

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0005.13
(396 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
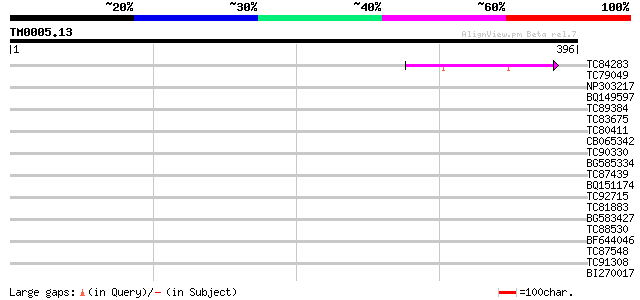
Score E
Sequences producing significant alignments: (bits) Value
TC84283 similar to GP|8777419|dbj|BAA97009.1 mitotic checkpoint ... 45 5e-05
TC79049 weakly similar to PIR|T10798|T10798 pherophorin-S - Volv... 35 0.051
NP303217 NP303217|AJ297721.1|CAC20726.1 putative repetitive prol... 35 0.066
BQ149597 similar to PIR|B84583|B84 hypothetical protein At2g1995... 35 0.066
TC89384 similar to GP|15450417|gb|AAK96502.1 At1g26300/F28B23_4 ... 33 0.25
TC83675 similar to PIR|T14321|T14321 nuclear matrix constituent ... 33 0.25
TC80411 weakly similar to PIR|T47635|T47635 probable protein - A... 33 0.25
CB065342 similar to SP|P34818|TRPI_ TrpBA operon transcriptional... 32 0.33
TC90330 weakly similar to PIR|D96593|D96593 myosin-like protein ... 32 0.33
BG585334 similar to GP|20161490|db beta-1 3 glucanase-like prote... 32 0.43
TC87439 weakly similar to GP|5139695|dbj|BAA81686.1 expressed in... 32 0.56
BQ151174 similar to GP|11036868|gb| PxORF73 peptide {Plutella xy... 31 0.73
TC92715 weakly similar to GP|23496006|gb|AAN35668.1 hypothetical... 31 0.73
TC81883 weakly similar to GP|7110148|gb|AAF36810.1| DNA repair-r... 31 0.95
BG583427 similar to PIR|T49091|T49 gamma response I protein - Ar... 30 1.2
TC88530 weakly similar to PIR|T01931|T01931 adenylyl cyclase - c... 30 1.2
BF644046 30 1.6
TC87548 similar to PIR|T47553|T47553 peroxiredoxin-like protein ... 30 2.1
TC91308 similar to GP|15450621|gb|AAK96582.1 At2g30530/T6B20.12 ... 30 2.1
BI270017 similar to GP|15450621|gb| At2g30530/T6B20.12 {Arabidop... 30 2.1
>TC84283 similar to GP|8777419|dbj|BAA97009.1 mitotic checkpoint
protein-like {Arabidopsis thaliana}, partial (21%)
Length = 829
Score = 45.1 bits (105), Expect = 5e-05
Identities = 34/112 (30%), Positives = 54/112 (47%), Gaps = 5/112 (4%)
Frame = +2
Query: 277 QDAEVLRSEVARLRALNAGFVEERK--AFLAQLKAKETSFAKELAEQSSRAEVSLEFRDR 334
+ AEVL+SE+ R+ A EER LA LK E A E SL +D
Sbjct: 29 EKAEVLKSEIKRIELTLAVVTEERNKLRILANLKNGEAGDESSSANPIQELESSLMKKDE 208
Query: 335 KISDLEKALGKLR---QEAQEESEAKVDLVASKDQTLSSMETKLEHLRAELA 383
I +LE L +LR EE + + + ++ + + S+E + + LR+E++
Sbjct: 209 YIKELESTLNELRVVNNRQHEEIKILNEKLHNEARRIKSLERESDRLRSEIS 364
>TC79049 weakly similar to PIR|T10798|T10798 pherophorin-S - Volvox carteri,
partial (6%)
Length = 957
Score = 35.0 bits (79), Expect = 0.051
Identities = 24/77 (31%), Positives = 35/77 (45%)
Frame = +1
Query: 161 NVSPHPVAATSSSDGDRAPVISAAFCGPEETPKPNQPGEPSLATPVSSPRAQDDSVDKPP 220
+V P P ++ SS + V F P +P P+ P PS P+ SP + PP
Sbjct: 721 SVPPQP*SSYSSPQFPSSQV--HQFFSPNPSPSPHSPLSPSSTLPLPSPTPR----PYPP 882
Query: 221 LSAKGFANREPPYSLSP 237
+K +N +PP S P
Sbjct: 883 CPSKPPSNPQPPSSSPP 933
>NP303217 NP303217|AJ297721.1|CAC20726.1 putative repetitive proline-rich
protein
Length = 525
Score = 34.7 bits (78), Expect = 0.066
Identities = 18/53 (33%), Positives = 26/53 (48%), Gaps = 4/53 (7%)
Frame = +1
Query: 192 PKPNQPGEPSLATPVSSPRAQDDSVDKPPLS----AKGFANREPPYSLSPTPA 240
PKP Q PS P+++P + + KPP++ K R PPY P P+
Sbjct: 355 PKPPQKKPPSRKRPINTPPNKKPPLPKPPVNKPPHKKPSNKRPPPYGNQPPPS 513
>BQ149597 similar to PIR|B84583|B84 hypothetical protein At2g19950 [imported]
- Arabidopsis thaliana, partial (23%)
Length = 634
Score = 34.7 bits (78), Expect = 0.066
Identities = 22/95 (23%), Positives = 50/95 (52%), Gaps = 13/95 (13%)
Frame = +3
Query: 302 AFLAQLKAKETSFAKELAEQSSRAEVSLEFRDRKISDLEKALGKLRQEAQEESEAKVDLV 361
A + ++ + T+ A EL ++ + EV ++++ D+E L + ++++ EE+ + +
Sbjct: 48 ARIQRIADERTAKATELEQKVALLEVECSSLNQELQDMEARLRREQKKSPEEANQVIQIQ 227
Query: 362 ASKDQT-------------LSSMETKLEHLRAELA 383
A +++ LSS+E +L+ LR E+A
Sbjct: 228 AWQEEVERARQGQREAENKLSSLEAELQKLRVEMA 332
>TC89384 similar to GP|15450417|gb|AAK96502.1 At1g26300/F28B23_4
{Arabidopsis thaliana}, partial (46%)
Length = 801
Score = 32.7 bits (73), Expect = 0.25
Identities = 15/50 (30%), Positives = 24/50 (48%)
Frame = +2
Query: 188 PEETPKPNQPGEPSLATPVSSPRAQDDSVDKPPLSAKGFANREPPYSLSP 237
P++ P P P L +P+SSP+ + + PP + PP S+ P
Sbjct: 167 PQKNPNL*PPPPPLLESPISSPKPPRNPKNSPPKLPRKQMRSNPPLSVKP 316
>TC83675 similar to PIR|T14321|T14321 nuclear matrix constituent protein 1 -
carrot, partial (17%)
Length = 905
Score = 32.7 bits (73), Expect = 0.25
Identities = 25/94 (26%), Positives = 41/94 (43%)
Frame = +3
Query: 297 VEERKAFLAQLKAKETSFAKELAEQSSRAEVSLEFRDRKISDLEKALGKLRQEAQEESEA 356
+ E K L Q KA E ++ +L + DLEKAL ++R E + A
Sbjct: 33 IVEVKDALEQEKAAHLFALSEAEKREENLRKALGVEKECVLDLEKALREMRSEHAKIKFA 212
Query: 357 KVDLVASKDQTLSSMETKLEHLRAELAKKDEALS 390
+A + ++S+E K + A+L D L+
Sbjct: 213 ADSKLAEANALIASVEEKSLEVEAKLRSADAKLA 314
>TC80411 weakly similar to PIR|T47635|T47635 probable protein - Arabidopsis
thaliana, partial (8%)
Length = 1045
Score = 32.7 bits (73), Expect = 0.25
Identities = 26/76 (34%), Positives = 34/76 (44%), Gaps = 3/76 (3%)
Frame = +3
Query: 187 GPEETPKPNQPGEPSLATPVSSPRAQDDSVDKPPLSAKGF--ANREP-PYSLSPTPAQVA 243
G EE P +P + PV P ++ D+ P+SAK A + P P + SP PAQV
Sbjct: 510 GTEEKPAQQKPVQQK---PVQQPSRKNQKKDEQPVSAKSPKPAVKSPKPATKSPKPAQVN 680
Query: 244 RMDEVVKQQGLDRVSE 259
R K SE
Sbjct: 681 RKPYSKKNNSESNPSE 728
>CB065342 similar to SP|P34818|TRPI_ TrpBA operon transcriptional activator.
[pv. syringae] {Pseudomonas syringae}, partial (31%)
Length = 631
Score = 32.3 bits (72), Expect = 0.33
Identities = 22/75 (29%), Positives = 37/75 (49%)
Frame = +3
Query: 321 QSSRAEVSLEFRDRKISDLEKALGKLRQEAQEESEAKVDLVASKDQTLSSMETKLEHLRA 380
++S + SLE + +I L KAL L+ +A EES+ + + S+ E+ L H R+
Sbjct: 15 RNSLRKASLESMESEIESLLKALENLKHDASEESQKSLKTIR------SNAESALRHSRS 176
Query: 381 ELAKKDEALSMMRTQ 395
L + E + Q
Sbjct: 177 LLGEAYEEVKTRTRQ 221
>TC90330 weakly similar to PIR|D96593|D96593 myosin-like protein
97843-94399 [imported] - Arabidopsis thaliana, partial
(36%)
Length = 1191
Score = 32.3 bits (72), Expect = 0.33
Identities = 45/172 (26%), Positives = 68/172 (39%), Gaps = 14/172 (8%)
Frame = +1
Query: 212 QDDSVDK-PPLSAKGFANREPPYSLSPTPAQVARMDEVVKQQGLDRVSEGAFATAFHAFH 270
QD V + PP+S++ E PY+ P P + +Q H
Sbjct: 55 QDTCVHREPPMSSRRGYPSEGPYARGPPP--------MPRQ---------------HIPP 165
Query: 271 HYKRSAQDAEVLRSEVARLRALNAGFVEERKAFLAQL-KAKETSFAKELAEQSSRAEVSL 329
H ++ EV +E+ RL A N +++R L AKE LA RAE +
Sbjct: 166 HPALLEEELEVQHAEMRRLVADNRRLIDDRMTLQRDLAAAKEELHRMNLAIGDIRAEHEM 345
Query: 330 EFRD------------RKISDLEKALGKLRQEAQEESEAKVDLVASKDQTLS 369
R+ R L+ + +LR E Q+ S K +L + K QTL+
Sbjct: 346 HSRELVDKNMKLESDLRSTEPLKNEVIQLRAEVQKLSSIKQEL-SGKVQTLT 498
>BG585334 similar to GP|20161490|db beta-1 3 glucanase-like protein {Oryza
sativa (japonica cultivar-group)}, partial (39%)
Length = 835
Score = 32.0 bits (71), Expect = 0.43
Identities = 16/55 (29%), Positives = 25/55 (45%), Gaps = 2/55 (3%)
Frame = +1
Query: 190 ETPKPNQP--GEPSLATPVSSPRAQDDSVDKPPLSAKGFANREPPYSLSPTPAQV 242
+T P P P+ +P S+P A + PP S ++N P + +PT V
Sbjct: 265 DTTSPTNPYSNPPTSTSPYSNPPASTNPYSNPPASTNPYSNPTSPTTTTPTSPTV 429
>TC87439 weakly similar to GP|5139695|dbj|BAA81686.1 expressed in cucumber
hypocotyls {Cucumis sativus}, partial (57%)
Length = 897
Score = 31.6 bits (70), Expect = 0.56
Identities = 22/65 (33%), Positives = 26/65 (39%), Gaps = 7/65 (10%)
Frame = +3
Query: 166 PVAATSSSDGDRAPVISAAFCGPEETPKPNQP-------GEPSLATPVSSPRAQDDSVDK 218
PVAA ++ APV S P +PK P P TPVS+P V
Sbjct: 168 PVAAPPTTPTTPAPVASPKSSPPATSPKAAAPTATPPAASSPPAVTPVSTPPPAPVPVKS 347
Query: 219 PPLSA 223
PP A
Sbjct: 348 PPTPA 362
>BQ151174 similar to GP|11036868|gb| PxORF73 peptide {Plutella xylostella
granulovirus}, partial (42%)
Length = 909
Score = 31.2 bits (69), Expect = 0.73
Identities = 18/64 (28%), Positives = 24/64 (37%)
Frame = +1
Query: 190 ETPKPNQPGEPSLATPVSSPRAQDDSVDKPPLSAKGFANREPPYSLSPTPAQVARMDEVV 249
+TPK N P P P +P KPP K EPP P P +
Sbjct: 199 QTPKNNTPPPPPHTPPDPTPPQPPPQPPKPPHHEKRPRTPEPPGGRPPPPTTPKQARPTT 378
Query: 250 KQQG 253
+++G
Sbjct: 379 QERG 390
>TC92715 weakly similar to GP|23496006|gb|AAN35668.1 hypothetical protein
{Plasmodium falciparum 3D7}, partial (2%)
Length = 537
Score = 31.2 bits (69), Expect = 0.73
Identities = 18/61 (29%), Positives = 28/61 (45%)
Frame = +1
Query: 177 RAPVISAAFCGPEETPKPNQPGEPSLATPVSSPRAQDDSVDKPPLSAKGFANREPPYSLS 236
R P ++ C P P + P + ++P PR++ S P S+ A R PP S +
Sbjct: 34 RHPSRRSSSCAPRARPSTS-PTTSTPSSPARRPRSRTSSSPPNPRSSSPSARRRPPASST 210
Query: 237 P 237
P
Sbjct: 211 P 213
>TC81883 weakly similar to GP|7110148|gb|AAF36810.1| DNA
repair-recombination protein {Arabidopsis thaliana},
partial (10%)
Length = 1057
Score = 30.8 bits (68), Expect = 0.95
Identities = 20/76 (26%), Positives = 40/76 (52%)
Frame = +1
Query: 320 EQSSRAEVSLEFRDRKISDLEKALGKLRQEAQEESEAKVDLVASKDQTLSSMETKLEHLR 379
E + R + ++ +K+ DLE LG E + E +++LVA + T ++ T+L +L
Sbjct: 454 EHADRIFLEIQTLQKKVEDLESELGCGGPEVRTLEEIQLELVALQG-TKDNLHTELTNLM 630
Query: 380 AELAKKDEALSMMRTQ 395
E D +S ++++
Sbjct: 631 DEQRNMDTDISGIQSR 678
>BG583427 similar to PIR|T49091|T49 gamma response I protein - Arabidopsis
thaliana, partial (3%)
Length = 730
Score = 30.4 bits (67), Expect = 1.2
Identities = 26/97 (26%), Positives = 44/97 (44%)
Frame = +1
Query: 286 VARLRALNAGFVEERKAFLAQLKAKETSFAKELAEQSSRAEVSLEFRDRKISDLEKALGK 345
V+ L L ++E K ++Q+ E F +L + + + R+I++LE L
Sbjct: 112 VSGLSTLLVASIQEAKDKISQI---EYVFCSQLYPNFQSTSKNSDSKRRRITELEHEL-- 276
Query: 346 LRQEAQEESEAKVDLVASKDQTLSSMETKLEHLRAEL 382
E + +DL+ SKD L S E K + A+L
Sbjct: 277 --DEGMTLQKNLLDLIRSKDSLLKSCEEKRKTAVAKL 381
>TC88530 weakly similar to PIR|T01931|T01931 adenylyl cyclase - common
tobacco, partial (57%)
Length = 1424
Score = 30.4 bits (67), Expect = 1.2
Identities = 16/38 (42%), Positives = 21/38 (55%)
Frame = +1
Query: 298 EERKAFLAQLKAKETSFAKELAEQSSRAEVSLEFRDRK 335
EERKAF A LK+ + +L + A VSLE R+
Sbjct: 1027 EERKAFAASLKSDILAMVNDLLDMGLEANVSLEDLSRR 1140
>BF644046
Length = 597
Score = 30.0 bits (66), Expect = 1.6
Identities = 13/24 (54%), Positives = 17/24 (70%)
Frame = +3
Query: 1 MSKAKSQLHPNSWAFLRCFEILCD 24
++ A SQLHPN AF+ FEI C+
Sbjct: 87 LNVASSQLHPNCCAFMCGFEISCE 158
>TC87548 similar to PIR|T47553|T47553 peroxiredoxin-like protein -
Arabidopsis thaliana, partial (76%)
Length = 911
Score = 29.6 bits (65), Expect = 2.1
Identities = 18/53 (33%), Positives = 24/53 (44%), Gaps = 3/53 (5%)
Frame = +3
Query: 192 PKPNQPGEPSLATPVSSPRAQDDS---VDKPPLSAKGFANREPPYSLSPTPAQ 241
PKP+ P PSL+ +S + DS PP S + PP S PA+
Sbjct: 114 PKPSLPTSPSLSNSTASTQNLSDSPLPPKSPPPSPSATNSPNPPSPTSTPPAK 272
>TC91308 similar to GP|15450621|gb|AAK96582.1 At2g30530/T6B20.12
{Arabidopsis thaliana}, partial (12%)
Length = 772
Score = 29.6 bits (65), Expect = 2.1
Identities = 20/73 (27%), Positives = 34/73 (46%), Gaps = 5/73 (6%)
Frame = +1
Query: 155 VFEKKGNVSPHPVAATSSSDGDRAPVISAAFCGPEETPKPNQPGEPSLA-----TPVSSP 209
+FE + +P+P+A +SSS D S+ + + + SL+ + +SSP
Sbjct: 172 IFEDEAIRNPNPIAVSSSSSNDGVDASSSDSLAAKAIRASSARRDSSLSSLYGHSNLSSP 351
Query: 210 RAQDDSVDKPPLS 222
R+Q PP S
Sbjct: 352 RSQPTPTAPPPSS 390
>BI270017 similar to GP|15450621|gb| At2g30530/T6B20.12 {Arabidopsis
thaliana}, partial (17%)
Length = 617
Score = 29.6 bits (65), Expect = 2.1
Identities = 20/73 (27%), Positives = 34/73 (46%), Gaps = 5/73 (6%)
Frame = +2
Query: 155 VFEKKGNVSPHPVAATSSSDGDRAPVISAAFCGPEETPKPNQPGEPSLA-----TPVSSP 209
+FE + +P+P+A +SSS D S+ + + + SL+ + +SSP
Sbjct: 98 IFEDEAIRNPNPIAVSSSSSNDGVDASSSDSLAAKAIRASSARRDSSLSSLYGHSNLSSP 277
Query: 210 RAQDDSVDKPPLS 222
R+Q PP S
Sbjct: 278 RSQPTPTAPPPSS 316
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.315 0.130 0.373
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 11,536,206
Number of Sequences: 36976
Number of extensions: 182305
Number of successful extensions: 1115
Number of sequences better than 10.0: 61
Number of HSP's better than 10.0 without gapping: 1039
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1103
length of query: 396
length of database: 9,014,727
effective HSP length: 98
effective length of query: 298
effective length of database: 5,391,079
effective search space: 1606541542
effective search space used: 1606541542
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 59 (27.3 bits)
Lotus: description of TM0005.13


