

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0004a.3
(148 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
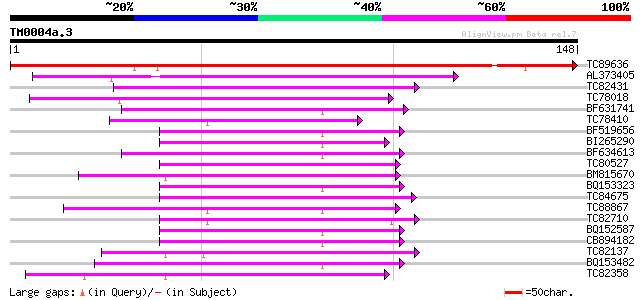
Score E
Sequences producing significant alignments: (bits) Value
TC89636 weakly similar to GP|10177420|dbj|BAB10705. auxin-induce... 199 3e-52
AL373405 similar to PIR|F86331|F863 hypothetical protein AAG1254... 73 5e-14
TC82431 weakly similar to GP|8777399|dbj|BAA96989.1 emb|CAB86483... 73 5e-14
TC78018 similar to PIR|T04499|T04499 auxin-induced protein homol... 64 2e-11
BF631741 similar to SP|P32295|ARG7_P Indole-3-acetic acid induce... 60 3e-10
TC78410 similar to PIR|A84906|A84906 probable auxin-regulated pr... 58 1e-09
BF519656 similar to SP|P32295|ARG7_P Indole-3-acetic acid induce... 57 4e-09
BI265290 similar to SP|P32295|ARG7_P Indole-3-acetic acid induce... 57 4e-09
BF634613 similar to SP|P33083|AX6B_S Auxin-induced protein 6B. [... 56 5e-09
TC80527 similar to GP|11994425|dbj|BAB02427. auxin-regulated pro... 56 5e-09
BM815670 similar to GP|20149050|gb| auxin-induced SAUR-like prot... 56 6e-09
BQ153323 similar to PIR|T06083|T0608 probable auxin-induced prot... 56 6e-09
TC84675 similar to GP|21553798|gb|AAM62891.1 unknown {Arabidopsi... 55 8e-09
TC88867 weakly similar to GP|9758890|dbj|BAB09466.1 auxin-induce... 55 8e-09
TC82710 similar to PIR|A84906|A84906 probable auxin-regulated pr... 55 1e-08
BQ152587 similar to SP|P33080|AX10_S Auxin-induced protein X10A.... 54 3e-08
CB894182 similar to SP|P33083|AX6B_S Auxin-induced protein 6B. [... 53 5e-08
TC82137 similar to PIR|A84906|A84906 probable auxin-regulated pr... 52 7e-08
BQ153482 similar to PIR|T06083|T0608 probable auxin-induced prot... 52 1e-07
TC82358 weakly similar to PIR|F84787|F84787 probable auxin-induc... 52 1e-07
>TC89636 weakly similar to GP|10177420|dbj|BAB10705. auxin-induced
protein-like {Arabidopsis thaliana}, partial (55%)
Length = 881
Score = 199 bits (506), Expect = 3e-52
Identities = 107/160 (66%), Positives = 125/160 (77%), Gaps = 12/160 (7%)
Frame = +2
Query: 1 MAGAMKKVDKIRQIVRLKQVMMRWKQISLRR-----HDSSDP------RPPSGCLYVYVG 49
MAG MKKVDKIRQIVRLKQ+M RWKQISLRR +++P +PPSG ++VYVG
Sbjct: 77 MAGTMKKVDKIRQIVRLKQLMTRWKQISLRRCSLRSETTTEPCVNPRRQPPSGFVFVYVG 256
Query: 50 HERQRFAIPARFLNLPVFAGLLDETEEEFGLRGSGGLILPCDVCFFTHIVKRLHKNENKY 109
ER RFAIPARFLN PVFAGLLD TEEEFGLRG+GGL+LPC V FFT IVKRLHKNE+KY
Sbjct: 257 SERHRFAIPARFLNFPVFAGLLDVTEEEFGLRGNGGLVLPCHVNFFTEIVKRLHKNEHKY 436
Query: 110 GKLTLEQFVITFSDAAANFDSCKD-QNVVVFTPLLQKAEV 148
GKL+LE+FV S+ +SCK+ +NVVV PLL++A V
Sbjct: 437 GKLSLEEFVKMLSEVTL-LESCKEKENVVVLAPLLERAFV 553
>AL373405 similar to PIR|F86331|F863 hypothetical protein AAG12548.1
[imported] - Arabidopsis thaliana, partial (64%)
Length = 455
Score = 72.8 bits (177), Expect = 5e-14
Identities = 44/114 (38%), Positives = 61/114 (52%), Gaps = 3/114 (2%)
Frame = +2
Query: 7 KVDKIRQIVRLKQVMMRWK---QISLRRHDSSDPRPPSGCLYVYVGHERQRFAIPARFLN 63
K KIR IVRL+Q++ RW+ +IS SD PSG + V VG RF + A +LN
Sbjct: 56 KCSKIRHIVRLRQMLRRWRNKARISSANRAPSDV--PSGHVAVCVGANYTRFVVRATYLN 229
Query: 64 LPVFAGLLDETEEEFGLRGSGGLILPCDVCFFTHIVKRLHKNENKYGKLTLEQF 117
P+F LL + EEE+G G L +PCD FF + + ++ + G E F
Sbjct: 230 HPIFQKLLVQAEEEYGFSNHGPLTIPCDEEFFEEALWFISRSGSNNGSNRFEDF 391
>TC82431 weakly similar to GP|8777399|dbj|BAA96989.1
emb|CAB86483.1~gene_id:MFB16.16~similar to unknown
protein {Arabidopsis thaliana}, partial (33%)
Length = 717
Score = 72.8 bits (177), Expect = 5e-14
Identities = 35/80 (43%), Positives = 46/80 (56%)
Frame = +1
Query: 28 SLRRHDSSDPRPPSGCLYVYVGHERQRFAIPARFLNLPVFAGLLDETEEEFGLRGSGGLI 87
S ++ S +PP GCL VYVG ERQRF I + N P+F LL++ E E+G R G L
Sbjct: 229 SHKKQKDSTKKPPHGCLCVYVGPERQRFVIKIKIFNHPLFKTLLEDVENEYGYRNDGPLW 408
Query: 88 LPCDVCFFTHIVKRLHKNEN 107
LPCDV F + + E+
Sbjct: 409 LPCDVDLFCEALVEIESAED 468
>TC78018 similar to PIR|T04499|T04499 auxin-induced protein homolog
F8F16.140 - Arabidopsis thaliana, partial (58%)
Length = 803
Score = 63.9 bits (154), Expect = 2e-11
Identities = 42/126 (33%), Positives = 58/126 (45%), Gaps = 31/126 (24%)
Frame = +2
Query: 6 KKVDKIRQIVRLKQVMMRWKQI-------------------------------SLRRHDS 34
KK +KIR+IVRL+Q++ +W+++ S R
Sbjct: 158 KKPNKIREIVRLQQILKKWRRVANTSKTYRSSSINNNSTTSKSIKFLKRTLSMSEREGGG 337
Query: 35 SDPRPPSGCLYVYVGHERQRFAIPARFLNLPVFAGLLDETEEEFGLRGSGGLILPCDVCF 94
S+ P G L V VG + RF IP +L F LL E EEEFG +G L +PC+V
Sbjct: 338 SNNAVPKGYLAVCVGVDLNRFVIPTEYLAHQAFHILLREAEEEFGFEQTGVLRIPCEVSV 517
Query: 95 FTHIVK 100
F I+K
Sbjct: 518 FESILK 535
>BF631741 similar to SP|P32295|ARG7_P Indole-3-acetic acid induced protein
ARG7. [Mung bean Vigna radiata] {Phaseolus aureus},
partial (78%)
Length = 450
Score = 60.1 bits (144), Expect = 3e-10
Identities = 32/76 (42%), Positives = 41/76 (53%), Gaps = 1/76 (1%)
Frame = +1
Query: 30 RRHDSSDPRPPSGCLYVYVGHERQRFAIPARFLNLPVFAGLLDETEEEFGL-RGSGGLIL 88
R S P GCL VYVG E +RF IP +LN P+F LL++ EE+F +GGL +
Sbjct: 1 RTSSSKGVEVPKGCLAVYVGEEMKRFVIPISYLNQPLFQDLLNQAEEQFEYDHPTGGLTI 180
Query: 89 PCDVCFFTHIVKRLHK 104
PC F I L +
Sbjct: 181 PCREDMFLDITSCLSR 228
>TC78410 similar to PIR|A84906|A84906 probable auxin-regulated protein
[imported] - Arabidopsis thaliana, partial (61%)
Length = 1073
Score = 58.2 bits (139), Expect = 1e-09
Identities = 28/68 (41%), Positives = 39/68 (57%), Gaps = 2/68 (2%)
Frame = +1
Query: 27 ISLRRHDSSDPRPPSGCLYVYVGH--ERQRFAIPARFLNLPVFAGLLDETEEEFGLRGSG 84
++ H + P GCL V VG E+Q+F IP ++N P+F LL E EEE+G G
Sbjct: 592 LNFHHHHEKNKDIPKGCLAVMVGQGEEQQKFVIPVIYINHPLFMQLLKEAEEEYGFDHKG 771
Query: 85 GLILPCDV 92
+I+PC V
Sbjct: 772 PIIIPCQV 795
>BF519656 similar to SP|P32295|ARG7_P Indole-3-acetic acid induced protein
ARG7. [Mung bean Vigna radiata] {Phaseolus aureus},
partial (80%)
Length = 483
Score = 56.6 bits (135), Expect = 4e-09
Identities = 29/65 (44%), Positives = 35/65 (53%), Gaps = 1/65 (1%)
Frame = +2
Query: 40 PSGCLYVYVGHERQRFAIPARFLNLPVFAGLLDETEEEFGL-RGSGGLILPCDVCFFTHI 98
P G L VYVG + +RF IP +LN F LL + EEEFG GGL +PC F HI
Sbjct: 137 PKGYLAVYVGEQMKRFVIPTSYLNQASFQNLLSQAEEEFGYDHPMGGLTIPCTEDVFLHI 316
Query: 99 VKRLH 103
+
Sbjct: 317 TSHFN 331
>BI265290 similar to SP|P32295|ARG7_P Indole-3-acetic acid induced protein
ARG7. [Mung bean Vigna radiata] {Phaseolus aureus},
partial (67%)
Length = 531
Score = 56.6 bits (135), Expect = 4e-09
Identities = 29/61 (47%), Positives = 35/61 (56%), Gaps = 1/61 (1%)
Frame = +3
Query: 40 PSGCLYVYVGHERQRFAIPARFLNLPVFAGLLDETEEEFGL-RGSGGLILPCDVCFFTHI 98
P GC+ VYVG +RF IP LN P F LL + EEEFG GGL +PC F +I
Sbjct: 150 PKGCVAVYVGENMKRFVIPIGCLNQPSFQDLLSKAEEEFGYHHPMGGLTIPCSEDSFLNI 329
Query: 99 V 99
+
Sbjct: 330 I 332
>BF634613 similar to SP|P33083|AX6B_S Auxin-induced protein 6B. [Soybean]
{Glycine max}, partial (82%)
Length = 368
Score = 56.2 bits (134), Expect = 5e-09
Identities = 30/75 (40%), Positives = 40/75 (53%), Gaps = 1/75 (1%)
Frame = +3
Query: 30 RRHDSSDPRPPSGCLYVYVGHERQRFAIPARFLNLPVFAGLLDETEEEFGL-RGSGGLIL 88
R+ S G + VYVG + RF +P +LN P F LL ++EEEFG GGL +
Sbjct: 3 RQATSKSVEVRKGYVAVYVGEKLVRFVVPVSYLNQPSFQDLLSQSEEEFGYDHPMGGLTI 182
Query: 89 PCDVCFFTHIVKRLH 103
PC F HI+ L+
Sbjct: 183 PCTEDVFQHIISSLN 227
>TC80527 similar to GP|11994425|dbj|BAB02427. auxin-regulated protein-like
{Arabidopsis thaliana}, partial (62%)
Length = 632
Score = 56.2 bits (134), Expect = 5e-09
Identities = 27/63 (42%), Positives = 38/63 (59%)
Frame = +1
Query: 40 PSGCLYVYVGHERQRFAIPARFLNLPVFAGLLDETEEEFGLRGSGGLILPCDVCFFTHIV 99
P G + +YVG E +RF + A LN PVF LL+E+ +E+G G L LPC V F ++
Sbjct: 262 PEGHVPIYVGDEMERFVVCAELLNHPVFIKLLNESAQEYGYEQKGVLRLPCHVLVFERVL 441
Query: 100 KRL 102
+ L
Sbjct: 442 EAL 450
>BM815670 similar to GP|20149050|gb| auxin-induced SAUR-like protein
{Capsicum annuum}, partial (80%)
Length = 611
Score = 55.8 bits (133), Expect = 6e-09
Identities = 30/87 (34%), Positives = 41/87 (46%), Gaps = 3/87 (3%)
Frame = +3
Query: 19 QVMMRWKQISLRRHDSSDPRP---PSGCLYVYVGHERQRFAIPARFLNLPVFAGLLDETE 75
Q++ R R+ + + P P G VYVG R R+ IP +L P F LL E
Sbjct: 3 QIVKRCSSFGKRQSYNEEGLPEDVPKGHFVVYVGENRTRYIIPISWLAHPQFQSLLQRAE 182
Query: 76 EEFGLRGSGGLILPCDVCFFTHIVKRL 102
+EFG GL +PCD FF + +
Sbjct: 183 DEFGFNHDMGLTIPCDEVFFESLTSMM 263
>BQ153323 similar to PIR|T06083|T0608 probable auxin-induced protein
T9A14.120 - Arabidopsis thaliana, partial (75%)
Length = 455
Score = 55.8 bits (133), Expect = 6e-09
Identities = 29/65 (44%), Positives = 35/65 (53%), Gaps = 1/65 (1%)
Frame = +1
Query: 40 PSGCLYVYVGHERQRFAIPARFLNLPVFAGLLDETEEEFGL-RGSGGLILPCDVCFFTHI 98
P G L VYVG + +RF IP +LN F LL + EEEFG GGL +PC F HI
Sbjct: 64 PKGYLAVYVGEQMKRFVIPTSYLNQASFQTLLSQVEEEFGYDHPMGGLTIPCTEDVFLHI 243
Query: 99 VKRLH 103
+
Sbjct: 244 TSHFN 258
>TC84675 similar to GP|21553798|gb|AAM62891.1 unknown {Arabidopsis
thaliana}, partial (46%)
Length = 599
Score = 55.5 bits (132), Expect = 8e-09
Identities = 25/67 (37%), Positives = 40/67 (59%)
Frame = +1
Query: 40 PSGCLYVYVGHERQRFAIPARFLNLPVFAGLLDETEEEFGLRGSGGLILPCDVCFFTHIV 99
P G L VYVG E +RF IP +L+ +F LL++ +EFG GGL +PC++ F +++
Sbjct: 154 PKGYLAVYVGPELRRFIIPTSYLSHSLFKMLLEKAADEFGFNQCGGLTIPCEIETFKYLL 333
Query: 100 KRLHKNE 106
+ +
Sbjct: 334 SCMENTQ 354
>TC88867 weakly similar to GP|9758890|dbj|BAB09466.1 auxin-induced
protein-like {Arabidopsis thaliana}, partial (83%)
Length = 453
Score = 55.5 bits (132), Expect = 8e-09
Identities = 32/90 (35%), Positives = 46/90 (50%), Gaps = 2/90 (2%)
Frame = +3
Query: 15 VRLKQVMMRWKQISLRRHDSSDPRPPSGCLYVYVGH-ERQRFAIPARFLNLPVFAGLLDE 73
+RL + ++ KQI P G + VYVG +++RF +P +LN P F LL
Sbjct: 72 IRLPFMALQAKQIFKSTSTQQQSNVPKGHIAVYVGELQKKRFVVPISYLNHPTFLDLLSS 251
Query: 74 TEEEFGL-RGSGGLILPCDVCFFTHIVKRL 102
EEEFG GGL +PC F ++ +L
Sbjct: 252 VEEEFGYNHPMGGLTIPCKEDAFINLTSQL 341
>TC82710 similar to PIR|A84906|A84906 probable auxin-regulated protein
[imported] - Arabidopsis thaliana, partial (58%)
Length = 721
Score = 54.7 bits (130), Expect = 1e-08
Identities = 30/76 (39%), Positives = 41/76 (53%), Gaps = 8/76 (10%)
Frame = +3
Query: 40 PSGCLYVYVGH--ERQRFAIPARFLNLPVFAGLLDETEEEFGLRGSGGLILPCDVCFFTH 97
P GCL + VG ++QRF +P + N P+F LL E EEE+G G + +PC V F +
Sbjct: 285 PKGCLAIKVGQGEDQQRFVVPVIYFNHPLFMQLLKEAEEEYGFDHKGAITIPCRVEEFRN 464
Query: 98 I------VKRLHKNEN 107
I K LH N +
Sbjct: 465 IRGLIDREKSLHHNHH 512
>BQ152587 similar to SP|P33080|AX10_S Auxin-induced protein X10A. [Soybean]
{Glycine max}, partial (92%)
Length = 475
Score = 53.5 bits (127), Expect = 3e-08
Identities = 29/65 (44%), Positives = 37/65 (56%), Gaps = 1/65 (1%)
Frame = +2
Query: 40 PSGCLYVYVGHERQRFAIPARFLNLPVFAGLLDETEEEFGL-RGSGGLILPCDVCFFTHI 98
P G L VYVG E +RF IP +L+ F LL++ EE+FG GGL +PC F I
Sbjct: 161 PKGYLAVYVGEEMKRFVIPISYLSQSSFQELLNQAEEQFGYDHPMGGLTIPCREDVFLDI 340
Query: 99 VKRLH 103
RL+
Sbjct: 341 TSRLN 355
>CB894182 similar to SP|P33083|AX6B_S Auxin-induced protein 6B. [Soybean]
{Glycine max}, partial (88%)
Length = 827
Score = 52.8 bits (125), Expect = 5e-08
Identities = 29/65 (44%), Positives = 34/65 (51%), Gaps = 1/65 (1%)
Frame = -1
Query: 40 PSGCLYVYVGHERQRFAIPARFLNLPVFAGLLDETEEEFGL-RGSGGLILPCDVCFFTHI 98
P G L VYVG E +RF I +LN F LL E+EFG GGL +PC F HI
Sbjct: 608 PKGYLAVYVGEEMKRFVIHMSYLNQTSFQDLLSRAEDEFGYDHPMGGLTIPCREEVFLHI 429
Query: 99 VKRLH 103
R +
Sbjct: 428 TSRFN 414
>TC82137 similar to PIR|A84906|A84906 probable auxin-regulated protein
[imported] - Arabidopsis thaliana, partial (56%)
Length = 674
Score = 52.4 bits (124), Expect = 7e-08
Identities = 29/89 (32%), Positives = 46/89 (51%), Gaps = 6/89 (6%)
Frame = +2
Query: 25 KQISLRRHDSSDPRP----PSGCLYVYVG--HERQRFAIPARFLNLPVFAGLLDETEEEF 78
K + L RH + P G + + VG E+QRF +P + N P+F LL E EEE+
Sbjct: 215 KNLHLHRHHLQHKQEFKGVPKGFMAIKVGLGEEQQRFVVPVMYFNHPLFIQLLKEAEEEY 394
Query: 79 GLRGSGGLILPCDVCFFTHIVKRLHKNEN 107
G G + +PC V F ++ + +++N
Sbjct: 395 GFDQKGTITIPCHVEEFRNVRGLIDRDKN 481
>BQ153482 similar to PIR|T06083|T0608 probable auxin-induced protein
T9A14.120 - Arabidopsis thaliana, partial (72%)
Length = 508
Score = 51.6 bits (122), Expect = 1e-07
Identities = 32/82 (39%), Positives = 39/82 (47%), Gaps = 1/82 (1%)
Frame = +3
Query: 23 RWKQISLRRHDSSDPRPPSGCLYVYVGHERQRFAIPARFLNLPVFAGLLDETEEEFGL-R 81
R I R S P G L VYVG E +RF IP +L F LL ++EE+F
Sbjct: 108 RLPSIIKRASSSKSVGVPKGYLAVYVGEEMKRFVIPISYLKQKSFQELLSQSEEQFEYDH 287
Query: 82 GSGGLILPCDVCFFTHIVKRLH 103
GGL +PC F I RL+
Sbjct: 288 PMGGLTIPCGEDVFLDITSRLN 353
>TC82358 weakly similar to PIR|F84787|F84787 probable auxin-induced protein
[imported] - Arabidopsis thaliana, partial (50%)
Length = 546
Score = 51.6 bits (122), Expect = 1e-07
Identities = 30/100 (30%), Positives = 51/100 (51%), Gaps = 5/100 (5%)
Frame = +3
Query: 5 MKKVDKIRQIVRLK----QVMMRWKQISLRRHDSSDPRP-PSGCLYVYVGHERQRFAIPA 59
+++ K R+++R+ + + S ++ + P+ P G L VYVG + +RF I
Sbjct: 54 LREFSKFRKVIRVSFLKCAIRASFCSSSQQKSNLHIPKDVPKGHLVVYVGEDCKRFVIKV 233
Query: 60 RFLNLPVFAGLLDETEEEFGLRGSGGLILPCDVCFFTHIV 99
LN P F LLD E+ FG L++PC+ F +I+
Sbjct: 234 GTLNHPPFKALLDHAEDAFGFTNGSKLLIPCNENVFLNIL 353
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.326 0.141 0.426
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 4,734,279
Number of Sequences: 36976
Number of extensions: 63067
Number of successful extensions: 453
Number of sequences better than 10.0: 67
Number of HSP's better than 10.0 without gapping: 441
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 442
length of query: 148
length of database: 9,014,727
effective HSP length: 87
effective length of query: 61
effective length of database: 5,797,815
effective search space: 353666715
effective search space used: 353666715
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 54 (25.4 bits)
Lotus: description of TM0004a.3


