

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0005.1
(346 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
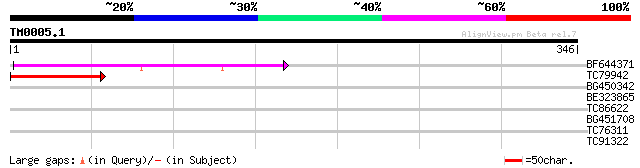
Score E
Sequences producing significant alignments: (bits) Value
BF644371 weakly similar to GP|8927657|gb| EST gb|N38213 comes fr... 65 5e-11
TC79942 similar to GP|8927657|gb|AAF82148.1| EST gb|N38213 comes... 44 1e-04
BG450342 similar to GP|8927657|gb EST gb|N38213 comes from this ... 34 0.095
BE323865 similar to PIR|E96814|E96 hypothetical protein T30F21.8... 30 1.0
TC86622 weakly similar to GP|20197623|gb|AAM15156.1 putative myo... 30 1.4
BG451708 29 2.3
TC76311 homologue to GP|3204132|emb|CAA07235.1 extensin {Cicer a... 29 3.0
TC91322 similar to GP|16550932|gb|AAL25648.1 inward-rectifying K... 28 6.8
>BF644371 weakly similar to GP|8927657|gb| EST gb|N38213 comes from this
gene. {Arabidopsis thaliana}, partial (4%)
Length = 673
Score = 64.7 bits (156), Expect = 5e-11
Identities = 47/194 (24%), Positives = 86/194 (44%), Gaps = 26/194 (13%)
Frame = +3
Query: 3 DTHFARLNAIYQQICSKLRQHESSPHEPKTFNVEKFRGHKRLYEDIFAMFRLRKSQITPE 62
+++ L+ +YQ+I +KL QH+S PH+PK+ +EK + K + E + ++ KS I+P
Sbjct: 3 ESYLPELSEMYQKIATKLHQHDSLPHQPKSDQLEKLKVFKMMLERLITFLQVSKSNISPS 182
Query: 63 ISERVDRVETLINSIIQ------------------HQNLSSHQGKHSADVQSKQQVLPQV 104
+ E++ E I + I H + S + VQS + +
Sbjct: 183 LKEKLGSYEKQIINFINTNRPRKISSLQPGQLPPPHMHSMSQTQPQATQVQSHENQMNTQ 362
Query: 105 LGTQNDHLAKQTSQRNNMCLQEQQ-------VAKQYGNSEQGVNQLSIKEGPGKAVQRQT 157
L T N T Q+NN+ + + N+ Q L + +G ++Q+ +
Sbjct: 363 LQTTNMQGPVPTMQQNNLTSMQHSSLXGVSTAQQNMMNTMQPSASLDLGQGNMSSLQQNS 542
Query: 158 LCA-QATEASGVST 170
+ A Q T S +S+
Sbjct: 543 VTAPQQTNVSSLSS 584
>TC79942 similar to GP|8927657|gb|AAF82148.1| EST gb|N38213 comes from this
gene. {Arabidopsis thaliana}, partial (9%)
Length = 879
Score = 43.5 bits (101), Expect = 1e-04
Identities = 18/58 (31%), Positives = 36/58 (62%)
Frame = +3
Query: 1 MKDTHFARLNAIYQQICSKLRQHESSPHEPKTFNVEKFRGHKRLYEDIFAMFRLRKSQ 58
MK+++ +L+ + Q+I +KL+QH+S P +PK+ +E + K + E + ++ K Q
Sbjct: 552 MKESYLPKLSEMIQKIATKLQQHDSLPQQPKSDELEMLKEFKMVLERLITFLQVSKEQ 725
>BG450342 similar to GP|8927657|gb EST gb|N38213 comes from this gene.
{Arabidopsis thaliana}, partial (10%)
Length = 669
Score = 33.9 bits (76), Expect = 0.095
Identities = 26/92 (28%), Positives = 42/92 (45%), Gaps = 7/92 (7%)
Frame = +2
Query: 153 VQRQTLCAQATEASGVSTNDMGFSSSALIK-------GSGNLNEVSHKPALISEKPSAAM 205
V Q + A A ++ G S+S L+ GN S + ++E+P M
Sbjct: 311 VGHQQTGSGAAPAQSLAIGTPGISASPLLAEFTGPDGAHGNALAPSSGKSTVTEEP---M 481
Query: 206 QRLIGVFTSMSSEALGASIGKIREVVHLNDAI 237
RLI +S++ AL A++ I V+ +ND I
Sbjct: 482 DRLIKAVSSLTPAALSAAVSDISSVISMNDRI 577
>BE323865 similar to PIR|E96814|E96 hypothetical protein T30F21.8 [imported]
- Arabidopsis thaliana, partial (34%)
Length = 403
Score = 30.4 bits (67), Expect = 1.0
Identities = 16/51 (31%), Positives = 27/51 (52%)
Frame = +1
Query: 237 IPAAKFIDEPLEMAGQQNQPSRIPQGRKMARNINAMAFDTSRSCARTHESF 287
+ A+ FID+ + + G + P+RI + K + +A T CA T E+F
Sbjct: 250 LQASHFIDDKIPVLGVNSDPTRIDEVEKFSGEFDATR-STGHLCAATVENF 399
>TC86622 weakly similar to GP|20197623|gb|AAM15156.1 putative myosin heavy
chain {Arabidopsis thaliana}, partial (17%)
Length = 2546
Score = 30.0 bits (66), Expect = 1.4
Identities = 71/313 (22%), Positives = 119/313 (37%), Gaps = 30/313 (9%)
Frame = +3
Query: 2 KDTHFARLNAIYQQICSKLRQHESSPHEPKTFNVEKFRGHKRLYEDIFAM----FRLRKS 57
K+ +F + A+Y+ S L+ S+ K V++ K ED+ + KS
Sbjct: 1167 KN*NFVQEIAVYESKLSDLQSKLSAALVEKDETVKEILASKNAAEDLVTQHNEEVQTLKS 1346
Query: 58 QITPEISERVDRVETLIN------SII--QHQNLSSHQGKHSADVQSKQQVLPQVLGTQN 109
QI+ I +R ET N SII + L HQ K+ ++S+ + L + ++
Sbjct: 1347 QISSVIDDRNLLNETNQNLKKELESIILDLEEKLKEHQ-KNEDSLKSEVETLKIEIAEKS 1523
Query: 110 ----------DHLAKQTSQRNNMCLQEQQVAKQ--------YGNSEQGVNQLSIKEGPGK 151
LAK S+ + Q A Q + + EQ ++++++ G
Sbjct: 1524 ALQSRLHEIEAQLAKAESRLHEEVGSVQAAASQREVDLSSKFEDYEQKISEINVLNGKVA 1703
Query: 152 AVQRQTLCAQATEASGVSTNDMGFSSSALIKGSGNLNEVSHKPALISEKPSAAMQRLIGV 211
++++ AQ T A+ A +K S + E+ K IS +Q+ +
Sbjct: 1704 ELEKELHLAQDTIANQKGEESQKLELEAALKNS--VEELETKKNEIS-----LLQKQVIE 1862
Query: 212 FTSMSSEALGASIGKIREVVHLNDAIPAAKFIDEPLEMAGQQNQPSRIPQGRKMARNINA 271
F +A K E V DA LE+ + S P RK + A
Sbjct: 1863 FEQKLQQADEKISVKGEEAVDKKDA----------LEVKSRDFSISS-PSKRKSKKKSEA 2009
Query: 272 MAFDTSRSCARTH 284
TS S + TH
Sbjct: 2010 TTPQTSTSSSETH 2048
>BG451708
Length = 608
Score = 29.3 bits (64), Expect = 2.3
Identities = 23/72 (31%), Positives = 35/72 (47%), Gaps = 4/72 (5%)
Frame = -1
Query: 276 TSRSCARTHESFNQLTDVEEPDLDLVASKVKRSRIEVIYLLSVFFFFMNAIYKTRYTIFY 335
T CARTHE F D+E + ++ + + E Y +SV + I +T+Y FY
Sbjct: 293 TQFDCARTHE*FAYTLDIEGFET*VLLIR*DCANKE--YRMSVCKDRLLTILETQYLPFY 120
Query: 336 P----VKSCYFL 343
P + +CY L
Sbjct: 119 PNTQGIPTCYRL 84
>TC76311 homologue to GP|3204132|emb|CAA07235.1 extensin {Cicer arietinum},
partial (77%)
Length = 973
Score = 28.9 bits (63), Expect = 3.0
Identities = 17/49 (34%), Positives = 25/49 (50%)
Frame = +1
Query: 75 NSIIQHQNLSSHQGKHSADVQSKQQVLPQVLGTQNDHLAKQTSQRNNMC 123
NS+ S HQG ++ + +K Q LP + N L K +S NN+C
Sbjct: 466 NSVPYKSCFSFHQGINATSIGNKVQHLPSCRSSLNKSL-KASSLSNNIC 609
>TC91322 similar to GP|16550932|gb|AAL25648.1 inward-rectifying K+ channel
{Eucalyptus camaldulensis}, partial (2%)
Length = 661
Score = 27.7 bits (60), Expect = 6.8
Identities = 15/52 (28%), Positives = 25/52 (47%), Gaps = 6/52 (11%)
Frame = -3
Query: 301 VASKVKRS------RIEVIYLLSVFFFFMNAIYKTRYTIFYPVKSCYFLRCS 346
+ KVKRS + ++ + +++ IYKT Y +F + CY CS
Sbjct: 377 ICCKVKRSITFFSLYFQTTLIIMLLYYY---IYKTFYNVFVAITHCYM*CCS 231
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.317 0.130 0.362
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 9,060,598
Number of Sequences: 36976
Number of extensions: 108356
Number of successful extensions: 667
Number of sequences better than 10.0: 16
Number of HSP's better than 10.0 without gapping: 666
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 666
length of query: 346
length of database: 9,014,727
effective HSP length: 97
effective length of query: 249
effective length of database: 5,428,055
effective search space: 1351585695
effective search space used: 1351585695
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 59 (27.3 bits)
Lotus: description of TM0005.1


