+++ + L + + E +D K F
Sbjct: 674 KEAVLNGV---------SLEDDKREQFNKIEQELERLSEKFG--ENVLDATKKF 802
>TC79093 similar to GP|4102190|gb|AAD01431.1| 35 kDa seed maturation protein
{Glycine max}, partial (57%)
Length = 1127
Score = 29.3 bits (64), Expect = 4.2
Identities = 35/148 (23%), Positives = 61/148 (40%), Gaps = 8/148 (5%)
Frame = +3
Query: 28 EIGESDSDRDNMLLQLEQECLDIYRRKV----DATRKHKAELCQWLADAEAELINLVSS- 82
E GE D + Q +E D K D T E + A+ E + ++
Sbjct: 429 EAGEKARDTTDYAAQKTRENADYAAEKARETKDTTANKAGEYTNYAAEKAREAKDTTANK 608
Query: 83 LGESSSFSRGKGTLKQQLATIRPVLE--DLRSKKDERVKEFSKIKS-QISQICVEIAGYE 139
GE ++++ K + AT+ E + + K ++ K+ + K+ + V AG
Sbjct: 609 AGEYTNYAAEKAKEAKD-ATVNKAGEYTNYAADKAKQAKDATVQKAGEAKDATVNKAGEY 785
Query: 140 QPKSSSEEVDQSDLTFKKLGELKSHLQD 167
+ ++ + V+ D T KLGELK D
Sbjct: 786 KDYTAEKAVEAKDTTVGKLGELKDSAAD 869
>TC78063 similar to GP|14194099|gb|AAK56244.1 AT3g52990/F8J2_160 {Arabidopsis
thaliana}, complete
Length = 1843
Score = 29.3 bits (64), Expect = 4.2
Identities = 27/108 (25%), Positives = 47/108 (43%), Gaps = 4/108 (3%)
Frame = +2
Query: 126 SQISQICVEIAGYEQPKSSSEEVDQSDLTFKK----LGELKSHLQDLQNEKILRQQKVKS 181
S + +IC E +E+V DL FKK +GE SH++ + + + KVK+
Sbjct: 1088 STVGKICAE----------AEKVYNQDLYFKKTVKFVGEPMSHMESIASSAVRAAVKVKA 1237
Query: 182 HISTISELSAVMSIDFWKTLSEIHPSLGDSSKGTPQSISNDTLASLTG 229
+ S + + L++ P++ S PQ +N + TG
Sbjct: 1238 SVIICFTSSGRAA----RLLAKYRPTMPVISVVIPQLKTNQLRWTFTG 1369
>TC90211 similar to SP|P93768|PSD3_TOBAC Probable 26S proteasome non-ATPase
regulatory subunit 3 (26S proteasome subunit S3),
partial (57%)
Length = 1009
Score = 28.9 bits (63), Expect = 5.5
Identities = 37/146 (25%), Positives = 64/146 (43%), Gaps = 5/146 (3%)
Frame = +1
Query: 154 TFKKLGELKSHLQDLQNEKILRQ--QKVKSHISTISELSAVMSIDFWKTLSEIHPSLGD- 210
T + L E+ S ++ N K +R+ + V+ ++ S+L+A + F + H +L
Sbjct: 127 TLQHLKEIASIIETGSNLKEVRKISRAVRLTLALRSKLTAPVLSSFLHYVLPPHSTLSSY 306
Query: 211 --SSKGTPQSISNDTLASLTGYIHSLKQEKQQRLLKVQELTKFLVDLWDLIETPIDEQKA 268
+SKG + DT S + Q + LL E+ + + L LI+ QK
Sbjct: 307 LPNSKGGDHEMEVDTATS------AAIQTPAKHLLPELEIYCYFLVLLFLID-----QKR 453
Query: 269 FNHVTRLISASVDEVSTHGCLSSDVI 294
+N SAS+D + + DVI
Sbjct: 454 YNEAKACSSASIDRLKNLNRRTVDVI 531
>TC76822 similar to PIR|T06459|T06459 62K sucrose-binding protein homolog -
garden pea, partial (97%)
Length = 1693
Score = 28.9 bits (63), Expect = 5.5
Identities = 14/41 (34%), Positives = 22/41 (53%)
Frame = +2
Query: 453 EKAPLLQSLEEYHVQRQLREEEKRKSREQKRLQEQHAVEQE 493
+K + E+YH +Q E EKR+ + Q+ + QH E E
Sbjct: 167 DKGICMDKCEDYHRMKQ--EREKRQHQHQREHEHQHEREHE 283
>TC89590 similar to GP|17473836|gb|AAL38344.1 unknown protein {Arabidopsis
thaliana}, partial (30%)
Length = 1754
Score = 28.5 bits (62), Expect = 7.2
Identities = 34/146 (23%), Positives = 66/146 (44%), Gaps = 11/146 (7%)
Frame = +3
Query: 242 LLKVQELTKFLVDLWDLIETPIDEQKAFNHVTRLISASVDEVSTHGCLSSDVIEQ----- 296
L ++ ++DL D++ + E+ ++ + + S +E + G ++ D++E
Sbjct: 198 LADAYDIHNHILDLKDML---LKEKMDYHSLLK----SANETAEPGNMTLDILELNRLRR 356
Query: 297 ---VEAEV--QRLNALKASKMKDLVFKRQNELEEIYRGVHMDVDSEAARQILTSLIESGN 351
+ + V RLN+L + K L K + E E G + VDS Q S +E N
Sbjct: 357 SLLIGSHVWDHRLNSLDSLIKKSLSSKVKQETESFADGKELRVDSLHKDQSFDSWLEQNN 536
Query: 352 IDLSDLLQSMDD-QIRQAKEQAQSRR 376
S +S + ++ + +Q +SRR
Sbjct: 537 SQPSKGHESHESHKLVEPDDQPKSRR 614
>BQ135590 similar to GP|23499166|em hypothetical protein {Plasmodium
falciparum 3D7}, partial (0%)
Length = 1238
Score = 28.5 bits (62), Expect = 7.2
Identities = 24/110 (21%), Positives = 49/110 (43%), Gaps = 2/110 (1%)
Frame = +3
Query: 436 TTKVKAWEMEKGIPFLYEKAPLLQSLEEYHVQRQLREEEKRKSREQKRLQEQHAVEQEAL 495
T K +K PF Y +P QSL + + ++K+K + + +++ +
Sbjct: 3 TPTKKIISKKKNFPFFYTSSPP*QSLPFFPKLYSPQLKKKKKIYYKNKTKKERGIPTGWG 182
Query: 496 FGSR--SATKKPLGQSTTANTIVGTPNGRRMLTPSSRYGTSGGKERRESV 543
G + +K+ L Q +T+ ++ +G+ ++ R GG RR+ V
Sbjct: 183 VGVHCVALSKQLLSQLSTSLAVICGVDGKWRVSGGKRPRERGGLRRRKYV 332
>BF633350 similar to GP|17473640|gb| unknown protein {Arabidopsis thaliana},
partial (15%)
Length = 513
Score = 28.5 bits (62), Expect = 7.2
Identities = 17/55 (30%), Positives = 30/55 (53%)
Frame = +2
Query: 261 TPIDEQKAFNHVTRLISASVDEVSTHGCLSSDVIEQVEAEVQRLNALKASKMKDL 315
T ++ K + +ISASV ++ H S ++ ++ + +NAL AS+ KDL
Sbjct: 50 TSMERHKEIDAFDEVISASVKKIHEHLKRRSFLLGFSQSPAEFINALIASQSKDL 214
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.312 0.128 0.347
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 14,176,610
Number of Sequences: 36976
Number of extensions: 160810
Number of successful extensions: 800
Number of sequences better than 10.0: 45
Number of HSP's better than 10.0 without gapping: 788
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 791
length of query: 566
length of database: 9,014,727
effective HSP length: 101
effective length of query: 465
effective length of database: 5,280,151
effective search space: 2455270215
effective search space used: 2455270215
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.9 bits)
S2: 61 (28.1 bits)
Lotus: description of TM0005.6


