

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0005.4
(283 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
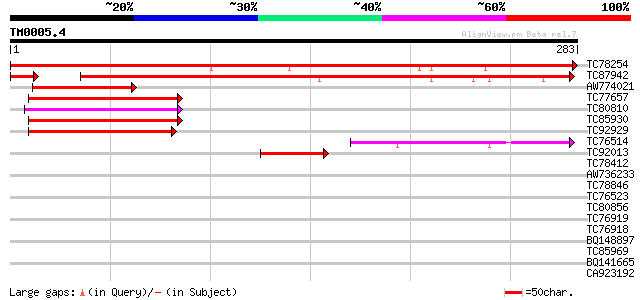
Score E
Sequences producing significant alignments: (bits) Value
TC78254 similar to GP|15795167|dbj|BAB03155. gene_id:MQC3.30~unk... 418 e-117
TC87942 similar to GP|15795167|dbj|BAB03155. gene_id:MQC3.30~unk... 362 e-102
AW774021 homologue to GP|15795167|dbj gene_id:MQC3.30~unknown pr... 97 9e-21
TC77657 homologue to PIR|E86221|E86221 hypothetical protein [imp... 59 2e-09
TC80810 homologue to PIR|T46203|T46203 transcription factor Hap5... 59 2e-09
TC85930 similar to GP|17065398|gb|AAL32853.1 Unknown protein {Ar... 59 2e-09
TC92929 similar to GP|18252953|gb|AAL62403.1 transcription facto... 57 6e-09
TC76514 homologue to PIR|T09648|T09648 nucleolin homolog nuM1 - ... 42 2e-04
TC92013 homologue to GP|11595557|emb|CAC18142. related to c-modu... 41 5e-04
TC78412 homologue to GP|17978995|gb|AAL47458.1 AT3g07590/MLP3_4 ... 40 0.001
AW736233 GP|21105451|gb small nuclear ribonucleoprotein D1 {Dani... 39 0.002
TC78846 homologue to PIR|B85036|B85036 small nuclear riboprotein... 39 0.002
TC76523 homologue to PIR|T09648|T09648 nucleolin homolog nuM1 - ... 38 0.005
TC80856 similar to GP|15706274|emb|CAC69997. HMG I/Y like protei... 37 0.007
TC76919 similar to GP|15724324|gb|AAL06555.1 At1g76010/T4O12_22 ... 37 0.011
TC76918 similar to GP|2231329|gb|AAB62000.1| bactinecin 11 {Ovis... 37 0.011
BQ148897 similar to GP|10177387|dbj gene_id:MNL12.7~unknown prot... 36 0.015
TC85969 similar to GP|21553908|gb|AAM62991.1 unknown {Arabidopsi... 35 0.043
BQ141665 similar to GP|18568267|gb putative polyprotein {Zea may... 34 0.056
CA923192 similar to GP|20196906|gb| expressed protein {Arabidops... 34 0.056
>TC78254 similar to GP|15795167|dbj|BAB03155. gene_id:MQC3.30~unknown
protein {Arabidopsis thaliana}, partial (51%)
Length = 1464
Score = 418 bits (1074), Expect = e-117
Identities = 219/305 (71%), Positives = 242/305 (78%), Gaps = 22/305 (7%)
Frame = +1
Query: 1 MKKKLDTRFPAARIKKIMQADEDVGKIALAVPVLVSKALELFLQDLCDRTYEITLQRGAK 60
MKKKLDTRFPAARIKKIMQADEDVGKIALAVPVLVSKALELFLQDLCDRTYEITLQRGAK
Sbjct: 262 MKKKLDTRFPAARIKKIMQADEDVGKIALAVPVLVSKALELFLQDLCDRTYEITLQRGAK 441
Query: 61 TMNSLHLKHCVQSYNVFDFLRDVVSKVPDYSHGHGHSEP-SADDRTIPKRRKAAGDDCND 119
TMNSLHLKHCVQSYNVFDFL+DVVSKVPDY HGHGH++ ADD+TIPKRRKAAGDD ND
Sbjct: 442 TMNSLHLKHCVQSYNVFDFLKDVVSKVPDYGHGHGHTDAGGADDQTIPKRRKAAGDDGND 621
Query: 120 SDEEVKRGKMPELSHTGST--GRGRGRGRGRGRGRGRTAVRETPHQAVEPEPCASVQQGI 177
SDEE KRGKM EL HT T GRGRGRGRGRGRGRGR RE HQ E EPC SVQQ
Sbjct: 622 SDEEAKRGKMLELGHTSPTGRGRGRGRGRGRGRGRGRAIQREGHHQETESEPCPSVQQVS 801
Query: 178 QHDTNTDMTIHDSSETKELPKENAAV-PAESAE-FHNLDLNANTNENEDKKASTTAKQEI 235
Q+ T+T++ I D +E+ ELPK N V PA++++ N+DLNAN NEN+DKKAST A +
Sbjct: 802 QNTTDTNVAILDGTESNELPKNNVVVLPADNSDSLLNIDLNANMNENDDKKASTVANLTV 981
Query: 236 S-----------------EPPTESQHEEIPGWSLSDVDKMAIDSVQLANLGTHIEEDEED 278
+ PP S HEEIPGWSLS+VDKMAIDS+QLA LGT ++ED+ED
Sbjct: 982 TIPEAANPEAANPPVSEPPPPDSSHHEEIPGWSLSEVDKMAIDSMQLAQLGTRMDEDDED 1161
Query: 279 YDEEG 283
YDEEG
Sbjct: 1162YDEEG 1176
>TC87942 similar to GP|15795167|dbj|BAB03155. gene_id:MQC3.30~unknown
protein {Arabidopsis thaliana}, partial (29%)
Length = 1222
Score = 362 bits (930), Expect(2) = e-102
Identities = 186/255 (72%), Positives = 206/255 (79%), Gaps = 8/255 (3%)
Frame = +1
Query: 36 SKALELFLQDLCDRTYEITLQRGAKTMNSLHLKHCVQSYNVFDFLRDVVSKVPDYSHGHG 95
SKALELFLQDLCDRTYEITLQRGAKTMN+LHLKHCVQSYNVFDFLRD+VS+VPDYSHGHG
Sbjct: 154 SKALELFLQDLCDRTYEITLQRGAKTMNALHLKHCVQSYNVFDFLRDIVSRVPDYSHGHG 333
Query: 96 HSEPSADDRTIPKRRKAAGDDCNDSDEEVKRGKMPELSHTGSTGRGRGRGRGRGRGRG-R 154
HSE ADDR +PKRRKAAGDDCNDSDEE KR KM EL HTGSTGRGRGRGRGRGRGRG R
Sbjct: 334 HSEAGADDRAVPKRRKAAGDDCNDSDEEAKRSKMLELGHTGSTGRGRGRGRGRGRGRGAR 513
Query: 155 TAVRETPHQAVEPEPCASVQQGIQHDTNTDMTIHDSSETKELPKENAAVPAESAE-FHNL 213
TA RET HQ VE EPC S+QQ + NT M I + SE KEL KEN A ES + N+
Sbjct: 514 TAERETLHQQVESEPCTSIQQTSKDVPNTSMAIDNGSEPKELSKENIAAHEESTQSLRNI 693
Query: 214 DLNANTNENEDKKASTT---AKQEISEP--PTESQHEEIPGWSLSDVDKMAIDSVQL-AN 267
DLNAN +ENE+K ++T + + EP T+ QHEEIPGWSLSDVDKMAID++QL AN
Sbjct: 694 DLNANLHENEEKNNTSTDIPTQAPLPEPAATTDMQHEEIPGWSLSDVDKMAIDTLQLAAN 873
Query: 268 LGTHIEEDEEDYDEE 282
LG ++EDEEDYDEE
Sbjct: 874 LGNRLDEDEEDYDEE 918
Score = 27.7 bits (60), Expect(2) = e-102
Identities = 11/14 (78%), Positives = 13/14 (92%)
Frame = +2
Query: 1 MKKKLDTRFPAARI 14
MKKKLDTRFPA ++
Sbjct: 122 MKKKLDTRFPAQKL 163
>AW774021 homologue to GP|15795167|dbj gene_id:MQC3.30~unknown protein
{Arabidopsis thaliana}, partial (21%)
Length = 704
Score = 96.7 bits (239), Expect = 9e-21
Identities = 48/52 (92%), Positives = 51/52 (97%)
Frame = +1
Query: 12 ARIKKIMQADEDVGKIALAVPVLVSKALELFLQDLCDRTYEITLQRGAKTMN 63
ARIKKIMQADED+GKIALAVP+LVSKALELFLQDLCDRTYEITL RGAKT+N
Sbjct: 547 ARIKKIMQADEDIGKIALAVPLLVSKALELFLQDLCDRTYEITLGRGAKTVN 702
Score = 27.3 bits (59), Expect = 6.8
Identities = 11/13 (84%), Positives = 12/13 (91%)
Frame = +2
Query: 1 MKKKLDTRFPAAR 13
M+KKLDTRFPA R
Sbjct: 95 MRKKLDTRFPAGR 133
>TC77657 homologue to PIR|E86221|E86221 hypothetical protein [imported] -
Arabidopsis thaliana, partial (62%)
Length = 1159
Score = 59.3 bits (142), Expect = 2e-09
Identities = 29/77 (37%), Positives = 48/77 (61%)
Frame = +2
Query: 10 PAARIKKIMQADEDVGKIALAVPVLVSKALELFLQDLCDRTYEITLQRGAKTMNSLHLKH 69
P ARIKKIM+ADEDV I+ PV+ ++A E+F+ +L R++ T + +T+ +
Sbjct: 410 PLARIKKIMKADEDVRMISAEAPVIFARACEMFILELTLRSWNHTEENKRRTLQKNDIAA 589
Query: 70 CVQSYNVFDFLRDVVSK 86
+ ++FDFL D+V +
Sbjct: 590 AITRTDIFDFLVDIVPR 640
>TC80810 homologue to PIR|T46203|T46203 transcription factor Hap5a -
Arabidopsis thaliana, partial (68%)
Length = 942
Score = 59.3 bits (142), Expect = 2e-09
Identities = 30/79 (37%), Positives = 48/79 (59%)
Frame = +3
Query: 8 RFPAARIKKIMQADEDVGKIALAVPVLVSKALELFLQDLCDRTYEITLQRGAKTMNSLHL 67
+ P ARIKKIM+ADEDV I+ P+L +KA ELF+ +L R++ + +T+ +
Sbjct: 183 QLPLARIKKIMKADEDVRMISAEAPILFAKACELFILELTIRSWLHAEENKRRTLQKNDI 362
Query: 68 KHCVQSYNVFDFLRDVVSK 86
+ ++FDFL D+V +
Sbjct: 363 AAAITRTDIFDFLVDIVPR 419
>TC85930 similar to GP|17065398|gb|AAL32853.1 Unknown protein {Arabidopsis
thaliana}, partial (62%)
Length = 1234
Score = 59.3 bits (142), Expect = 2e-09
Identities = 29/77 (37%), Positives = 48/77 (61%)
Frame = +3
Query: 10 PAARIKKIMQADEDVGKIALAVPVLVSKALELFLQDLCDRTYEITLQRGAKTMNSLHLKH 69
P ARIKKIM+ADEDV I+ PV+ ++A E+F+ +L R++ T + +T+ +
Sbjct: 432 PLARIKKIMKADEDVKMISAEAPVVFARACEMFILELTLRSWNHTEENKRRTLQKNDIAA 611
Query: 70 CVQSYNVFDFLRDVVSK 86
+ ++FDFL D+V +
Sbjct: 612 AITGTDIFDFLVDIVPR 662
>TC92929 similar to GP|18252953|gb|AAL62403.1 transcription factor putative
{Arabidopsis thaliana}, partial (58%)
Length = 598
Score = 57.4 bits (137), Expect = 6e-09
Identities = 30/74 (40%), Positives = 46/74 (61%)
Frame = +1
Query: 10 PAARIKKIMQADEDVGKIALAVPVLVSKALELFLQDLCDRTYEITLQRGAKTMNSLHLKH 69
P ARIKKIM+ADEDV I+ PV+ +KA E+F+ +L R++ T + +T+ +
Sbjct: 376 PLARIKKIMKADEDVRMISAEAPVVFAKACEMFILELTLRSWIHTEENKRRTLQKNDVAA 555
Query: 70 CVQSYNVFDFLRDV 83
+ +VFDFL D+
Sbjct: 556 AIARNDVFDFLGDI 597
>TC76514 homologue to PIR|T09648|T09648 nucleolin homolog nuM1 - alfalfa,
partial (61%)
Length = 1288
Score = 42.4 bits (98), Expect = 2e-04
Identities = 31/120 (25%), Positives = 57/120 (46%), Gaps = 8/120 (6%)
Frame = +3
Query: 171 ASVQQGIQHDTNTDMTIHDSSE-----TKELPKENAAVPAESAEFHNLDLNANTNENEDK 225
A ++ + +TD + + E K +P +N +VPA+ A+ + D ++ +++ EDK
Sbjct: 606 APAKKAASDEEDTDESSDEDEEDEKPAAKAVPSKNGSVPAKKADTESSDEDSESSDEEDK 785
Query: 226 KASTTAKQEISEP---PTESQHEEIPGWSLSDVDKMAIDSVQLANLGTHIEEDEEDYDEE 282
K + A + +S P S EE SD D+ A + A + ++D D D+E
Sbjct: 786 KPAAKASKNVSAPTKKAASSSDEESD--EESDEDEDAKPVSKPAAVAKKSKKDSSDSDDE 959
>TC92013 homologue to GP|11595557|emb|CAC18142. related to c-module-binding
factor {Neurospora crassa}, partial (2%)
Length = 1437
Score = 41.2 bits (95), Expect = 5e-04
Identities = 20/34 (58%), Positives = 22/34 (63%)
Frame = -2
Query: 126 RGKMPELSHTGSTGRGRGRGRGRGRGRGRTAVRE 159
RG+ G GRGRGRGRGRGRGRG +V E
Sbjct: 272 RGRGGGRGRGGGRGRGRGRGRGRGRGRGN*SVAE 171
Score = 37.4 bits (85), Expect = 0.007
Identities = 18/39 (46%), Positives = 22/39 (56%)
Frame = -2
Query: 116 DCNDSDEEVKRGKMPELSHTGSTGRGRGRGRGRGRGRGR 154
+ + S+ E G P + GRG GRGRG GRGRGR
Sbjct: 338 EISSSESESGIGGSPAMRGPSMRGRGGGRGRGGGRGRGR 222
Score = 34.7 bits (78), Expect = 0.043
Identities = 17/32 (53%), Positives = 19/32 (59%)
Frame = -2
Query: 139 GRGRGRGRGRGRGRGRTAVRETPHQAVEPEPC 170
GRGRGRGRGRGRG A E+ + E C
Sbjct: 227 GRGRGRGRGRGRGN*SVAEGESLMPRADCESC 132
>TC78412 homologue to GP|17978995|gb|AAL47458.1 AT3g07590/MLP3_4
{Arabidopsis thaliana}, complete
Length = 740
Score = 40.0 bits (92), Expect = 0.001
Identities = 20/33 (60%), Positives = 22/33 (66%)
Frame = +1
Query: 122 EEVKRGKMPELSHTGSTGRGRGRGRGRGRGRGR 154
EE R K + + GRGRGRGRGRGRGRGR
Sbjct: 349 EEAPRVKPKKPTAGKPLGRGRGRGRGRGRGRGR 447
>AW736233 GP|21105451|gb small nuclear ribonucleoprotein D1 {Danio rerio},
partial (14%)
Length = 510
Score = 39.3 bits (90), Expect = 0.002
Identities = 16/19 (84%), Positives = 18/19 (94%)
Frame = +1
Query: 137 STGRGRGRGRGRGRGRGRT 155
+ GRGRGRGRGRGRGRGR+
Sbjct: 223 NAGRGRGRGRGRGRGRGRS 279
Score = 35.0 bits (79), Expect = 0.033
Identities = 14/22 (63%), Positives = 16/22 (72%)
Frame = +1
Query: 134 HTGSTGRGRGRGRGRGRGRGRT 155
+ GRGRGRGRGRGRGR +
Sbjct: 220 YNAGRGRGRGRGRGRGRGRSNS 285
Score = 31.2 bits (69), Expect = 0.47
Identities = 14/24 (58%), Positives = 15/24 (62%)
Frame = +1
Query: 131 ELSHTGSTGRGRGRGRGRGRGRGR 154
E G+ GRGRGRGRGRGR
Sbjct: 193 ESQEVGTEYYNAGRGRGRGRGRGR 264
>TC78846 homologue to PIR|B85036|B85036 small nuclear riboprotein Sm-D1-like
protein [imported] - Arabidopsis thaliana, partial (98%)
Length = 642
Score = 38.9 bits (89), Expect = 0.002
Identities = 21/34 (61%), Positives = 23/34 (66%), Gaps = 1/34 (2%)
Frame = +1
Query: 122 EEVKRGKMPELSHTGST-GRGRGRGRGRGRGRGR 154
EE R K + + G GRGRGRGRGRGRGRGR
Sbjct: 352 EEAPRVKPKKPTAGGKPLGRGRGRGRGRGRGRGR 453
>TC76523 homologue to PIR|T09648|T09648 nucleolin homolog nuM1 - alfalfa,
partial (69%)
Length = 1615
Score = 37.7 bits (86), Expect = 0.005
Identities = 15/45 (33%), Positives = 29/45 (64%)
Frame = +2
Query: 194 KELPKENAAVPAESAEFHNLDLNANTNENEDKKASTTAKQEISEP 238
K +P +N +VPA+ A+ + D ++ +++ EDKK + A + +S P
Sbjct: 11 KAVPSKNGSVPAKKADTESSDEDSESSDEEDKKPAAKASKNVSAP 145
>TC80856 similar to GP|15706274|emb|CAC69997. HMG I/Y like protein {Glycine
max}, partial (35%)
Length = 1552
Score = 37.4 bits (85), Expect = 0.007
Identities = 39/142 (27%), Positives = 53/142 (36%), Gaps = 19/142 (13%)
Frame = +2
Query: 126 RGKMPELSHTGSTGRGRGRGRGRGRGR-------------GRTAVRETPHQAVEPEPCAS 172
RG+ G GRGRG GRGRGR R GR R TP A PE A
Sbjct: 1064 RGRGRGRGRGGVAGRGRGGGRGRGRPRLNLLAQSFGKRPVGRPKKRTTPATASAPENAA- 1240
Query: 173 VQQGIQHDTNTDMTIHDSSETKE-----LPKENAAVPAES-AEFHNLDLNANTNENEDKK 226
+ D H + KE P N P + A L++ + +E K
Sbjct: 1241 -----KEDDLKRKLEHFQGKVKESLAALRPHFNHESPVTAIAAIQELEILVAMDLSEPLK 1405
Query: 227 ASTTAKQEISEPPTESQHEEIP 248
T +Q+ + Q +++P
Sbjct: 1406 DETLPQQQNLSAQDQPQQQQLP 1471
>TC76919 similar to GP|15724324|gb|AAL06555.1 At1g76010/T4O12_22
{Arabidopsis thaliana}, partial (42%)
Length = 1207
Score = 36.6 bits (83), Expect = 0.011
Identities = 15/15 (100%), Positives = 15/15 (100%)
Frame = +3
Query: 139 GRGRGRGRGRGRGRG 153
GRGRGRGRGRGRGRG
Sbjct: 573 GRGRGRGRGRGRGRG 617
Score = 36.2 bits (82), Expect = 0.015
Identities = 15/15 (100%), Positives = 15/15 (100%)
Frame = +3
Query: 140 RGRGRGRGRGRGRGR 154
RGRGRGRGRGRGRGR
Sbjct: 570 RGRGRGRGRGRGRGR 614
Score = 32.7 bits (73), Expect = 0.16
Identities = 14/15 (93%), Positives = 14/15 (93%)
Frame = +3
Query: 140 RGRGRGRGRGRGRGR 154
R RGRGRGRGRGRGR
Sbjct: 564 RMRGRGRGRGRGRGR 608
Score = 32.7 bits (73), Expect = 0.16
Identities = 15/19 (78%), Positives = 15/19 (78%)
Frame = +3
Query: 140 RGRGRGRGRGRGRGRTAVR 158
RGRGR RGR RGRGR A R
Sbjct: 789 RGRGRERGRARGRGRDAGR 845
Score = 29.6 bits (65), Expect = 1.4
Identities = 13/16 (81%), Positives = 13/16 (81%)
Frame = +3
Query: 139 GRGRGRGRGRGRGRGR 154
G R RGRGRGRGRGR
Sbjct: 555 GSPRMRGRGRGRGRGR 602
Score = 28.9 bits (63), Expect = 2.3
Identities = 13/18 (72%), Positives = 13/18 (72%)
Frame = +3
Query: 139 GRGRGRGRGRGRGRGRTA 156
GR RGR RGRGR GR A
Sbjct: 798 GRERGRARGRGRDAGRGA 851
Score = 28.9 bits (63), Expect = 2.3
Identities = 12/14 (85%), Positives = 12/14 (85%)
Frame = +3
Query: 139 GRGRGRGRGRGRGR 152
GRGR RGR RGRGR
Sbjct: 792 GRGRERGRARGRGR 833
Score = 28.9 bits (63), Expect = 2.3
Identities = 12/15 (80%), Positives = 12/15 (80%)
Frame = +3
Query: 139 GRGRGRGRGRGRGRG 153
GRGRGRGRGRG G
Sbjct: 585 GRGRGRGRGRGMYNG 629
Score = 27.7 bits (60), Expect = 5.2
Identities = 12/16 (75%), Positives = 12/16 (75%)
Frame = +3
Query: 139 GRGRGRGRGRGRGRGR 154
G G R RGRGRGRGR
Sbjct: 549 GEGSPRMRGRGRGRGR 596
>TC76918 similar to GP|2231329|gb|AAB62000.1| bactinecin 11 {Ovis aries},
partial (13%)
Length = 576
Score = 36.6 bits (83), Expect = 0.011
Identities = 16/19 (84%), Positives = 17/19 (89%)
Frame = +1
Query: 140 RGRGRGRGRGRGRGRTAVR 158
RG+GRGRGRGRGRGR A R
Sbjct: 214 RGQGRGRGRGRGRGRDAGR 270
Score = 35.0 bits (79), Expect = 0.033
Identities = 15/18 (83%), Positives = 15/18 (83%)
Frame = +1
Query: 139 GRGRGRGRGRGRGRGRTA 156
GRGRGRGRGRGR GR A
Sbjct: 223 GRGRGRGRGRGRDAGRVA 276
Score = 34.3 bits (77), Expect = 0.056
Identities = 14/14 (100%), Positives = 14/14 (100%)
Frame = +1
Query: 140 RGRGRGRGRGRGRG 153
RGRGRGRGRGRGRG
Sbjct: 1 RGRGRGRGRGRGRG 42
Score = 32.3 bits (72), Expect = 0.21
Identities = 13/13 (100%), Positives = 13/13 (100%)
Frame = +1
Query: 139 GRGRGRGRGRGRG 151
GRGRGRGRGRGRG
Sbjct: 4 GRGRGRGRGRGRG 42
Score = 32.0 bits (71), Expect = 0.28
Identities = 13/13 (100%), Positives = 13/13 (100%)
Frame = +1
Query: 142 RGRGRGRGRGRGR 154
RGRGRGRGRGRGR
Sbjct: 1 RGRGRGRGRGRGR 39
Score = 28.9 bits (63), Expect = 2.3
Identities = 12/15 (80%), Positives = 12/15 (80%)
Frame = +1
Query: 139 GRGRGRGRGRGRGRG 153
GRGRGRGRGRG G
Sbjct: 10 GRGRGRGRGRGMYNG 54
Score = 26.9 bits (58), Expect = 8.9
Identities = 14/20 (70%), Positives = 14/20 (70%), Gaps = 1/20 (5%)
Frame = +1
Query: 136 GSTGRGRGRGRGRG-RGRGR 154
G GRGRGR GR RGRGR
Sbjct: 91 GYGGRGRGRAWGRAFRGRGR 150
>BQ148897 similar to GP|10177387|dbj gene_id:MNL12.7~unknown protein
{Arabidopsis thaliana}, partial (48%)
Length = 620
Score = 36.2 bits (82), Expect = 0.015
Identities = 16/42 (38%), Positives = 26/42 (61%)
Frame = +2
Query: 6 DTRFPAARIKKIMQADEDVGKIALAVPVLVSKALELFLQDLC 47
+ FP +R+KKI+ D+DV +++ LVS++ ELF C
Sbjct: 32 EPEFPKSRVKKIITLDKDVKRVSSEALFLVSRSTELFFTVTC 157
>TC85969 similar to GP|21553908|gb|AAM62991.1 unknown {Arabidopsis thaliana},
partial (31%)
Length = 2756
Score = 34.7 bits (78), Expect = 0.043
Identities = 34/116 (29%), Positives = 48/116 (41%), Gaps = 26/116 (22%)
Frame = -1
Query: 167 PEPCASVQQGIQHDTNTDMTIHDSSETKELPKENAAVPAESAEFHNLDLN--------AN 218
P P ASVQ QH + T+H S + + P AA+ A H+ + AN
Sbjct: 1112 PHPVASVQYH-QHQQPFEQTVHQSPKPFD-PAAAAAIVAAVEP*HHRKHHLQHAP**EAN 939
Query: 219 TNENEDKKASTTAK-------QEISEPPT-----------ESQHEEIPGWSLSDVD 256
+ + E KK+S K QE +E P E H+E+P W+L D
Sbjct: 938 SLKTETKKSSQRTKKSPS*ANQEQNEHPASSSYNSRKIQVEKHHQELPQWNLVQYD 771
>BQ141665 similar to GP|18568267|gb putative polyprotein {Zea mays}, partial
(1%)
Length = 1195
Score = 34.3 bits (77), Expect = 0.056
Identities = 27/92 (29%), Positives = 41/92 (44%), Gaps = 5/92 (5%)
Frame = +1
Query: 97 SEPSADDRTIPKRRKAAGDDCN-----DSDEEVKRGKMPELSHTGSTGRGRGRGRGRGRG 151
+E + RT PK +K N ++E +RGK G GR RG G GR
Sbjct: 190 TEGTPHTRTKPKEKKGRRRKKNMGRRTGGEQETERGK-----REGQRGRNE-RGPGPGRS 351
Query: 152 RGRTAVRETPHQAVEPEPCASVQQGIQHDTNT 183
R +R+TPH+ P+ ++G + N+
Sbjct: 352 EQRNQLRKTPHRWHWPKTTGR*REGTRGAGNS 447
>CA923192 similar to GP|20196906|gb| expressed protein {Arabidopsis
thaliana}, partial (16%)
Length = 762
Score = 34.3 bits (77), Expect = 0.056
Identities = 15/29 (51%), Positives = 20/29 (68%)
Frame = -3
Query: 126 RGKMPELSHTGSTGRGRGRGRGRGRGRGR 154
RG++ G+ GRG+G+G GRGRG GR
Sbjct: 421 RGRVSGGGRGGARGRGQGQGWGRGRGSGR 335
Score = 29.6 bits (65), Expect = 1.4
Identities = 13/23 (56%), Positives = 16/23 (69%)
Frame = -3
Query: 136 GSTGRGRGRGRGRGRGRGRTAVR 158
G G RGRG+G+G GRGR + R
Sbjct: 403 GGRGGARGRGQGQGWGRGRGSGR 335
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.310 0.129 0.366
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 7,336,963
Number of Sequences: 36976
Number of extensions: 92757
Number of successful extensions: 1285
Number of sequences better than 10.0: 105
Number of HSP's better than 10.0 without gapping: 729
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 992
length of query: 283
length of database: 9,014,727
effective HSP length: 95
effective length of query: 188
effective length of database: 5,502,007
effective search space: 1034377316
effective search space used: 1034377316
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.7 bits)
S2: 58 (26.9 bits)
Lotus: description of TM0005.4


